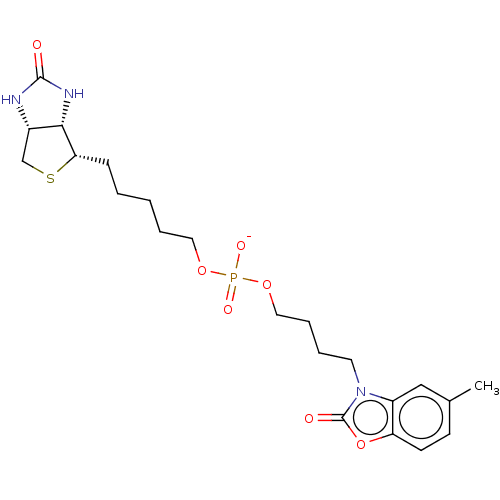
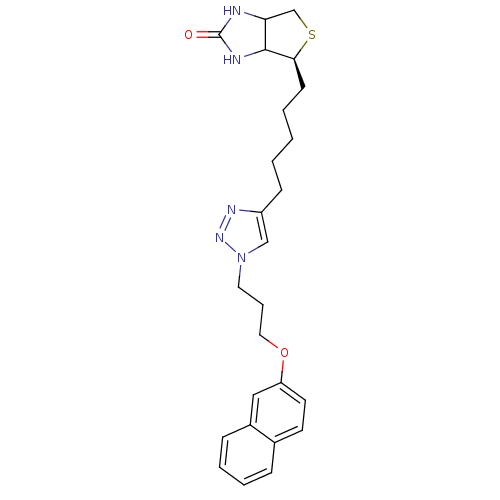
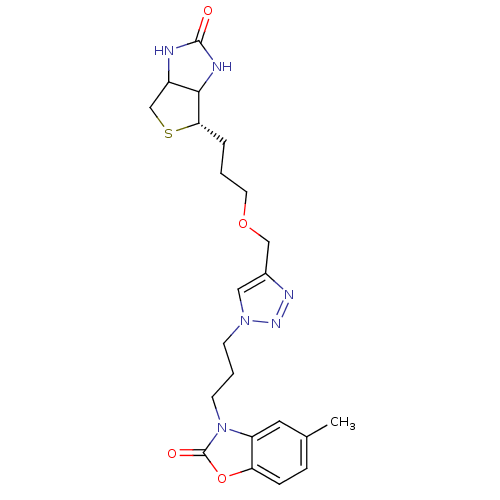
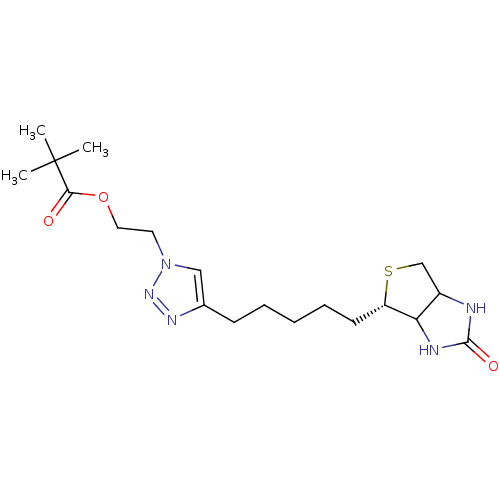
Affinity DataKi: 1.20nMAssay Description:Compound was tested for inhibitory activity against the binding of [3H]-spiperone to Dopamine receptor D2 in rat striatal membranesMore data for this Ligand-Target Pair
Affinity DataKi: 5.20nMAssay Description:Compound was tested for its ability to displace [3H]- spiperone from D2 binding site in rat striatal membranesMore data for this Ligand-Target Pair
Affinity DataKi: 5.5nMAssay Description:Compound was tested for its ability to displace [3H]- spiperone from Dopamine receptor D2 in rat striatal membranesMore data for this Ligand-Target Pair
Affinity DataKi: 30nM ΔG°: -44.7kJ/mole IC50: 203nMpH: 8.0 T: 2°CAssay Description:Quantitation of BPL catalysed 3H-biotin incorporation into the biotin domain substrate was performed as previously described by Polyak et al, J. Biol...More data for this Ligand-Target Pair
Affinity DataKi: 50nMAssay Description:Tested for Competitive binding inhibition of [3H]-Spiperone to Dopamine receptor D2 in rat striatal membrane.More data for this Ligand-Target Pair
Affinity DataKi: 90nM ΔG°: -41.8kJ/mole IC50: 530nMpH: 8.0 T: 2°CAssay Description:Quantitation of BPL catalysed 3H-biotin incorporation into the biotin domain substrate was performed as previously described by Polyak et al, J. Biol...More data for this Ligand-Target Pair
Affinity DataKi: 90nMAssay Description:In vitro enzyme inhibition using biotin protein ligase.More data for this Ligand-Target Pair
Affinity DataKi: 140nMAssay Description:Displacement of [3H]-biotin from human BPL after 20 mins by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 200nMAssay Description:Displacement of [3H]-biotin from human BPL after 20 mins by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 225nM ΔG°: -39.5kJ/molepH: 8.0 T: 2°CAssay Description:Quantitation of BPL catalysed 3H-biotin incorporation into the biotin domain substrate was performed as previously described by Polyak et al, J. Biol...More data for this Ligand-Target Pair
Affinity DataKi: 280nMAssay Description:Compound was tested for inhibitory activity against the binding of [3H]-spiperone toDopamine receptor D2 in rat striatal membranesMore data for this Ligand-Target Pair
Affinity DataKi: 420nM ΔG°: -37.9kJ/molepH: 8.0 T: 2°CAssay Description:Quantitation of BPL catalysed 3H-biotin incorporation into the biotin domain substrate was performed as previously described by Polyak et al, J. Biol...More data for this Ligand-Target Pair
Affinity DataKi: 660nM ΔG°: -36.7kJ/mole IC50: 4.00E+3nMpH: 8.0 T: 2°CAssay Description:Quantitation of BPL catalysed 3H-biotin incorporation into the biotin domain substrate was performed as previously described by Polyak et al, J. Biol...More data for this Ligand-Target Pair
