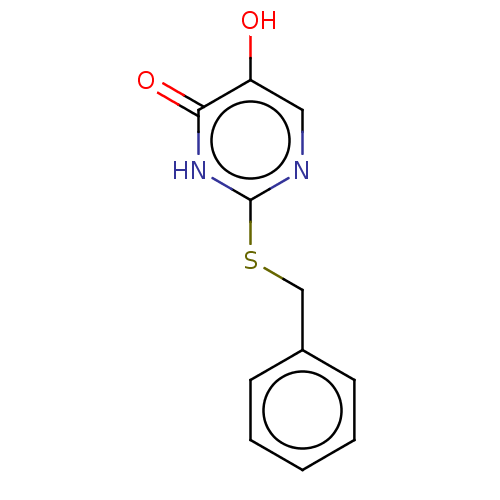
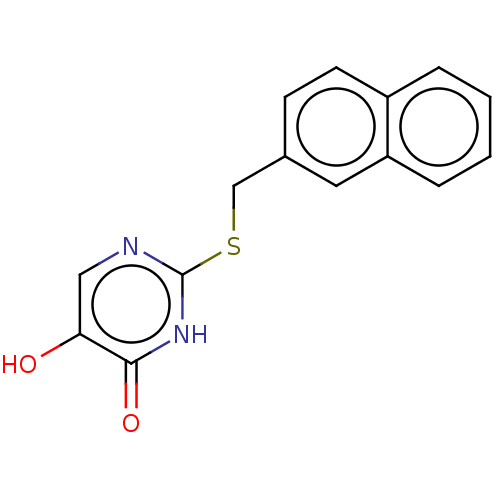
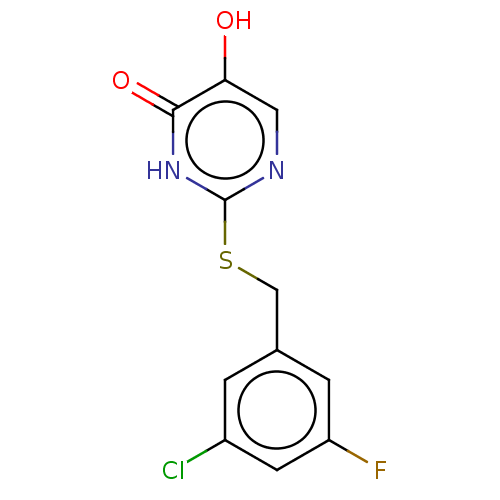
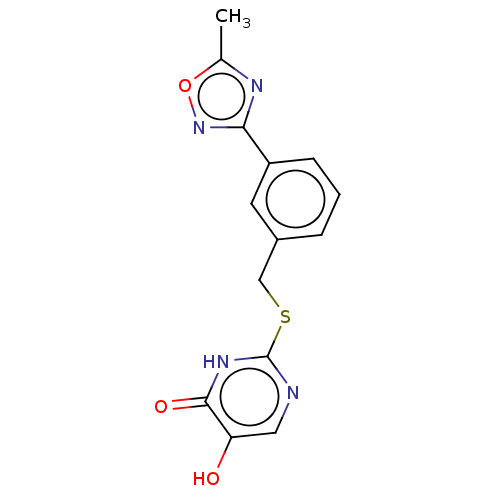
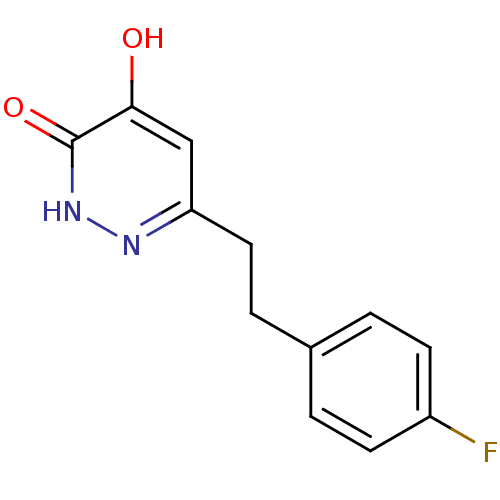
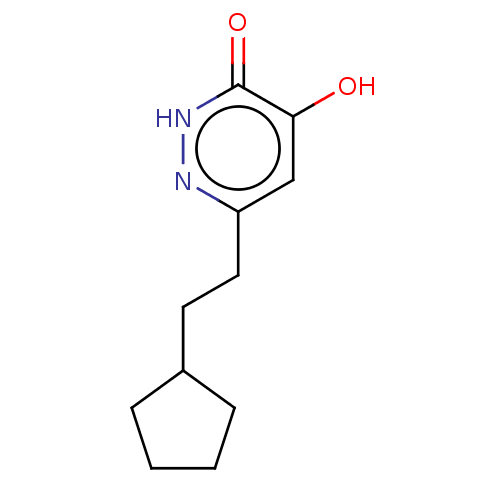
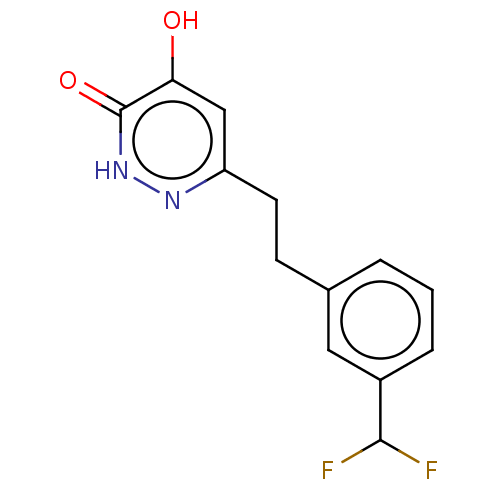
Affinity DataIC50: 13nMpH: 7.4 T: 2°CAssay Description:Human DAAO enzyme was supplied by the Takeda Pharmaceutical Company (Osaka) and each batch was tested and used at concentrations giving comparable le...More data for this Ligand-Target Pair
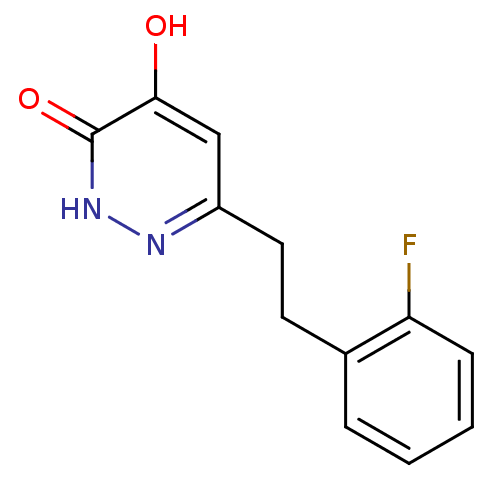
Affinity DataIC50: 12nMpH: 7.4 T: 2°CAssay Description:Human DAAO enzyme was supplied by the Takeda Pharmaceutical Company (Osaka) and each batch was tested and used at concentrations giving comparable le...More data for this Ligand-Target Pair
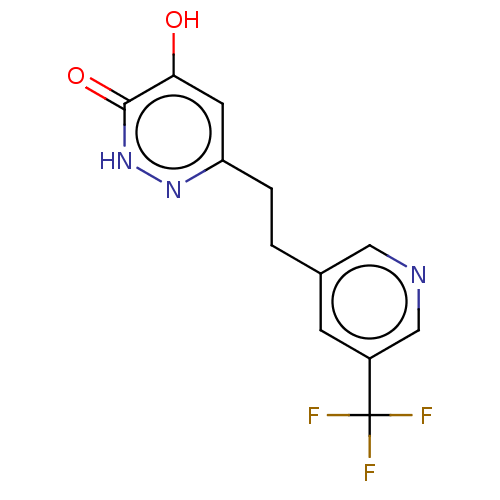
Affinity DataIC50: 10nMpH: 7.4 T: 2°CAssay Description:Human DAAO enzyme was supplied by the Takeda Pharmaceutical Company (Osaka) and each batch was tested and used at concentrations giving comparable le...More data for this Ligand-Target Pair
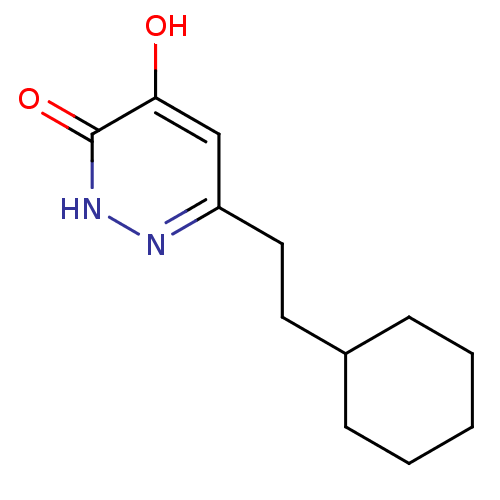
Affinity DataIC50: 19nMpH: 7.4 T: 2°CAssay Description:Human DAAO enzyme was supplied by the Takeda Pharmaceutical Company (Osaka) and each batch was tested and used at concentrations giving comparable le...More data for this Ligand-Target Pair
Affinity DataIC50: 120nMpH: 7.4Assay Description:The functional activity of compounds inhibiting the DAAO enzyme was determined by utilizing the co-product of the catalysis of D-Serine, H2O2 which c...More data for this Ligand-Target Pair
Affinity DataIC50: 230nMpH: 7.4Assay Description:The functional activity of compounds inhibiting the DAAO enzyme was determined by utilizing the co-product of the catalysis of D-Serine, H2O2 which c...More data for this Ligand-Target Pair
Affinity DataIC50: 220nMpH: 7.4Assay Description:The functional activity of compounds inhibiting the DAAO enzyme was determined by utilizing the co-product of the catalysis of D-Serine, H2O2 which c...More data for this Ligand-Target Pair
Affinity DataIC50: 120nMpH: 7.4Assay Description:The functional activity of compounds inhibiting the DAAO enzyme was determined by utilizing the co-product of the catalysis of D-Serine, H2O2 which c...More data for this Ligand-Target Pair
Affinity DataIC50: 170nMpH: 7.4Assay Description:The functional activity of compounds inhibiting the DAAO enzyme was determined by utilizing the co-product of the catalysis of D-Serine, H2O2 which c...More data for this Ligand-Target Pair
