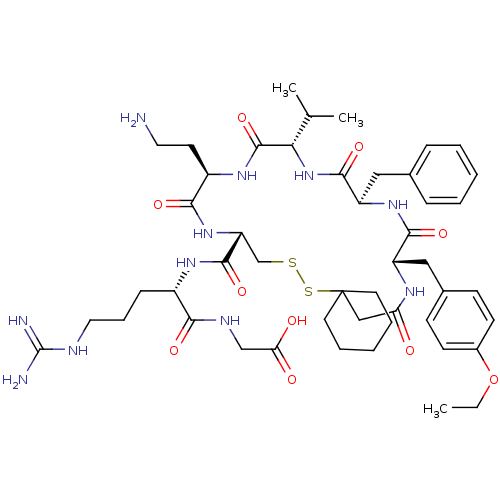
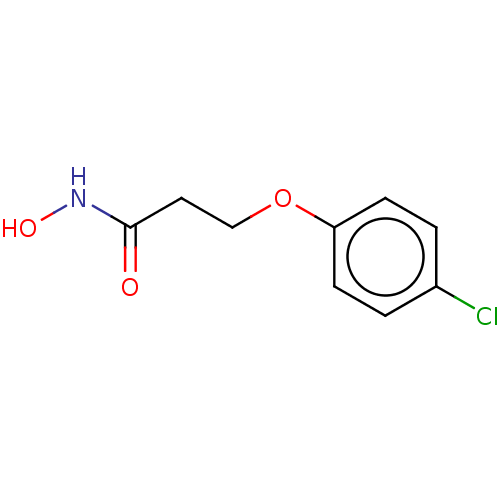
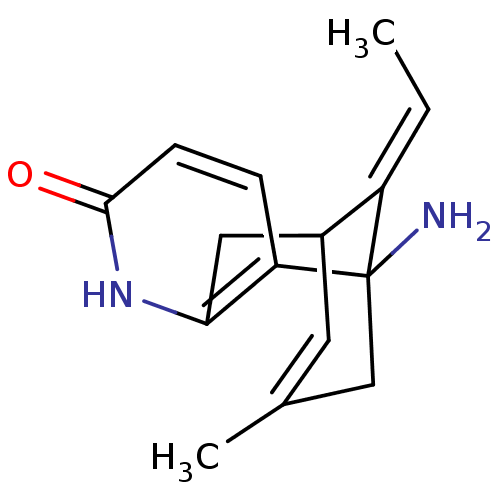
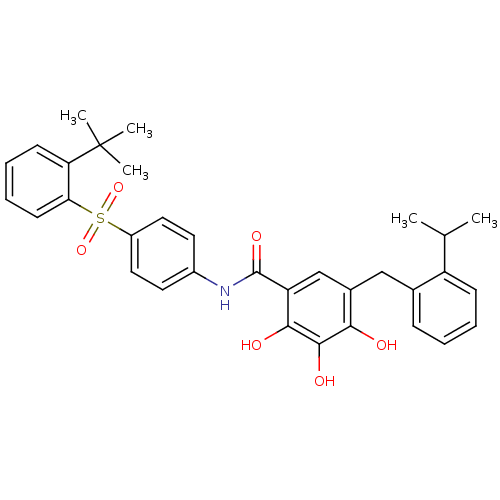
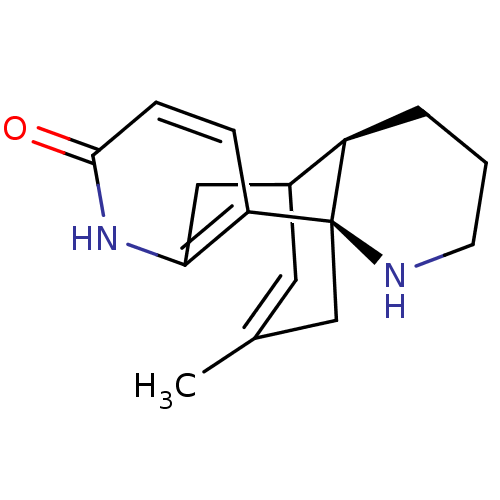
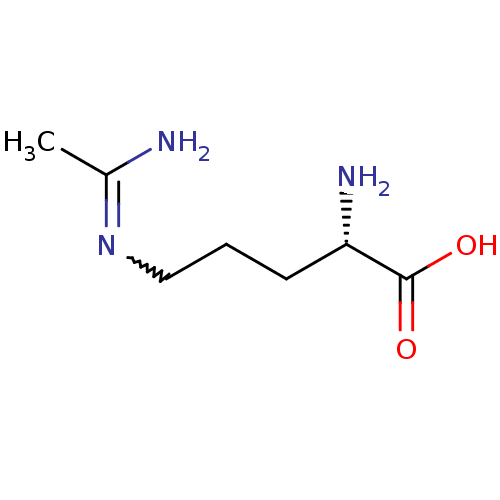
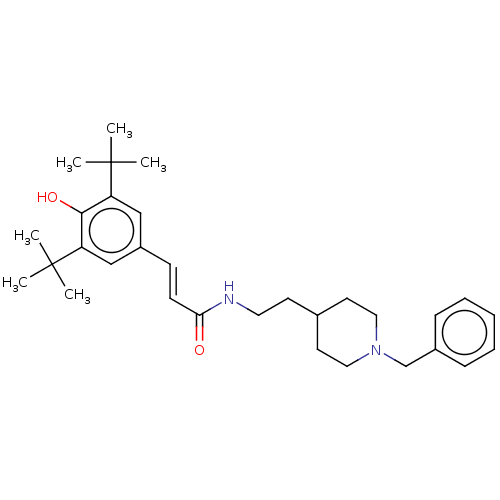
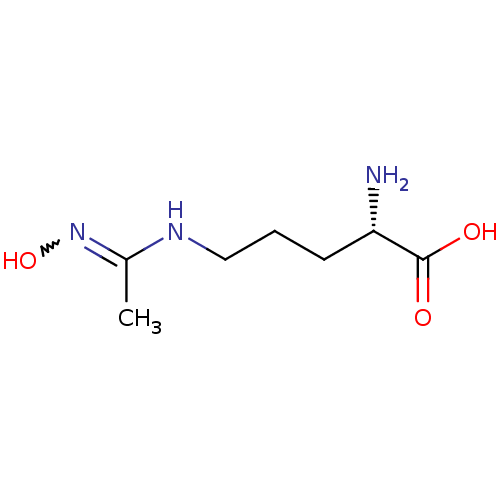
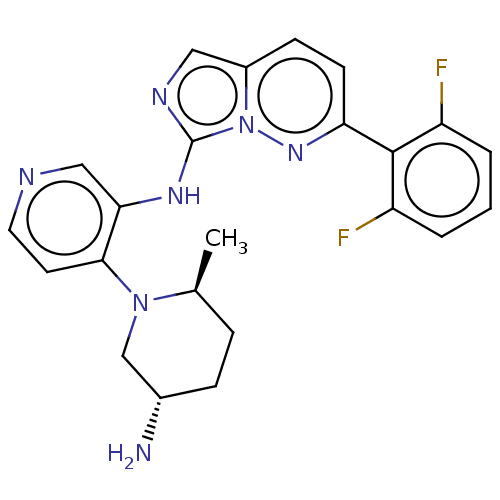
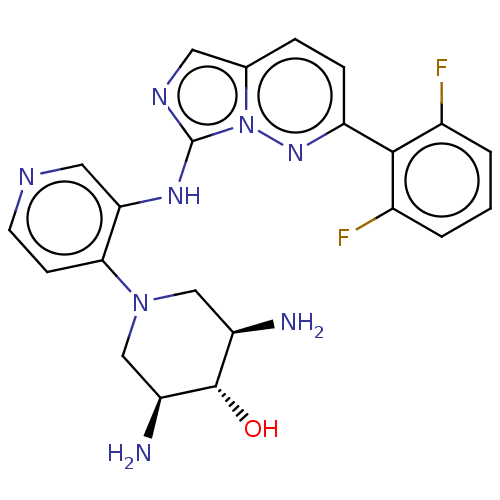
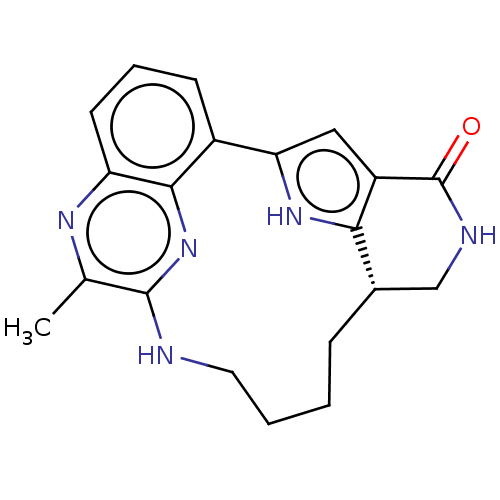
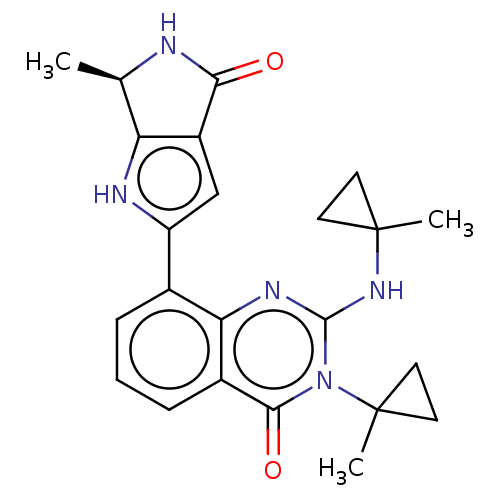
TargetAdenylate cyclase type 4(Homo sapiens (Human))
Research And Development Division
Curated by ChEMBL
Research And Development Division
Curated by ChEMBL
Affinity DataKi: 0.880nMAssay Description:Binding affinity for dopamine receptor D2 was evaluated by the ability to displace [3H]spiperoneMore data for this Ligand-Target Pair
Affinity DataKi: 1.20nMAssay Description:Inhibition of LVP stimulated adenylate cyclase activity in pig kidney medullary membraneMore data for this Ligand-Target Pair
TargetAdenylate cyclase type 4(Homo sapiens (Human))
Research And Development Division
Curated by ChEMBL
Research And Development Division
Curated by ChEMBL
Affinity DataKi: 1.70nMAssay Description:Binding affinity for dopamine receptor D2 was evaluated by the ability to displace [3H]spiperoneMore data for this Ligand-Target Pair
TargetAdenylate cyclase type 4(Homo sapiens (Human))
Research And Development Division
Curated by ChEMBL
Research And Development Division
Curated by ChEMBL
Affinity DataKi: 1.70nMAssay Description:Binding affinity for dopamine receptor D2 was evaluated by the ability to displace [3H]spiperoneMore data for this Ligand-Target Pair
TargetAdenylate cyclase type 4(Homo sapiens (Human))
Research And Development Division
Curated by ChEMBL
Research And Development Division
