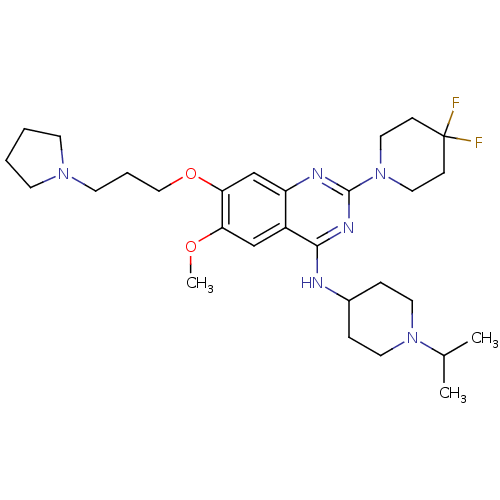
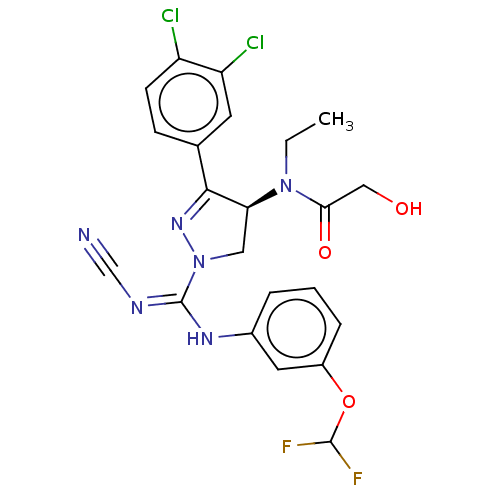
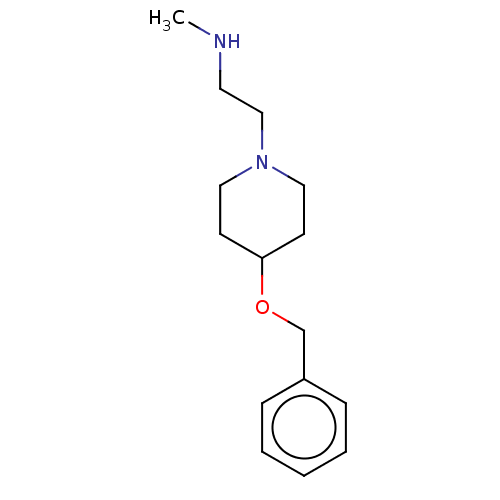
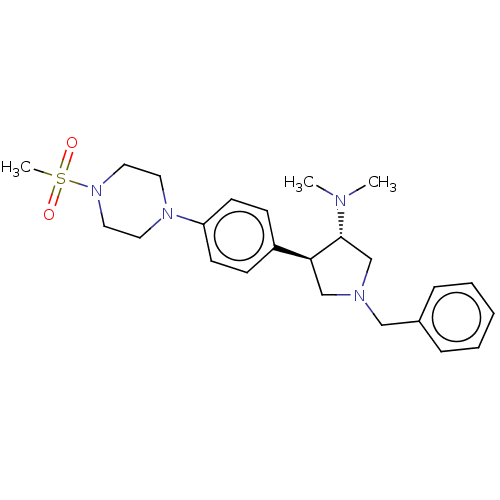
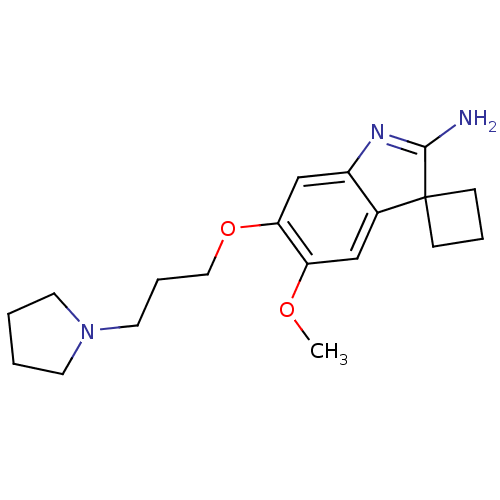
Affinity DataKi: 0.400nM ΔG°: -53.6kJ/molepH: 7.5 T: 2°CAssay Description:For the assay, compounds were dispensed in assay-ready plates using a three-fold serial dilution from 50 μM to ~850 pM using an Echo 550 Acousti...More data for this Ligand-Target Pair
Affinity DataKi: 0.5nM ΔG°: -53.1kJ/molepH: 7.5 T: 2°CAssay Description:For the assay, compounds were dispensed in assay-ready plates using a three-fold serial dilution from 50 μM to ~850 pM using an Echo 550 Acousti...More data for this Ligand-Target Pair
Affinity DataKi: 0.800nM IC50: 4nMAssay Description:In brief, the tritiated S-adenosyl-L-methionine (3HSAM, PerkinElmer Life Sciences) was used as the donor of methyl group. The (3H) methylated biotin ...More data for this Ligand-Target Pair
Affinity DataKi: 1nM ΔG°: -51.4kJ/molepH: 7.5 T: 2°CAssay Description:For the assay, compounds were dispensed in assay-ready plates using a three-fold serial dilution from 50 μM to ~850 pM using an Echo 550 Acousti...More data for this Ligand-Target Pair
Affinity DataKi: 1.30nM IC50: 5nMAssay Description:In brief, the tritiated S-adenosyl-L-methionine (3HSAM, PerkinElmer Life Sciences) was used as the donor of methyl group. The (3H) methylated biotin ...More data for this Ligand-Target Pair
Affinity DataKi: 3nM IC50: 9nMAssay Description:In brief, the tritiated S-adenosyl-L-methionine (3HSAM, PerkinElmer Life Sciences) was used as the donor of methyl group. The (3H) methylated biotin ...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EHMT2(Homo sapiens (Human))
University Of North Carolina At Chapel Hill
Curated by ChEMBL
University Of North Carolina At Chapel Hill
Curated by ChEMBL
Affinity DataKi: 3.70nMAssay Description:Non-competitive inhibition of lysine methyltransferase G9a (unknown origin) using SAM as substrate by Michaelis-Menten kinetic assayMore data for this Ligand-Target Pair
Affinity DataKi: 4nM ΔG°: -47.9kJ/molepH: 7.5 T: 2°CAssay Description:For the assay, compounds were dispensed in assay-ready plates using a three-fold serial dilution from 50 μM to ~850 pM using an Echo 550 Acousti...More data for this Ligand-Target Pair
Affinity DataKi: 8nMAssay Description:Competitive inhibition of full length 6xHis-tagged SMYD2 (unknown origin) expressed in Escherichia coli using varying levels of Btn-Ahx-GSRAHSSHLKSKK...More data for this Ligand-Target Pair
Affinity DataKi: 11nM IC50: 30nMAssay Description:In brief, the tritiated S-adenosyl-L-methionine (3HSAM, PerkinElmer Life Sciences) was used as the donor of methyl group. The (3H) methylated biotin ...More data for this Ligand-Target Pair