Affinity DataIC50: 160nMpH: 7.4Assay Description:The functional activity of compounds inhibiting the DAAO enzyme was determined by utilizing the co-product of the catalysis of D-Serine, H2O2 which c...More data for this Ligand-Target Pair
Affinity DataIC50: 150nMpH: 7.4Assay Description:The functional activity of compounds inhibiting the DAAO enzyme was determined by utilizing the co-product of the catalysis of D-Serine, H2O2 which c...More data for this Ligand-Target Pair
Affinity DataIC50: 150nMpH: 7.4Assay Description:The functional activity of compounds inhibiting the DAAO enzyme was determined by utilizing the co-product of the catalysis of D-Serine, H2O2 which c...More data for this Ligand-Target Pair
Affinity DataIC50: 500nMpH: 7.4Assay Description:The functional activity of compounds inhibiting the DAAO enzyme was determined by utilizing the co-product of the catalysis of D-Serine, H2O2 which c...More data for this Ligand-Target Pair
Affinity DataIC50: 1.20E+3nMpH: 7.4Assay Description:The functional activity of compounds inhibiting the DAAO enzyme was determined by utilizing the co-product of the catalysis of D-Serine, H2O2 which c...More data for this Ligand-Target Pair
Affinity DataIC50: 540nMpH: 7.4Assay Description:The functional activity of compounds inhibiting the DAAO enzyme was determined by utilizing the co-product of the catalysis of D-Serine, H2O2 which c...More data for this Ligand-Target Pair
Affinity DataIC50: 190nMpH: 7.4Assay Description:The functional activity of compounds inhibiting the DAAO enzyme was determined by utilizing the co-product of the catalysis of D-Serine, H2O2 which c...More data for this Ligand-Target Pair
Affinity DataIC50: 470nMpH: 7.4Assay Description:The functional activity of compounds inhibiting the DAAO enzyme was determined by utilizing the co-product of the catalysis of D-Serine, H2O2 which c...More data for this Ligand-Target Pair
Affinity DataIC50: 1.60E+3nMpH: 7.4Assay Description:The functional activity of compounds inhibiting the DAAO enzyme was determined by utilizing the co-product of the catalysis of D-Serine, H2O2 which c...More data for this Ligand-Target Pair
Affinity DataIC50: 740nMpH: 7.4Assay Description:The functional activity of compounds inhibiting the DAAO enzyme was determined by utilizing the co-product of the catalysis of D-Serine, H2O2 which c...More data for this Ligand-Target Pair