Curated by ChEMBL
Affinity DataKi: 1.90nMAssay Description:Binding affinity against sigma receptorMore data for this Ligand-Target Pair
Affinity DataKi: 2.5nMAssay Description:Inhibition of LVP stimulated adenylate cyclase activity in pig kidney medullary membraneMore data for this Ligand-Target Pair
TargetAdenylate cyclase type 4(Homo sapiens (Human))
Research And Development Division
Curated by ChEMBL
Research And Development Division
Curated by ChEMBL
Affinity DataKi: 2.5nMAssay Description:Binding affinity for dopamine receptor D2 was evaluated by the ability to displace [3H]spiperoneMore data for this Ligand-Target Pair
TargetAdenylate cyclase type 4(Homo sapiens (Human))
Research And Development Division
Curated by ChEMBL
Research And Development Division
Curated by ChEMBL
Affinity DataKi: 2.60nMAssay Description:Binding affinity for dopamine receptor D2 was evaluated by the ability to displace [3H]spiperoneMore data for this Ligand-Target Pair
Affinity DataKi: 2.70nMAssay Description:Inhibition of LVP stimulated adenylate cyclase activity in pig kidney medullary membraneMore data for this Ligand-Target Pair
TargetAdenylate cyclase type 4(Homo sapiens (Human))
Research And Development Division
Curated by ChEMBL
Research And Development Division
Curated by ChEMBL
Affinity DataKi: 3.90nMAssay Description:Binding affinity for dopamine receptor D2 was evaluated by the ability to displace [3H]spiperoneMore data for this Ligand-Target Pair
Affinity DataKi: 3.90nMAssay Description:Compound was tested for inhibition against V2 vasopressin receptor in pig renal medullary membrane preparations.More data for this Ligand-Target Pair
Affinity DataKi: 3.90nMAssay Description:Inhibition of LVP stimulated adenylate cyclase activity in pig kidney medullary membraneMore data for this Ligand-Target Pair