Affinity DataKi: 16nMAssay Description:Inhibition of human PRMT6 pre-incubated for 1 to 30 mins using [3H]SAM as donor and [3H]methylated biotin-labeled peptide as substrate by scintillati...More data for this Ligand-Target Pair
Affinity DataKi: 17nM IC50: 42nMAssay Description:In brief, the tritiated S-adenosyl-L-methionine (3HSAM, PerkinElmer Life Sciences) was used as the donor of methyl group. The (3H) methylated biotin ...More data for this Ligand-Target Pair
Affinity DataKi: 23nM IC50: 83nMAssay Description:In brief, the tritiated S-adenosyl-L-methionine (3HSAM, PerkinElmer Life Sciences) was used as the donor of methyl group. The (3H) methylated biotin ...More data for this Ligand-Target Pair
Affinity DataKi: 28nMAssay Description:Uncompetitive inhibition of full length 6xHis-tagged SMYD2 (unknown origin) expressed in Escherichia coli using fixed levels of Btn-Ahx-GSRAHSSHLKSKK...More data for this Ligand-Target Pair
Affinity DataKi: 55nM IC50: 119nMAssay Description:In brief, the tritiated S-adenosyl-L-methionine (3HSAM, PerkinElmer Life Sciences) was used as the donor of methyl group. The (3H) methylated biotin ...More data for this Ligand-Target Pair
TargetSigma non-opioid intracellular receptor 1(Cavia porcellus (Guinea pig))
Icahn School Of Medicine At Mount Sinai
Curated by ChEMBL
Icahn School Of Medicine At Mount Sinai
Curated by ChEMBL
Affinity DataKi: 64nMAssay Description:Displacement of [3H]pentazocine from guinea pig sigma1 receptorMore data for this Ligand-Target Pair
TargetHistamine H3 receptor(Homo sapiens (Human))
Icahn School Of Medicine At Mount Sinai
Curated by ChEMBL
Icahn School Of Medicine At Mount Sinai
Curated by ChEMBL
Affinity DataKi: 87nMAssay Description:Displacement of [3H]alpha-methylhistamine from human histamine H3 receptor expressed HEK Flp-In cell membranesMore data for this Ligand-Target Pair
Affinity DataKi: 110nM IC50: 230nMAssay Description:In brief, the tritiated S-adenosyl-L-methionine (3HSAM, PerkinElmer Life Sciences) was used as the donor of methyl group. The (3H) methylated biotin ...More data for this Ligand-Target Pair
Affinity DataKi: 120nM IC50: 250nMAssay Description:In brief, the tritiated S-adenosyl-L-methionine (3HSAM, PerkinElmer Life Sciences) was used as the donor of methyl group. The (3H) methylated biotin ...More data for this Ligand-Target Pair
Affinity DataKi: 120nM IC50: 260nMAssay Description:In brief, the tritiated S-adenosyl-L-methionine (3HSAM, PerkinElmer Life Sciences) was used as the donor of methyl group. The (3H) methylated biotin ...More data for this Ligand-Target Pair
