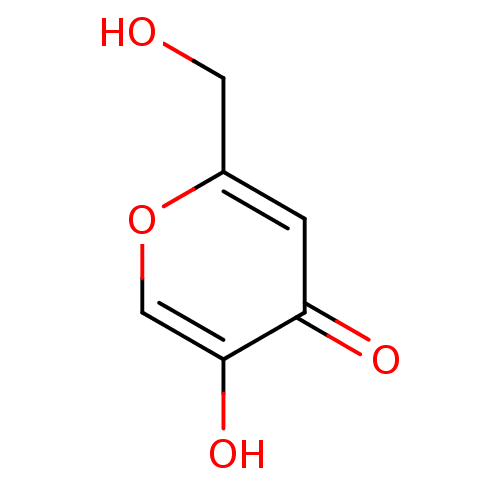
BDBM50031467 5-HYDROXY-2-(HYDROXYMETHYL)-4H-PYRAN-4-ONE::5-Hydroxy-2-hydroxymethyl-pyran-4-one::CHEMBL287556::Kojic Acid::US10857082, Kojic acid::US9422261, Kojic acid::US9682910, kojic acid
SMILES OCc1cc(=O)c(O)co1
InChI Key InChIKey=BEJNERDRQOWKJM-UHFFFAOYSA-N
Activity Spreadsheet -- Enzyme Inhibition Constant Data from BindingDB
 Found 160 hits for monomerid = 50031467
Found 160 hits for monomerid = 50031467
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
University Of Technology Sydney
Curated by ChEMBL
University Of Technology Sydney
Curated by ChEMBL
Affinity DataKi: 8.89E+3nMAssay Description:Competitive inhibition of mushroom tyrosinase assessed as inhibition of L-DOPA oxidase activity at 5 uM active compound by Lineweaver-Burk plot analy...More data for this Ligand-Target Pair
TargetPolyphenol oxidase 1 [N65D,S176D,K211T,P277A,P406L](Agaricus bisporus (Mushroom))
University Of Technology Sydney
University Of Technology Sydney
Affinity DataKi: 1.22E+4nM IC50: 2.75E+4nMpH: 6.8Assay Description:The mushroom tyrosinase inhibition activity of hydroxy naphthylchalcone compounds was measured using L-DOPA as substrate. The activity assay used 3 m...More data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
University Of Technology Sydney
Curated by ChEMBL
University Of Technology Sydney
Curated by ChEMBL
Affinity DataKi: 1.22E+4nMAssay Description:Competitive inhibition of mushroom tyrosinase polyphenol oxidase activity using L-DOPA as substrate by Dixon plot analysisMore data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
University Of Technology Sydney
Curated by ChEMBL
University Of Technology Sydney
Curated by ChEMBL
Affinity DataKi: 1.49E+4nMAssay Description:Competitive inhibition of mushroom tyrosinase diphenolase activity assessed as L-DOPA conversion to dopachrome by Lineweaver-Burk plot analysisMore data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
University Of Technology Sydney
Curated by ChEMBL
University Of Technology Sydney
Curated by ChEMBL
Affinity DataKi: 2.00E+4nMAssay Description:Inhibition of mushroom tyrosinase diphenolase activityMore data for this Ligand-Target Pair
Affinity DataKi: 2.10E+4nMAssay Description:Competitive inhibition of pig kidney DAAO using D-Alanine as substrate by Michaelis-Menten plot analysisMore data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
University Of Technology Sydney
Curated by ChEMBL
University Of Technology Sydney
Curated by ChEMBL
Affinity DataKi: 2.23E+4nMAssay Description:Inhibition of mashroom tyrosinaseMore data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
University Of Technology Sydney
Curated by ChEMBL
University Of Technology Sydney
Curated by ChEMBL
Affinity DataKi: 3.00E+4nMAssay Description:Inhibition of mushroom tyrosinase assessed as inhibition constant for compound-free enzyme using as L-tyrosine substrate by spectrophotometric analys...More data for this Ligand-Target Pair
Affinity DataKi: 3.50E+5nMAssay Description:Inhibition of recombinant human tyrosinase expressed in baculovirus infected Sf9 cells assessed as diphenolase activity using L-DOPA as substrate by ...More data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
University Of Technology Sydney
Curated by ChEMBL
University Of Technology Sydney
Curated by ChEMBL
Affinity DataKi: 5.83E+5nMAssay Description:Inhibition of mushroom tyrosinase assessed as oxidation of L-DOPAMore data for this Ligand-Target Pair
Affinity DataKi: 5.50E+6nMAssay Description:Inhibition of tyrosinase in mouse Melan-a cells assessed as decrease in L-DOPA Vmax at 120 uM by Lineweaver-Burk plotMore data for this Ligand-Target Pair
Affinity DataIC50: 1.67E+4nMpH: 6.8 T: 2°CAssay Description:Tyrosinase inhibition assays were performed in a 96-well microplate format using SpectraMax 340 (Molecular Devices, CA, USA) microplate reader accord...More data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
University Of Technology Sydney
Curated by ChEMBL
University Of Technology Sydney
Curated by ChEMBL
Affinity DataIC50: 1.25E+4nMT: 2°CAssay Description:Mushroom tyrosinase using either L-DOPA or L-tyrosine as substrate. In spectrophotometric experiments, enzyme activity was monitored by dopachrome f...More data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
University Of Technology Sydney
Curated by ChEMBL
University Of Technology Sydney
Curated by ChEMBL
Affinity DataIC50: 2.30E+4nMpH: 6.8 T: 2°CAssay Description:All compounds were dissolved in DMSO (final DMSO concentration in test solution was 2.0%). Phosphate buffer, pH 6.8, was use to dilute the DMSO stoc...More data for this Ligand-Target Pair
TargetUncharacterized protein YOR062C(Agaricus bisporus (Mushroom))
Hunan University Of Science And Technology