Affinity DataKi: 820nMAssay Description:Tested for Competitive binding inhibition of [3H]-Spiperone to Dopamine receptor D2 in rat striatal membrane.More data for this Ligand-Target Pair
Affinity DataKi: 1.17E+3nM ΔG°: -35.2kJ/mole IC50: 7.00E+3nMpH: 8.0 T: 2°CAssay Description:Quantitation of BPL catalysed 3H-biotin incorporation into the biotin domain substrate was performed as previously described by Polyak et al, J. Biol...More data for this Ligand-Target Pair
Affinity DataKi: 1.17E+3nMAssay Description:In vitro enzyme inhibition using biotin protein ligase.More data for this Ligand-Target Pair
Affinity DataKi: 1.17E+3nMAssay Description:In vitro enzyme inhibition using biotin protein ligase.More data for this Ligand-Target Pair
Affinity DataKi: 1.83E+3nM ΔG°: -34.1kJ/mole IC50: 1.16E+4nMpH: 8.0 T: 2°CAssay Description:Quantitation of BPL catalysed 3H-biotin incorporation into the biotin domain substrate was performed as previously described by Polyak et al, J. Biol...More data for this Ligand-Target Pair
Affinity DataKi: 2.15E+3nMAssay Description:Compound was tested for inhibitory activity against the binding of [3H]-spiperone to Dopamine receptor D2 in rat striatal membranesMore data for this Ligand-Target Pair
Affinity DataKi: 3.50E+3nMAssay Description:Displacement of [3H]-biotin from human BPL after 20 mins by scintillation countingMore data for this Ligand-Target Pair
TargetMetallo-beta-lactamase VIM-2(Pseudomonas aeruginosa (g-Proteobacteria))
Louis Stokes Cleveland Veterans Affairs Medical Center
Louis Stokes Cleveland Veterans Affairs Medical Center
Affinity DataKi: 3.70E+3nMAssay Description:The Ki for each inhibitor was determined by direct competition assays under steady-state conditions. The initial velocity was measured in the presenc...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase VIM-2(Pseudomonas aeruginosa (g-Proteobacteria))
Louis Stokes Cleveland Veterans Affairs Medical Center
Louis Stokes Cleveland Veterans Affairs Medical Center
Affinity DataKi: 3.80E+3nMAssay Description:The Ki for each inhibitor was determined by direct competition assays under steady-state conditions. The initial velocity was measured in the presenc...More data for this Ligand-Target Pair
Affinity DataKi: 3.85E+3nMAssay Description:Displacement of [3H]-biotin from human BPL after 20 mins by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 4.88E+3nMAssay Description:Tested for Competitive binding inhibition of [3H]-Spiperone to Dopamine receptor D2 in rat striatal membrane.More data for this Ligand-Target Pair
TargetVIM-24(Klebsiella pneumoniae (Enterobacteria))
Louis Stokes Cleveland Veterans Affairs Medical Center
Louis Stokes Cleveland Veterans Affairs Medical Center
Affinity DataKi: 4.90E+3nMAssay Description:The Ki for each inhibitor was determined by direct competition assays under steady-state conditions. The initial velocity was measured in the presenc...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase VIM-2(Pseudomonas aeruginosa (g-Proteobacteria))
Louis Stokes Cleveland Veterans Affairs Medical Center