Affinity DataKi: 550nM IC50: 1.10E+3nMAssay Description:In brief, the tritiated S-adenosyl-L-methionine (3HSAM, PerkinElmer Life Sciences) was used as the donor of methyl group. The (3H) methylated biotin ...More data for this Ligand-Target Pair
Affinity DataKi: 1.20E+3nM ΔG°: -33.8kJ/molepH: 7.5 T: 2°CAssay Description:For the assay, compounds were dispensed in assay-ready plates using a three-fold serial dilution from 50 μM to ~850 pM using an Echo 550 Acousti...More data for this Ligand-Target Pair
Affinity DataKi: 1.50E+3nM IC50: 3.00E+3nMAssay Description:In brief, the tritiated S-adenosyl-L-methionine (3HSAM, PerkinElmer Life Sciences) was used as the donor of methyl group. The (3H) methylated biotin ...More data for this Ligand-Target Pair
Affinity DataKi: 1.83E+3nM ΔG°: -32.7kJ/molepH: 7.5 T: 2°CAssay Description:For the assay, compounds were dispensed in assay-ready plates using a three-fold serial dilution from 50 μM to ~850 pM using an Echo 550 Acousti...More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+4nM IC50: >2.00E+4nMAssay Description:In brief, the tritiated S-adenosyl-L-methionine (3HSAM, PerkinElmer Life Sciences) was used as the donor of methyl group. The (3H) methylated biotin ...More data for this Ligand-Target Pair
Affinity DataKi: >3.75E+4nM IC50: >7.50E+4nMAssay Description:In brief, the tritiated S-adenosyl-L-methionine (3HSAM, PerkinElmer Life Sciences) was used as the donor of methyl group. The (3H) methylated biotin ...More data for this Ligand-Target Pair
Affinity DataKi: >5.00E+4nM IC50: >1.00E+5nMAssay Description:In brief, the tritiated S-adenosyl-L-methionine (3HSAM, PerkinElmer Life Sciences) was used as the donor of methyl group. The (3H) methylated biotin ...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EHMT2(Homo sapiens (Human))
University Of North Carolina At Chapel Hill
Curated by ChEMBL
University Of North Carolina At Chapel Hill
Curated by ChEMBL
Affinity DataIC50: 2nMAssay Description:Inhibition of G9a assessed as hydrolysis of S-adenosyl-L-homocysteine after 2 mins by SAHH-coupled fluorescence assayMore data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EHMT2(Homo sapiens (Human))
University Of North Carolina At Chapel Hill
Curated by ChEMBL
University Of North Carolina At Chapel Hill
Curated by ChEMBL
Affinity DataIC50: 3nMAssay Description:Inhibition of G9a (unknown origin)More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EHMT2(Homo sapiens (Human))
University Of North Carolina At Chapel Hill
Curated by ChEMBL
University Of North Carolina At Chapel Hill
Curated by ChEMBL
Affinity DataIC50: 3nMAssay Description:Inhibition of lysine methyltransferase G9a (unknown origin) using [3H]-SAM as substrate after 0.25 hrs by scintillation proximity assayMore data for this Ligand-Target Pair
