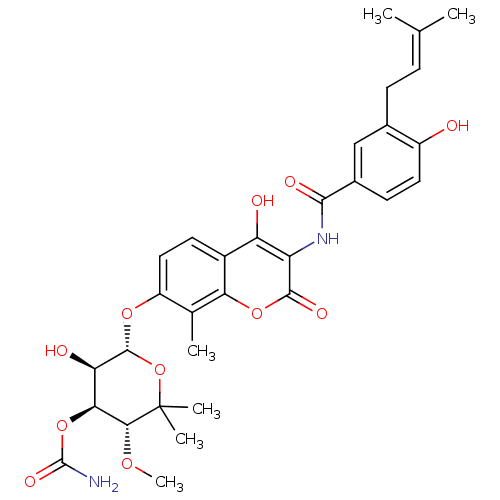
BDBM282 (3R,4S,5R,6R)-5-hydroxy-6-[(2-hydroxy-3-{[4-hydroxy-3-(3-methylbut-2-en-1-yl)benzene]amido}-8-methyl-4-oxo-4H-chromen-7-yl)oxy]-3-methoxy-2,2-dimethyloxan-4-yl carbamate::Coumarin-Based, DNA Gyrase Inhibitor::Novobiocin::US20230364057, Compound Novobiocin
SMILES [#6]-[#8]-[#6@@H]1-[#6@@H](-[#8]-[#6](-[#7])=O)-[#6@@H](-[#8])-[#6@H](-[#8]-c2ccc3c(-[#8])c(-[#7]-[#6](=O)-c4ccc(-[#8])c(-[#6]\[#6]=[#6](\[#6])-[#6])c4)c(=O)oc3c2-[#6])-[#8]C1([#6])[#6]
InChI Key InChIKey=YJQPYGGHQPGBLI-KGSXXDOSSA-N
Activity Spreadsheet -- Enzyme Inhibition Constant Data from BindingDB
 Found 8 hits for monomerid = 282
Found 8 hits for monomerid = 282
Affinity DataKi: 10nM ΔG°: -11.1kcal/molepH: 7.5 T: 2°CAssay Description:Enzymatic hydrolysis of ATP to ADP was coupled to the conversion of NADH to NAD+. The decrease in NADH absorbance was monitored at 340 nm for 20 min ...More data for this Ligand-Target Pair
Affinity DataKi: 14nMAssay Description:Enzymatic hydrolysis of ATP to ADP was coupled to the conversion of NADH to NAD+. The decrease in NADH absorbance was monitored at 340 nm for 20 min ...More data for this Ligand-Target Pair
Affinity DataKi: 110nMAssay Description:Enzymatic hydrolysis of ATP to ADP was coupled to the conversion of NADH to NAD+. The decrease in NADH absorbance was monitored at 340 nm for 20 min ...More data for this Ligand-Target Pair
Affinity DataIC50: 180nMAssay Description:Inhibition of Mycobacterium smegmatis GyrB ATPase activity expressed in Escherichia coli BL21 (DE3) pLysS cells after 100 mins by inorganic phosphate...More data for this Ligand-Target Pair
Affinity DataIC50: 30nMpH: 7.6 T: 2°CAssay Description:Supercoiling assay was performed using the commercially available kit (DNA gyrase supercoiling assay kit: SAS4001) from Inspiralis Pvt.limited, Norwi...More data for this Ligand-Target Pair
Affinity DataIC50: 125nMpH: 7.5Assay Description:Initially, equimolar quantities of about 0.5�M each of E. coli gyraseA and S. aureus gyraseB were incubated for a period of 45 min with salmon sperm ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.85E+3nMAssay Description:SDFQ HTS primary assay using pAB1_FL905 was performed in 2 �L of (1 xDNA gyrase buffer: 20 mM Tris-Acetate pH 7.9, 50 mM KAc, 10 mM MgCl2, 2 mM DTT, ...More data for this Ligand-Target Pair
Affinity DataKd: 5.30E+5nMpH: 7.5Assay Description:Binding of inhibitors and substrates to immobilized AcrB.More data for this Ligand-Target Pair
Activity Spreadsheet -- ITC Data from BindingDB
 Found 1 hit for monomerid = 282
Found 1 hit for monomerid = 282
ITC DataΔG°: -10.2kcal/mole −TΔS°: 2.17kcal/mole ΔH°: -12.4kcal/mole logk: 2.80E+7
pH: 7.5 T: 25.00°C
pH: 7.5 T: 25.00°C
 3D Structure (crystal)
3D Structure (crystal)