TargetAdenylate cyclase type 4(Homo sapiens (Human))
Research And Development Division
Curated by ChEMBL
Research And Development Division
Curated by ChEMBL
Affinity DataKi: 4.20nMAssay Description:Binding affinity against sigma receptorMore data for this Ligand-Target Pair
Affinity DataKi: 5.30nMAssay Description:Compound was tested for inhibition against V2 vasopressin receptor in pig renal medullary membrane preparations.More data for this Ligand-Target Pair
Affinity DataKi: 5.80nMAssay Description:Inhibition of LVP stimulated adenylate cyclase activity in pig kidney medullary membraneMore data for this Ligand-Target Pair
Affinity DataKi: 6.40nMAssay Description:Inhibition of LVP stimulated adenylate cyclase activity in pig kidney medullary membraneMore data for this Ligand-Target Pair
Affinity DataKi: 6.40nMAssay Description:Compound was tested for inhibition against V2 vasopressin receptor in pig renal medullary membrane preparations.More data for this Ligand-Target Pair
TargetAdenylate cyclase type 4(Homo sapiens (Human))
Research And Development Division
Curated by ChEMBL
Research And Development Division
Curated by ChEMBL
Affinity DataKi: 7.80nMAssay Description:Binding affinity for dopamine receptor D2 was evaluated by the ability to displace [3H]spiperoneMore data for this Ligand-Target Pair
Affinity DataKi: 8.90nMAssay Description:Compound was tested for inhibition against V2 vasopressin receptor in pig renal medullary membrane preparations.More data for this Ligand-Target Pair
Affinity DataKi: 13nMAssay Description:Inhibition of LVP stimulated adenylate cyclase activity in pig kidney medullary membraneMore data for this Ligand-Target Pair