Hunan University Of Science And Technology
Affinity DataIC50: 1.90E+4nMpH: 6.8Assay Description:The synthesized compounds were tested for diphenolase inhibitory activity of tyrosinase using L-DOPA (dihydroxyphenylalanine) as substrate. Mushroom ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.36E+4nMpH: 6.8Assay Description:Briefly, 1,3,4-thiadiazole derivatives were tested for diphenolase inhibitory activity of tyrosinase using L-DOPA (dihydroxyphenylalanine) as substra...More data for this Ligand-Target Pair
Affinity DataIC50: 1.67E+4nMpH: 6.8 T: 2°CAssay Description:Tyrosinase inhibition assays were performed in a 96-well microplate format using the SpectraMax 340 microplate reader (Molecular Devices, CA) accordi...More data for this Ligand-Target Pair
Affinity DataIC50: 1.50E+4nMpH: 6.8Assay Description:Reader: Synergy HT program: tyrosinase 280-490 kinetics: kinetics over 45 minutes, reading at t=10 minutes, Tests in transparent 96-well plates, Phos...More data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+4nMT: 2°CAssay Description:All test compounds were initially dissolved in 100% DMSO at a concentration of 400 mM. The compounds were further diluted in 100% DMSO to a concentra...More data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
University Of Technology Sydney
Curated by ChEMBL
University Of Technology Sydney
Curated by ChEMBL
Affinity DataIC50: 2.70E+4nMAssay Description:Inhibition of mushroom tyrosinaseMore data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
University Of Technology Sydney
Curated by ChEMBL
University Of Technology Sydney
Curated by ChEMBL
Affinity DataIC50: 2.35E+5nMAssay Description:Inhibition of mushroom tyrosinaseMore data for this Ligand-Target Pair
Affinity DataIC50: 1.25E+5nMAssay Description:Inhibition of tyrosinase (unknown origin)More data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
University Of Technology Sydney
Curated by ChEMBL
University Of Technology Sydney
Curated by ChEMBL
Affinity DataIC50: 7.14E+4nMAssay Description:Inhibition of mushroom tyrosinase using L-DOPA as substrate after 2 mins by spectrophotometryMore data for this Ligand-Target Pair
Affinity DataIC50: 1.25E+5nMAssay Description:Inhibition of tyrosinase (unknown origin)More data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
University Of Technology Sydney
Curated by ChEMBL
University Of Technology Sydney
Curated by ChEMBL
Affinity DataIC50: 2.50E+5nMAssay Description:Inhibition of mushroom tyrosinase by spectrophotometryMore data for this Ligand-Target Pair
Affinity DataIC50: 2.29E+4nMAssay Description:Inhibitory activity against mushroom tyrosinaseMore data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
University Of Technology Sydney
Curated by ChEMBL
University Of Technology Sydney
Curated by ChEMBL
Affinity DataIC50: 1.26E+4nMAssay Description:Inhibition of mushroom tyrosinase using L-tyrosine as substrate by spectrophotometryMore data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
University Of Technology Sydney
Curated by ChEMBL
University Of Technology Sydney
Curated by ChEMBL
Affinity DataIC50: 3.28E+4nMAssay Description:Inhibition of mushroom tyrosinase using L-DOPA as substrate by spectrophotometry relative to controlMore data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
University Of Technology Sydney
Curated by ChEMBL
University Of Technology Sydney
Curated by ChEMBL
Affinity DataIC50: >3.00E+4nMAssay Description:Inhibition of mushroom tyrosinase using L-tyrosine as substrate after 20 mins by microplate readerMore data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
University Of Technology Sydney
Curated by ChEMBL
University Of Technology Sydney
Curated by ChEMBL
Affinity DataIC50: 2.36E+4nMAssay Description:Binding affinity for 5-hydroxytryptamine 7 receptorMore data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
University Of Technology Sydney
Curated by ChEMBL
University Of Technology Sydney
Curated by ChEMBL
Affinity DataIC50: 2.73E+4nMAssay Description:Competitive inhibition of mushroom tyrosinase assessed as inhibition of L-DOPA oxidase activityMore data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
University Of Technology Sydney
Curated by ChEMBL
University Of Technology Sydney
Curated by ChEMBL
Affinity DataIC50: 2.80E+4nMAssay Description:Inhibition of mushroom tyrosinase using L-DOPA as substrate preincubated for 10 mins by spectrophotometryMore data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
University Of Technology Sydney
Curated by ChEMBL
University Of Technology Sydney
Curated by ChEMBL
Affinity DataIC50: 2.85E+4nMAssay Description:Inhibition of mushroom tyrosinase diphenolase activity using L-DOPA as substrate preincubated for 30 mins followed by substrate addition measured for...More data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
