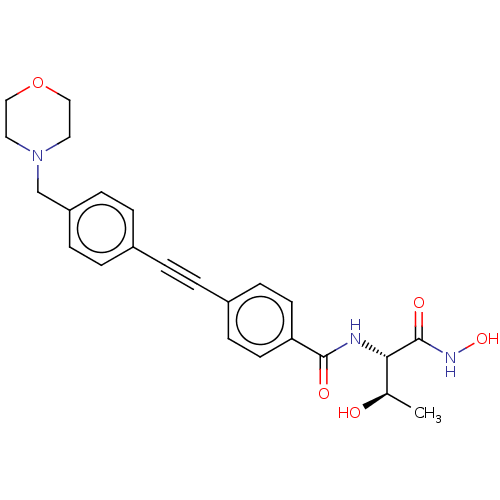
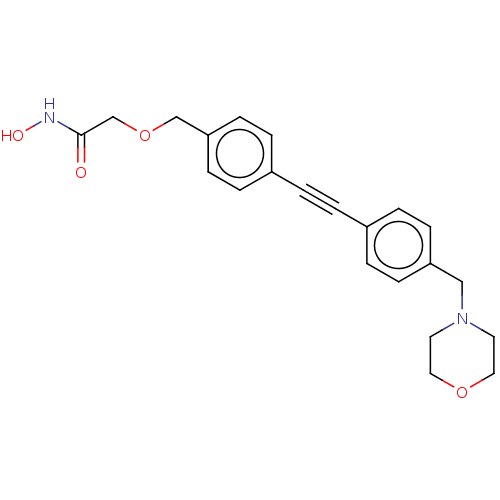
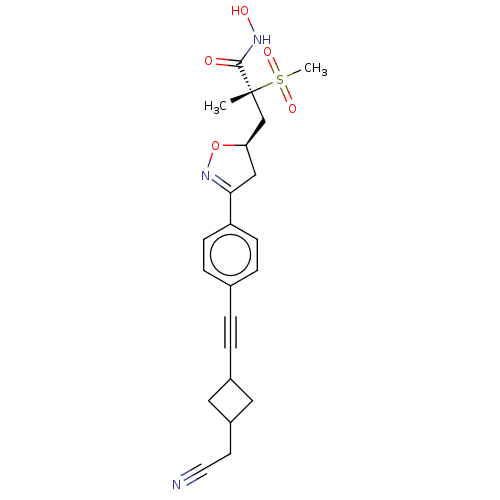
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli)
Duke University
Curated by ChEMBL
Duke University
Curated by ChEMBL
Affinity DataKi: 0.0240nMAssay Description:Inhibition of Escherichia coli LpxC enzyme using UDP-3-O-[(R)-3-hydroxymyristoyl]-N-acetyl glucosamine and [gamma-32P] UDP-3-O-[(R)-3-hydroxymyristoy...More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli)
Duke University
Curated by ChEMBL
Duke University
Curated by ChEMBL
Affinity DataKi: 0.0240nMAssay Description:Inhibition of Escherichia coli LpxC enzyme using UDP-3-O-[(R)-3-hydroxymyristoyl]-N-acetyl glucosamine and [gamma-32P] UDP-3-O-[(R)-3-hydroxymyristoy...More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli)
Duke University
Curated by ChEMBL
Duke University
Curated by ChEMBL
Affinity DataKi: 0.0530nMAssay Description:Inhibition of Escherichia coli LpxCMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli)
Duke University
Curated by ChEMBL
Duke University
Curated by ChEMBL
Affinity DataKi: 0.0690nMAssay Description:Inhibition of Escherichia coli LpxCMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli)
Duke University
Curated by ChEMBL
Duke University
Curated by ChEMBL
Affinity DataKi: 0.0860nMAssay Description:Inhibition of Escherichia coli LpxC Q202W/G210S mutantMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli)
Duke University
Curated by ChEMBL
Duke University
Curated by ChEMBL
Affinity DataKi: 0.200nMAssay Description:Inhibition of Escherichia coli LpxCMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli)
Duke University
Curated by ChEMBL
Duke University
Curated by ChEMBL
Affinity DataKi: 0.200nMAssay Description:Inhibition of Escherichia coli LpxCMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli)
Duke University
Curated by ChEMBL
Duke University
Curated by ChEMBL
Affinity DataKi: 0.340nMAssay Description:Inhibition of Escherichia coli LpxC Q202W/G210S mutantMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli)
Duke University
Curated by ChEMBL
Duke University
Curated by ChEMBL
Affinity DataKi: 0.5nMAssay Description:Inhibition of Escherichia coli LpxC enzymeMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli)
Duke University
Curated by ChEMBL
Duke University
Curated by ChEMBL
Affinity DataKi: 0.5nMAssay Description:Inhibition of Escherichia coli LpxCMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli)
Duke University
Curated by ChEMBL
Duke University
Curated by ChEMBL
Affinity DataKi: 0.550nMAssay Description:Inhibition of Escherichia coli LpxCMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli)
Duke University
Curated by ChEMBL
Duke University
Curated by ChEMBL
Affinity DataKi: 0.550nMAssay Description:Inhibition of Escherichia coli LpxCMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Rhizobium leguminosarum bv. trifolii (strain WSM13...)