Affinity DataIC50: 2.80E+3nMpH: 7.4Assay Description:The functional activity of compounds inhibiting the DAAO enzyme was determined by utilizing the co-product of the catalysis of D-Serine, H2O2 which c...More data for this Ligand-Target Pair
Affinity DataIC50: 3.20E+3nMpH: 7.4Assay Description:The functional activity of compounds inhibiting the DAAO enzyme was determined by utilizing the co-product of the catalysis of D-Serine, H2O2 which c...More data for this Ligand-Target Pair
Affinity DataIC50: 1.20E+3nMpH: 7.4Assay Description:The functional activity of compounds inhibiting the DAAO enzyme was determined by utilizing the co-product of the catalysis of D-Serine, H2O2 which c...More data for this Ligand-Target Pair
Affinity DataIC50: 2.70E+3nMpH: 7.4Assay Description:The functional activity of compounds inhibiting the DAAO enzyme was determined by utilizing the co-product of the catalysis of D-Serine, H2O2 which c...More data for this Ligand-Target Pair
Affinity DataIC50: 1.80E+3nMpH: 7.4Assay Description:The functional activity of compounds inhibiting the DAAO enzyme was determined by utilizing the co-product of the catalysis of D-Serine, H2O2 which c...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMpH: 7.4 T: 2°CAssay Description:Human DAAO enzyme was supplied by the Takeda Pharmaceutical Company (Osaka) and each batch was tested and used at concentrations giving comparable le...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMpH: 7.4 T: 2°CAssay Description:Human DAAO enzyme was supplied by the Takeda Pharmaceutical Company (Osaka) and each batch was tested and used at concentrations giving comparable le...More data for this Ligand-Target Pair
Affinity DataIC50: 21nMpH: 7.4 T: 2°CAssay Description:Human DAAO enzyme was supplied by the Takeda Pharmaceutical Company (Osaka) and each batch was tested and used at concentrations giving comparable le...More data for this Ligand-Target Pair
Affinity DataIC50: 3.70nMpH: 7.4 T: 2°CAssay Description:Human DAAO enzyme was supplied by the Takeda Pharmaceutical Company (Osaka) and each batch was tested and used at concentrations giving comparable le...More data for this Ligand-Target Pair
Affinity DataIC50: 30nMpH: 7.4 T: 2°CAssay Description:Human DAAO enzyme was supplied by the Takeda Pharmaceutical Company (Osaka) and each batch was tested and used at concentrations giving comparable le...More data for this Ligand-Target Pair