Louis Stokes Cleveland Veterans Affairs Medical Center
Affinity DataKi: 5.40E+3nMAssay Description:The Ki for each inhibitor was determined by direct competition assays under steady-state conditions. The initial velocity was measured in the presenc...More data for this Ligand-Target Pair
TargetVIM-24(Klebsiella pneumoniae (Enterobacteria))
Louis Stokes Cleveland Veterans Affairs Medical Center
Louis Stokes Cleveland Veterans Affairs Medical Center
Affinity DataKi: 6.00E+3nMAssay Description:The Ki for each inhibitor was determined by direct competition assays under steady-state conditions. The initial velocity was measured in the presenc...More data for this Ligand-Target Pair
Affinity DataKi: 6.43E+3nMAssay Description:Displacement of [3H]-biotin from human BPL after 20 mins by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 8.57E+3nMAssay Description:Tested for Competitive binding inhibition of [3H]-Spiperone to Dopamine receptor D2 in rat striatal membrane.More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+4nM ΔG°: >-29.7kJ/mole IC50: >5.00E+4nMpH: 8.0 T: 2°CAssay Description:Quantitation of BPL catalysed 3H-biotin incorporation into the biotin domain substrate was performed as previously described by Polyak et al, J. Biol...More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+4nMAssay Description:In vitro enzyme inhibition using biotin protein ligase.More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+4nMAssay Description:In vitro enzyme inhibition using biotin protein ligase.More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+4nMAssay Description:In vitro enzyme inhibition using biotin protein ligase.More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+4nMAssay Description:In vitro enzyme inhibition using biotin protein ligase.More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+4nMAssay Description:In vitro enzyme inhibition using biotin protein ligase.More data for this Ligand-Target Pair
TargetVIM-24(Klebsiella pneumoniae (Enterobacteria))
Louis Stokes Cleveland Veterans Affairs Medical Center
Louis Stokes Cleveland Veterans Affairs Medical Center
Affinity DataKi: 1.10E+4nMAssay Description:The Ki for each inhibitor was determined by direct competition assays under steady-state conditions. The initial velocity was measured in the presenc...More data for this Ligand-Target Pair
TargetVIM-24(Klebsiella pneumoniae (Enterobacteria))
Louis Stokes Cleveland Veterans Affairs Medical Center
Louis Stokes Cleveland Veterans Affairs Medical Center
Affinity DataKi: 1.20E+4nMAssay Description:The Ki for each inhibitor was determined by direct competition assays under steady-state conditions. The initial velocity was measured in the presenc...More data for this Ligand-Target Pair
Affinity DataKi: 1.20E+4nMAssay Description:Displacement of [3H]-biotin from human BPL after 20 mins by scintillation countingMore data for this Ligand-Target Pair
TargetMetallo-beta-lactamase VIM-2(Pseudomonas aeruginosa (g-Proteobacteria))