Duke University Medical Center
Curated by ChEMBL
Duke University Medical Center
Curated by ChEMBL
Affinity DataKi: 0.700nMAssay Description:Inhibition of Rhizobium leguminosarum LpxC W206Q/S214G mutantMore data for this Ligand-Target Pair
Affinity DataKi: 1nM ΔG°: -51.4kJ/molepH: 7.0 T: 2°CAssay Description:OctetRedMore data for this Ligand-Target Pair
Affinity DataKi: 1nM ΔG°: -51.4kJ/molepH: 7.0 T: 2°CAssay Description:OctetRedMore data for this Ligand-Target Pair
Affinity DataKi: 1nM ΔG°: -51.4kJ/molepH: 7.0 T: 2°CAssay Description:OctetRedMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli)
Duke University
Curated by ChEMBL
Duke University
Curated by ChEMBL
Affinity DataKi: 2nMAssay Description:Inhibition of Escherichia coli LpxCMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli)
Duke University
Curated by ChEMBL
Duke University
Curated by ChEMBL
Affinity DataKi: 2nMAssay Description:Inhibition of Escherichia coli LpxCMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Rhizobium leguminosarum bv. trifolii (strain WSM13...)
Duke University Medical Center
Curated by ChEMBL
Duke University Medical Center
Curated by ChEMBL
Affinity DataKi: 3nMAssay Description:Inhibition of Rhizobium leguminosarum LpxC W206Q/S214G mutantMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Rhizobium leguminosarum bv. trifolii (strain WSM13...)
Duke University Medical Center
Curated by ChEMBL
Duke University Medical Center
Curated by ChEMBL
Affinity DataKi: 3.5nMAssay Description:Inhibition of Rhizobium leguminosarum LpxC S214G mutantMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli)
Duke University
Curated by ChEMBL
Duke University
Curated by ChEMBL
Affinity DataKi: 5nMAssay Description:Inhibition of Escherichia coli LpxC Q202W/G210S mutantMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli)
Duke University
Curated by ChEMBL
Duke University
Curated by ChEMBL
Affinity DataKi: 8nMAssay Description:Inhibition of Escherichia coli LpxC using UDP-3-O-[(R)-3-hydroxymyristoyl]-N-acetylglucosamine as substrate incubated for 30 mins prior to enzyme add...More data for this Ligand-Target Pair
Affinity DataKi: 10nM ΔG°: -45.7kJ/molepH: 7.0 T: 2°CAssay Description:OctetRedMore data for this Ligand-Target Pair
Affinity DataKi: 10nM ΔG°: -45.7kJ/molepH: 7.0 T: 2°CAssay Description:OctetRedMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli)
Duke University
Curated by ChEMBL
Duke University
Curated by ChEMBL
Affinity DataKi: 20nMAssay Description:Inhibition of Escherichia coli LpxCMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli)
Duke University
Curated by ChEMBL
Duke University
Curated by ChEMBL
Affinity DataKi: 50nMAssay Description:Inhibition of Escherichia coli LpxC enzymeMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli)
Duke University
Curated by ChEMBL
Duke University
Curated by ChEMBL
Affinity DataKi: 50nMAssay Description:Inhibition of Escherichia coli LpxCMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli)
Duke University
Curated by ChEMBL
Duke University
Curated by ChEMBL
Affinity DataKi: 50nMAssay Description:Inhibition of Escherichia coli LpxCMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli)
Duke University
Curated by ChEMBL
Duke University
Curated by ChEMBL
Affinity DataKi: 66nMAssay Description:Inhibition of Escherichia coli LpxC using UDP-3-O-[(R)-3-hydroxymyristoyl]-N-acetylglucosamine as substrate incubated for 30 mins prior to enzyme add...More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli)
Duke University
Curated by ChEMBL
Duke University
Curated by ChEMBL
Affinity DataKi: 95nMAssay Description:Inhibition of Escherichia coli LpxC using UDP-3-O-[(R)-3-hydroxymyristoyl]-N-acetylglucosamine as substrate incubated for 30 mins prior to enzyme add...More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Rhizobium leguminosarum bv. trifolii (strain WSM13...)