Affinity DataKi: 23nMAssay Description:Inhibition of LVP stimulated adenylate cyclase activity in pig kidney medullary membraneMore data for this Ligand-Target Pair
Affinity DataKi: 38nMAssay Description:Mixed-type inhibition of human AChE using acetylthiocholine chloride as substrate incubated for 5 mins followed by substrate addition by Lineweaver-B...More data for this Ligand-Target Pair
Affinity DataKi: 41nMAssay Description:Inhibition of LVP stimulated adenylate cyclase activity in pig kidney medullary membraneMore data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataKi: 43nMAssay Description:Mixed-type inhibition of Helicobacter pylori ATCC 43504 urease assessed as enzyme-inhibitor complex using urea as substrate preincubated for 1.5 hrsMore data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataKi: 54nMAssay Description:Mixed-type inhibition of Helicobacter pylori ATCC 43504 urease assessed as enzyme-substrate-inhibitor complex using urea as substrate preincubated fo...More data for this Ligand-Target Pair
TargetAcetylcholinesterase(Tetronarce californica (Pacific electric ray) (Tor...)
Weizmann Institute Of Science
Weizmann Institute Of Science
Affinity DataKi: 175nM ΔG°: -38.6kJ/molepH: 7.0 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at ...More data for this Ligand-Target Pair
TargetInduced myeloid leukemia cell differentiation protein Mcl-1(Homo sapiens (Human))
Xihua University
Curated by ChEMBL
Xihua University
Curated by ChEMBL
Affinity DataKi: 260nMAssay Description:Inhibition of FAM-Bid BH3 peptide binding to recombinant human MCL1 by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 290nMAssay Description:Inhibition of FAM-Bid BH3 peptide binding to recombinant human BCL2 by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetAcetylcholinesterase(Tetronarce californica (Pacific electric ray) (Tor...)