University Of Technology Sydney
Curated by ChEMBL
University Of Technology Sydney
Curated by ChEMBL
Affinity DataIC50: 1.63E+4nMAssay Description:Inhibition of mushroom tyrosinaseMore data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
University Of Technology Sydney
Curated by ChEMBL
University Of Technology Sydney
Curated by ChEMBL
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of mushroom tyrosinase using L-DOPA as substrate assessed as dopaquinone production after 5 mins by microplate reader analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 1.33E+5nMAssay Description:Inhibition of tyrosinaseMore data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
University Of Technology Sydney
Curated by ChEMBL
University Of Technology Sydney
Curated by ChEMBL
Affinity DataIC50: 3.82E+4nMAssay Description:Inhibition of mushroom tyrosinaseMore data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
University Of Technology Sydney
Curated by ChEMBL
University Of Technology Sydney
Curated by ChEMBL
Affinity DataIC50: 1.20E+4nMAssay Description:Inhibition of mushroom tyrosinaseMore data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
University Of Technology Sydney
Curated by ChEMBL
University Of Technology Sydney
Curated by ChEMBL
Affinity DataIC50: 1.67E+4nMAssay Description:Inhibition of mushroom tyrosinase using L-DOPA as substrate preincubated for 10 mins followed by substrate addition measured after 20 minsMore data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
University Of Technology Sydney
Curated by ChEMBL
University Of Technology Sydney
Curated by ChEMBL
Affinity DataIC50: 2.43E+5nMAssay Description:Inhibition of Mushroom tyrosinaseMore data for this Ligand-Target Pair
Affinity DataIC50: 9.15E+3nMAssay Description:Inhibition of tyrosinase (unknown origin) using L-DOPA as substrate assessed as formation of dopachrome by spectrophotometric analysisMore data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
University Of Technology Sydney
Curated by ChEMBL
University Of Technology Sydney
Curated by ChEMBL
Affinity DataIC50: 1.12E+4nMAssay Description:Inhibition of mushroom tyrosinase assessed as reduction in L-DOPA oxidase activity using L-DOPA substrate at 5 uM by Dixon plotMore data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
University Of Technology Sydney
Curated by ChEMBL
University Of Technology Sydney
Curated by ChEMBL
Affinity DataIC50: 2.37E+4nMAssay Description:Inhibition of mushroom tyrosinase assessed as reduction in L-DOPA oxidase activity using L-DOPA substrateMore data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
University Of Technology Sydney
Curated by ChEMBL
University Of Technology Sydney
Curated by ChEMBL
Affinity DataIC50: 2.00E+5nMAssay Description:Inhibition of mushroom tyrosinase activity after 10 minsMore data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
University Of Technology Sydney
Curated by ChEMBL
University Of Technology Sydney
Curated by ChEMBL
Affinity DataIC50: 7.40E+3nMAssay Description:Inhibition of mushroom tyrosinaseMore data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
University Of Technology Sydney
Curated by ChEMBL
University Of Technology Sydney
Curated by ChEMBL
Affinity DataIC50: 1.60E+4nMAssay Description:Inhibition of mushroom tyrosinase using L-DOPA as substrate assessed as formation of L-DOPA chrome preincubated for 10 mins followed by addition of s...More data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
University Of Technology Sydney
Curated by ChEMBL
University Of Technology Sydney
Curated by ChEMBL
Affinity DataIC50: 2.75E+4nMAssay Description:Inhibition of mushroom tyrosinase diphenolase activity using L-DOPA as substrateMore data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
University Of Technology Sydney
Curated by ChEMBL
University Of Technology Sydney
Curated by ChEMBL
Affinity DataIC50: 3.23E+4nMAssay Description:Inhibition of mushroom tyrosinaseMore data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
University Of Technology Sydney
Curated by ChEMBL
University Of Technology Sydney
Curated by ChEMBL
Affinity DataIC50: 2.65E+4nMAssay Description:Inhibition of diphenolase activity of mushroom tyrosinase using L-DOPA as substrate preincubated with substrate for 10 mins followed by protein addit...More data for this Ligand-Target Pair
TargetPolyphenol oxidase 2(Agaricus bisporus (Common mushroom))
University Of Technology Sydney
Curated by ChEMBL
University Of Technology Sydney
Curated by ChEMBL
Affinity DataIC50: 1.32E+4nMAssay Description:Inhibition of monophenolase activity of mushroom tyrosinase using L-tyrosine as substrate preincubated with substrate for 10 mins followed by protein...More data for this Ligand-Target Pair