Duke University Medical Center
Curated by ChEMBL
Duke University Medical Center
Curated by ChEMBL
Affinity DataKi: 110nMAssay Description:Inhibition of Rhizobium leguminosarum LpxC W206Q/S214G mutantMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli)
Duke University
Curated by ChEMBL
Duke University
Curated by ChEMBL
Affinity DataKi: 358nMAssay Description:Inhibition of Escherichia coli LpxC using UDP-3-O-[(R)-3-hydroxymyristoyl]-N-acetylglucosamine as substrate incubated for 30 mins prior to enzyme add...More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli)
Duke University
Curated by ChEMBL
Duke University
Curated by ChEMBL
Affinity DataKi: 650nMAssay Description:Inhibition of Escherichia coli LpxCMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli)
Duke University
Curated by ChEMBL
Duke University
Curated by ChEMBL
Affinity DataKi: 650nMAssay Description:Inhibition of Escherichia coli LpxCMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli)
Duke University
Curated by ChEMBL
Duke University
Curated by ChEMBL
Affinity DataKi: 1.45E+3nMAssay Description:Inhibition of Escherichia coli LpxC using UDP-3-O-[(R)-3-hydroxymyristoyl]-N-acetylglucosamine as substrate incubated for 30 mins prior to enzyme add...More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli)
Duke University
Curated by ChEMBL
Duke University
Curated by ChEMBL
Affinity DataKi: 4.40E+3nMAssay Description:Inhibition of Escherichia coli LpxC using UDP-3-O-[(R)-3-hydroxymyristoyl]-N-acetylglucosamine as substrate incubated for 30 mins prior to enzyme add...More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli)
Duke University
Curated by ChEMBL
Duke University
Curated by ChEMBL
Affinity DataKi: 2.73E+4nMAssay Description:Inhibition of Escherichia coli LpxC using UDP-3-O-[(R)-3-hydroxymyristoyl]-N-acetylglucosamine as substrate incubated for 30 mins prior to enzyme add...More data for this Ligand-Target Pair
Affinity DataIC50: <0.000100nMpH: 7.5 T: 2°CAssay Description:The P. aeruginosa LpxC protein is produced according to the general method of Hyland et al (Journal of Bacteriology 1997 179, 2029-2037: Cloning, exp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.00100nMpH: 7.5 T: 2°CAssay Description:The P. aeruginosa LpxC protein is produced according to the general method of Hyland et al (Journal of Bacteriology 1997 179, 2029-2037: Cloning, exp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.00100nMpH: 7.5 T: 2°CAssay Description:The P. aeruginosa LpxC protein is produced according to the general method of Hyland et al (Journal of Bacteriology 1997 179, 2029-2037: Cloning, exp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.00100nMpH: 7.5 T: 2°CAssay Description:The P. aeruginosa LpxC protein is produced according to the general method of Hyland et al (Journal of Bacteriology 1997 179, 2029-2037: Cloning, exp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.00100nMpH: 7.5 T: 2°CAssay Description:The P. aeruginosa LpxC protein is produced according to the general method of Hyland et al (Journal of Bacteriology 1997 179, 2029-2037: Cloning, exp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.00100nMpH: 7.5 T: 2°CAssay Description:The P. aeruginosa LpxC protein is produced according to the general method of Hyland et al (Journal of Bacteriology 1997 179, 2029-2037: Cloning, exp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.00100nMpH: 7.5 T: 2°CAssay Description:The P. aeruginosa LpxC protein is produced according to the general method of Hyland et al (Journal of Bacteriology 1997 179, 2029-2037: Cloning, exp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.00100nMpH: 7.5 T: 2°CAssay Description:The P. aeruginosa LpxC protein is produced according to the general method of Hyland et al (Journal of Bacteriology 1997 179, 2029-2037: Cloning, exp...More data for this Ligand-Target Pair
Affinity DataIC50: <0.00100nMpH: 7.5 T: 2°CAssay Description:The P. aeruginosa LpxC protein is produced according to the general method of Hyland et al (Journal of Bacteriology 1997 179, 2029-2037: Cloning, exp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.00100nMpH: 7.5 T: 2°CAssay Description:The P. aeruginosa LpxC protein is produced according to the general method of Hyland et al (Journal of Bacteriology 1997 179, 2029-2037: Cloning, exp...More data for this Ligand-Target Pair
Affinity DataIC50: <0.00100nMpH: 7.5 T: 2°CAssay Description:The P. aeruginosa LpxC protein is produced according to the general method of Hyland et al (Journal of Bacteriology 1997 179, 2029-2037: Cloning, exp...More data for this Ligand-Target Pair
Affinity DataIC50: <0.00100nMpH: 7.5 T: 2°CAssay Description:The P. aeruginosa LpxC protein is produced according to the general method of Hyland et al (Journal of Bacteriology 1997 179, 2029-2037: Cloning, exp...More data for this Ligand-Target Pair
Affinity DataIC50: <0.00100nMpH: 7.5 T: 2°CAssay Description:The P. aeruginosa LpxC protein is produced according to the general method of Hyland et al (Journal of Bacteriology 1997 179, 2029-2037: Cloning, exp...More data for this Ligand-Target Pair




 3D Structure (crystal)
3D Structure (crystal)