Weizmann Institute Of Science
Weizmann Institute Of Science
Affinity DataKi: 334nM ΔG°: -37.0kJ/molepH: 7.0 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at ...More data for this Ligand-Target Pair
Affinity DataKi: 1.11E+3nMAssay Description:Inhibition of FAM-Bid BH3 peptide binding to recombinant human BCL-XL by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 1.40E+3nMAssay Description:Mixed-type inhibition of human AChE using acetylthiocholine chloride as substrate incubated for 5 mins followed by substrate addition by Lineweaver-B...More data for this Ligand-Target Pair
Affinity DataKi: 1.42E+3nMAssay Description:Compound was tested for competitive antagonist of Nitric oxide synthaseMore data for this Ligand-Target Pair
TargetCocaine esterase(Homo sapiens (Human))
Liaoning University Of Traditional Chinese Medicine
Curated by ChEMBL
Liaoning University Of Traditional Chinese Medicine
Curated by ChEMBL
Affinity DataKi: 1.76E+3nMAssay Description:Fixed inhibition of human carboxylesterase 2 using 4-benzoyl-N-butyl-1,8-naphthalimide as substrate by Dixon and Lineweaver-Burk plot analysisMore data for this Ligand-Target Pair
Affinity DataKi: 2.16E+3nMAssay Description:The compound was tested for competitive antagonist of Nitric oxide synthaseMore data for this Ligand-Target Pair
Affinity DataKi: 3.20E+3nMAssay Description:Mixed-type inhibition of human AChE using acetylthiocholine chloride as substrate incubated for 5 mins followed by substrate addition by Lineweaver-B...More data for this Ligand-Target Pair
TargetAcetylcholinesterase(Tetronarce californica (Pacific electric ray) (Tor...)
Weizmann Institute Of Science
Weizmann Institute Of Science
Affinity DataKi: 4.30E+3nM ΔG°: -30.6kJ/molepH: 7.0 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at ...More data for this Ligand-Target Pair
Affinity DataKi: 3.77E+4nMAssay Description:The compound was tested for competitive antagonist of Nitric oxide synthaseMore data for this Ligand-Target Pair
Affinity DataKi: 4.11E+4nMAssay Description:The compound was tested for competitive antagonist of Nitric oxide synthaseMore data for this Ligand-Target Pair