TargetProtein arginine N-methyltransferase 1(Homo sapiens)
Icahn School Of Medicine At Mount Sinai
Curated by ChEMBL
Icahn School Of Medicine At Mount Sinai
Curated by ChEMBL
Affinity DataIC50: 4nMAssay Description:Inhibition of PRMT1 (unknown origin) by scintillation proximity assayMore data for this Ligand-Target Pair
TargetProtein arginine N-methyltransferase 3(Homo sapiens)
Icahn School Of Medicine At Mount Sinai
Curated by ChEMBL
Icahn School Of Medicine At Mount Sinai
Curated by ChEMBL
Affinity DataIC50: 4nMAssay Description:Inhibition of PRMT3 (unknown origin) by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 4nMAssay Description:Inhibition of PRMT4 (unknown origin) by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 4nMAssay Description:Inhibition of PRMT6 (unknown origin) by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 4nMAssay Description:Inhibition of PRMT8 (unknown origin) by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 4nMAssay Description:Inhibition of human PRMT8 assessed as inhibition of methylation activity using biotin-labeled peptide as substrate and [3H]-SAM by scintillation prox...More data for this Ligand-Target Pair
Affinity DataIC50: 4nMAssay Description:Inhibition of human PRMT1 assessed as inhibition of methylation activity using biotin-labeled peptide as substrate and [3H]-SAM by scintillation prox...More data for this Ligand-Target Pair
Affinity DataIC50: 4nMAssay Description:Inhibition of human PRMT4 assessed as inhibition of methylation activity using biotin-labeled peptide as substrate and [3H]-SAM by scintillation prox...More data for this Ligand-Target Pair
Affinity DataIC50: 4nMAssay Description:Inhibition of human PRMT3 assessed as inhibition of methylation activity using biotin-labeled peptide as substrate and [3H]-SAM by scintillation prox...More data for this Ligand-Target Pair
Affinity DataIC50: 4nMAssay Description:Inhibition of human PRMT6 assessed as inhibition of methylation activity using biotin-labeled peptide as substrate and [3H]-SAM by scintillation prox...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EHMT2(Homo sapiens (Human))
University Of North Carolina At Chapel Hill
Curated by ChEMBL
University Of North Carolina At Chapel Hill
Curated by ChEMBL
Affinity DataIC50: 4nMAssay Description:Inhibition of G9a assessed as hydrolysis of S-adenosyl-L-homocysteine after 2 mins by SAHH-coupled fluorescence assayMore data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EHMT2(Homo sapiens (Human))
University Of North Carolina At Chapel Hill
Curated by ChEMBL
University Of North Carolina At Chapel Hill
Curated by ChEMBL