Affinity DataIC50: 9.70nMpH: 7.4 T: 2°CAssay Description:Human DAAO enzyme was supplied by the Takeda Pharmaceutical Company (Osaka) and each batch was tested and used at concentrations giving comparable le...More data for this Ligand-Target Pair
Affinity DataIC50: 13nMpH: 7.4 T: 2°CAssay Description:Human DAAO enzyme was supplied by the Takeda Pharmaceutical Company (Osaka) and each batch was tested and used at concentrations giving comparable le...More data for this Ligand-Target Pair
Affinity DataIC50: 11nMpH: 7.4 T: 2°CAssay Description:Human DAAO enzyme was supplied by the Takeda Pharmaceutical Company (Osaka) and each batch was tested and used at concentrations giving comparable le...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMpH: 7.4 T: 2°CAssay Description:Human DAAO enzyme was supplied by the Takeda Pharmaceutical Company (Osaka) and each batch was tested and used at concentrations giving comparable le...More data for this Ligand-Target Pair
Affinity DataIC50: 22nMpH: 7.4 T: 2°CAssay Description:Human DAAO enzyme was supplied by the Takeda Pharmaceutical Company (Osaka) and each batch was tested and used at concentrations giving comparable le...More data for this Ligand-Target Pair
Affinity DataIC50: 16nMpH: 7.4 T: 2°CAssay Description:Human DAAO enzyme was supplied by the Takeda Pharmaceutical Company (Osaka) and each batch was tested and used at concentrations giving comparable le...More data for this Ligand-Target Pair
Affinity DataIC50: 23nMpH: 7.4 T: 2°CAssay Description:Human DAAO enzyme was supplied by the Takeda Pharmaceutical Company (Osaka) and each batch was tested and used at concentrations giving comparable le...More data for this Ligand-Target Pair
Affinity DataIC50: 31nMpH: 7.4 T: 2°CAssay Description:Human DAAO enzyme was supplied by the Takeda Pharmaceutical Company (Osaka) and each batch was tested and used at concentrations giving comparable le...More data for this Ligand-Target Pair
Affinity DataIC50: 41nMpH: 7.4 T: 2°CAssay Description:Human DAAO enzyme was supplied by the Takeda Pharmaceutical Company (Osaka) and each batch was tested and used at concentrations giving comparable le...More data for this Ligand-Target Pair
Affinity DataIC50: 16nMpH: 7.4 T: 2°CAssay Description:Human DAAO enzyme was supplied by the Takeda Pharmaceutical Company (Osaka) and each batch was tested and used at concentrations giving comparable le...More data for this Ligand-Target Pair
Affinity DataIC50: 52nMpH: 7.4 T: 2°CAssay Description:Human DAAO enzyme was supplied by the Takeda Pharmaceutical Company (Osaka) and each batch was tested and used at concentrations giving comparable le...More data for this Ligand-Target Pair