Affinity DataIC50: 0.0200nMAssay Description:Inhibition of full length recombinant PIM1 (unknown origin) assessed as reduction in BAD phosphorylation at Ser112 residues preincubated for 30 mins ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.0210nMAssay Description:Inhibition of recombinant Pim1 (unknown origin) by electrochemiluminescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 0.0240nMAssay Description:Inhibition of recombinant Pim1 (unknown origin) by electrochemiluminescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 0.0270nMAssay Description:Inhibition of Pim3 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 0.0300nMAssay Description:Inhibition of full length recombinant PIM1 (unknown origin) assessed as reduction in BAD phosphorylation at Ser112 residues preincubated for 30 mins ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.0300nMAssay Description:Inhibition of full length recombinant PIM1 (unknown origin) assessed as reduction in BAD phosphorylation at Ser112 residues preincubated for 30 mins ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.0350nMAssay Description:Inhibition of recombinant Pim1 (unknown origin) by electrochemiluminescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 0.0370nMAssay Description:Inhibition of recombinant Pim1 (unknown origin) by electrochemiluminescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 0.0400nMAssay Description:Inhibition of recombinant Pim1 (unknown origin) by electrochemiluminescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 0.0500nMAssay Description:Inhibition of human full-length Pim2 expressed in Escherichia coli assessed as phosphorylation of biotinylated BAD peptide at Ser 112 preincubated fo...More data for this Ligand-Target Pair
Affinity DataIC50: 0.0500nMAssay Description:Inhibition of full length recombinant PIM1 (unknown origin) assessed as reduction in BAD phosphorylation at Ser112 residues preincubated for 30 mins ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.0500nMAssay Description:Inhibition of full length recombinant PIM2 (unknown origin) assessed as reduction in BAD phosphorylation at Ser112 residues preincubated for 30 mins ...More data for this Ligand-Target Pair






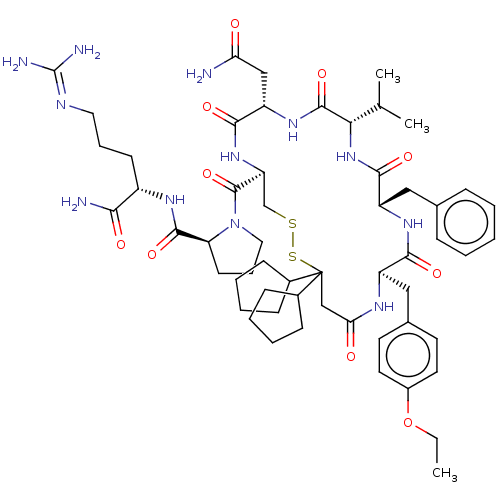
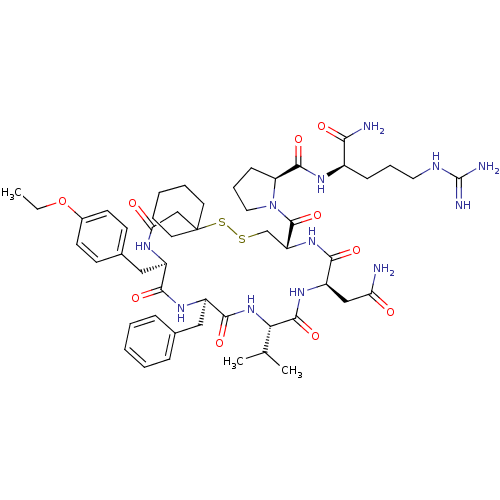

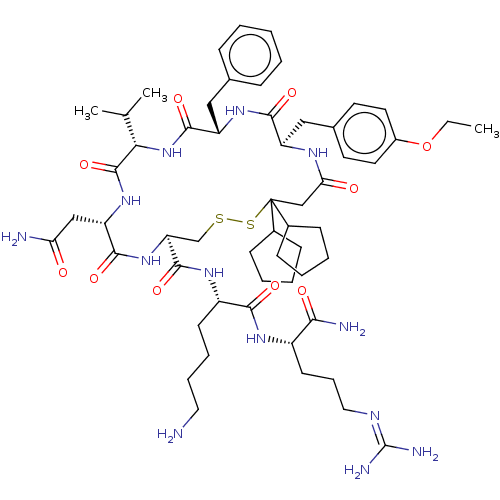
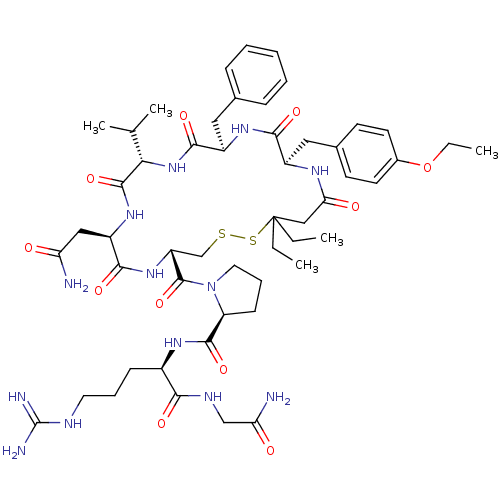


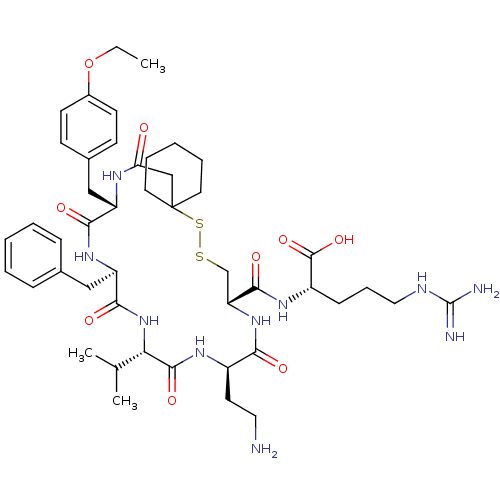
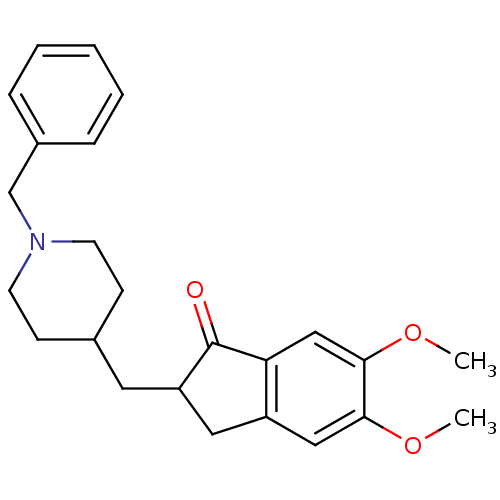
 3D Structure (crystal)
3D Structure (crystal)