Affinity DataIC50: 4nMAssay Description:Inhibition of lysine methyltransferase G9a (unknown origin) using [3H]-SAM as substrate after 0.25 hrs by scintillation proximity assayMore data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EHMT2(Homo sapiens (Human))
University Of North Carolina At Chapel Hill
Curated by ChEMBL
University Of North Carolina At Chapel Hill
Curated by ChEMBL
Affinity DataIC50: 4nMAssay Description:Inhibition of G9a assessed as hydrolysis of S-adenosyl-L-homocysteine after 2 mins by SAHH-coupled fluorescence assayMore data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EHMT2(Homo sapiens (Human))
University Of North Carolina At Chapel Hill
Curated by ChEMBL
University Of North Carolina At Chapel Hill
Curated by ChEMBL
Affinity DataIC50: 5nMAssay Description:Inhibition of G9a assessed as hydrolysis of S-adenosyl-L-homocysteine after 2 mins by SAHH-coupled fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 6nMAssay Description:Inhibition of recombinant human SMYD2 (1 to 433 residues) expressed in baculovirus infected Sf9 insect cells using biotinylated-p53 peptide (361 to 3...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EHMT2(Homo sapiens (Human))
University Of North Carolina At Chapel Hill
Curated by ChEMBL
University Of North Carolina At Chapel Hill
Curated by ChEMBL
Affinity DataIC50: 6nMAssay Description:Inhibition of G9a assessed as hydrolysis of S-adenosyl-L-homocysteine after 2 mins by SAHH-coupled fluorescence assayMore data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EHMT2(Homo sapiens (Human))
University Of North Carolina At Chapel Hill
Curated by ChEMBL
University Of North Carolina At Chapel Hill
Curated by ChEMBL
Affinity DataIC50: 7nMAssay Description:Inhibition of G9a assessed as hydrolysis of S-adenosyl-L-homocysteine after 2 mins by SAHH-coupled fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 7.51nMpH: 7.5 T: 2°CAssay Description:For the assay, compounds or nonbiotinylated H3(23-34) peptide were dispensed in assay-ready plates using a three-fold serial dilution from 50 μM...More data for this Ligand-Target Pair
Affinity DataIC50: 8nMAssay Description:Inhibition of recombinant human SMYD2 (1 to 433 residues) expressed in baculovirus infected Sf9 insect cells using biotinylated-p53 peptide (361 to 3...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EHMT2(Homo sapiens (Human))
University Of North Carolina At Chapel Hill
Curated by ChEMBL
University Of North Carolina At Chapel Hill
Curated by ChEMBL
Affinity DataIC50: 8nMAssay Description:Inhibition of G9a assessed as hydrolysis of S-adenosyl-L-homocysteine after 2 mins by SAHH-coupled fluorescence assayMore data for this Ligand-Target Pair

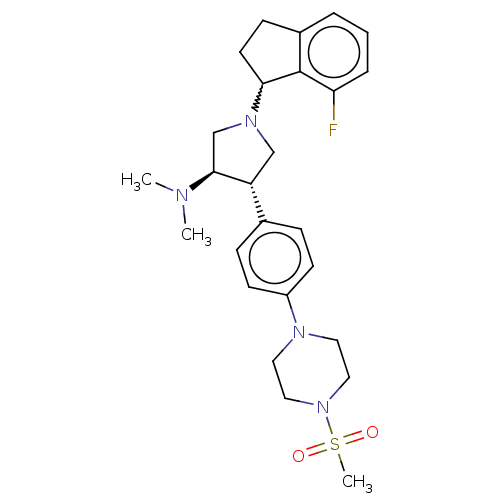
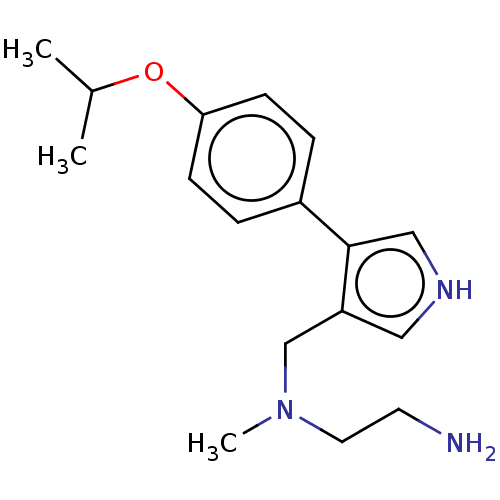
 3D Structure (crystal)
3D Structure (crystal)