Affinity DataIC50: 13nMpH: 7.4 T: 2°CAssay Description:Human DAAO enzyme was supplied by the Takeda Pharmaceutical Company (Osaka) and each batch was tested and used at concentrations giving comparable le...More data for this Ligand-Target Pair
Affinity DataIC50: 14nMpH: 7.4 T: 2°CAssay Description:Human DAAO enzyme was supplied by the Takeda Pharmaceutical Company (Osaka) and each batch was tested and used at concentrations giving comparable le...More data for this Ligand-Target Pair
Affinity DataIC50: 12nMpH: 7.4 T: 2°CAssay Description:Human DAAO enzyme was supplied by the Takeda Pharmaceutical Company (Osaka) and each batch was tested and used at concentrations giving comparable le...More data for this Ligand-Target Pair
Affinity DataIC50: 8.40nMpH: 7.4 T: 2°CAssay Description:Human DAAO enzyme was supplied by the Takeda Pharmaceutical Company (Osaka) and each batch was tested and used at concentrations giving comparable le...More data for this Ligand-Target Pair
Affinity DataIC50: 21nMpH: 7.4 T: 2°CAssay Description:Human DAAO enzyme was supplied by the Takeda Pharmaceutical Company (Osaka) and each batch was tested and used at concentrations giving comparable le...More data for this Ligand-Target Pair
Affinity DataIC50: 13nMpH: 7.4 T: 2°CAssay Description:Human DAAO enzyme was supplied by the Takeda Pharmaceutical Company (Osaka) and each batch was tested and used at concentrations giving comparable le...More data for this Ligand-Target Pair
Affinity DataIC50: 14nMpH: 7.4 T: 2°CAssay Description:Human DAAO enzyme was supplied by the Takeda Pharmaceutical Company (Osaka) and each batch was tested and used at concentrations giving comparable le...More data for this Ligand-Target Pair
Affinity DataIC50: 6nMpH: 7.4 T: 2°CAssay Description:Human DAAO enzyme was supplied by the Takeda Pharmaceutical Company (Osaka) and each batch was tested and used at concentrations giving comparable le...More data for this Ligand-Target Pair
Affinity DataIC50: 45nMpH: 7.4 T: 2°CAssay Description:Human DAAO enzyme was supplied by the Takeda Pharmaceutical Company (Osaka) and each batch was tested and used at concentrations giving comparable le...More data for this Ligand-Target Pair
Affinity DataIC50: 22nMpH: 7.4 T: 2°CAssay Description:Human DAAO enzyme was supplied by the Takeda Pharmaceutical Company (Osaka) and each batch was tested and used at concentrations giving comparable le...More data for this Ligand-Target Pair