Louis Stokes Cleveland Veterans Affairs Medical Center
Louis Stokes Cleveland Veterans Affairs Medical Center
Affinity DataKi: 1.40E+4nMAssay Description:The Ki for each inhibitor was determined by direct competition assays under steady-state conditions. The initial velocity was measured in the presenc...More data for this Ligand-Target Pair
Affinity DataKi: >2.00E+4nMAssay Description:Displacement of [3H]-biotin from human BPL after 20 mins by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: >2.00E+4nMAssay Description:Displacement of [3H]-biotin from human BPL after 20 mins by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: >2.00E+4nMAssay Description:Displacement of [3H]-biotin from human BPL after 20 mins by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: >3.00E+4nM ΔG°: >-26.9kJ/molepH: 8.0 T: 2°CAssay Description:Quantitation of BPL catalysed 3H-biotin incorporation into the biotin domain substrate was performed as previously described by Polyak et al, J. Biol...More data for this Ligand-Target Pair
Affinity DataKi: >3.00E+4nM ΔG°: >-26.9kJ/molepH: 8.0 T: 2°CAssay Description:Quantitation of BPL catalysed 3H-biotin incorporation into the biotin domain substrate was performed as previously described by Polyak et al, J. Biol...More data for this Ligand-Target Pair
Affinity DataKi: >3.00E+4nM ΔG°: >-26.9kJ/molepH: 8.0 T: 2°CAssay Description:Quantitation of BPL catalysed 3H-biotin incorporation into the biotin domain substrate was performed as previously described by Polyak et al, J. Biol...More data for this Ligand-Target Pair
Affinity DataKi: >3.00E+4nM ΔG°: >-26.9kJ/molepH: 8.0 T: 2°CAssay Description:Quantitation of BPL catalysed 3H-biotin incorporation into the biotin domain substrate was performed as previously described by Polyak et al, J. Biol...More data for this Ligand-Target Pair
Affinity DataKi: >3.00E+4nM ΔG°: >-26.9kJ/molepH: 8.0 T: 2°CAssay Description:Quantitation of BPL catalysed 3H-biotin incorporation into the biotin domain substrate was performed as previously described by Polyak et al, J. Biol...More data for this Ligand-Target Pair
Affinity DataKi: >3.00E+4nM ΔG°: >-26.9kJ/molepH: 8.0 T: 2°CAssay Description:Quantitation of BPL catalysed 3H-biotin incorporation into the biotin domain substrate was performed as previously described by Polyak et al, J. Biol...More data for this Ligand-Target Pair
Affinity DataKi: >3.00E+4nM ΔG°: >-26.9kJ/molepH: 8.0 T: 2°CAssay Description:Quantitation of BPL catalysed 3H-biotin incorporation into the biotin domain substrate was performed as previously described by Polyak et al, J. Biol...More data for this Ligand-Target Pair
Affinity DataKi: >3.00E+4nM ΔG°: >-26.9kJ/molepH: 8.0 T: 2°CAssay Description:Quantitation of BPL catalysed 3H-biotin incorporation into the biotin domain substrate was performed as previously described by Polyak et al, J. Biol...More data for this Ligand-Target Pair


 3D Structure (crystal)
3D Structure (crystal)