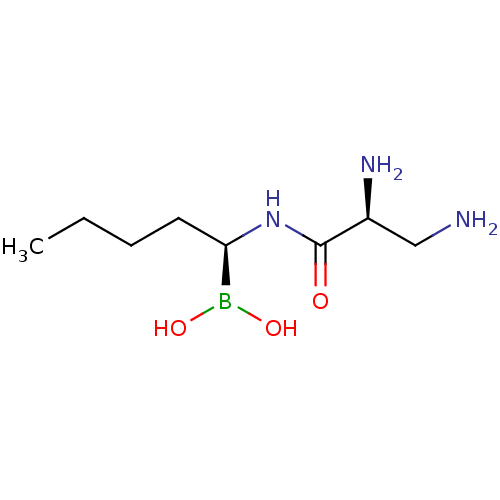
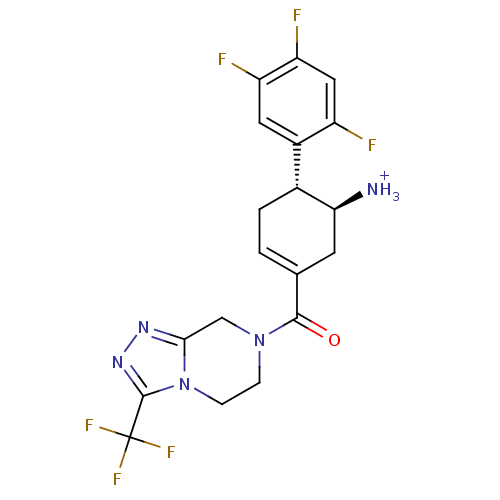
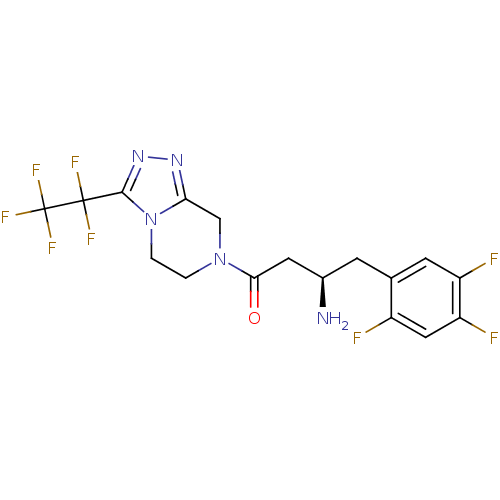
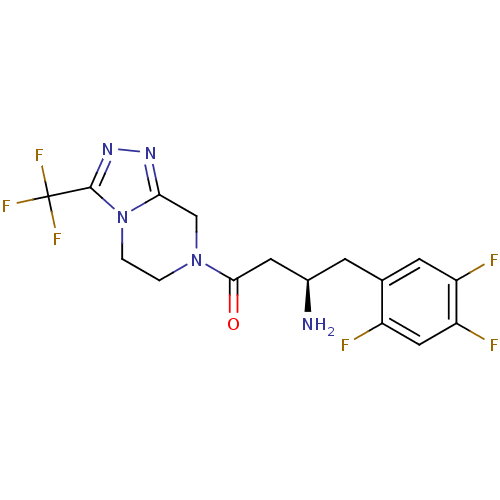
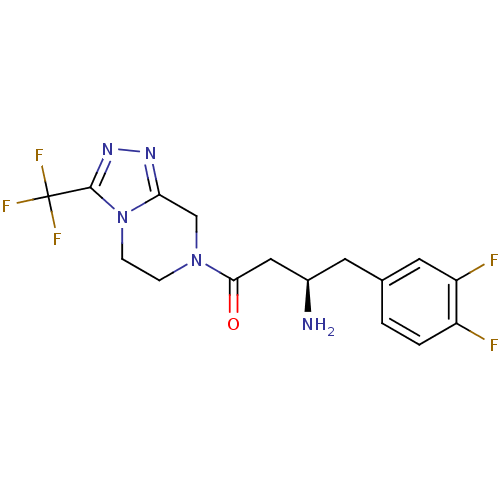
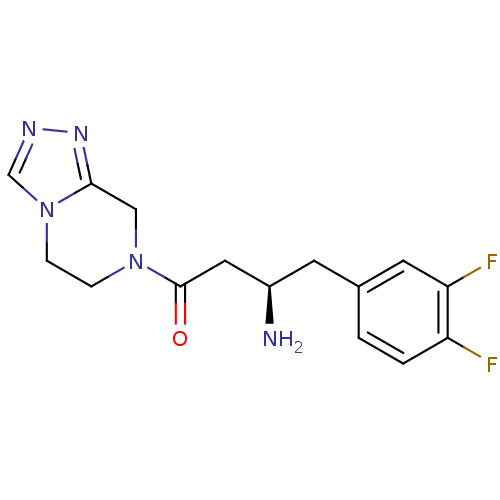
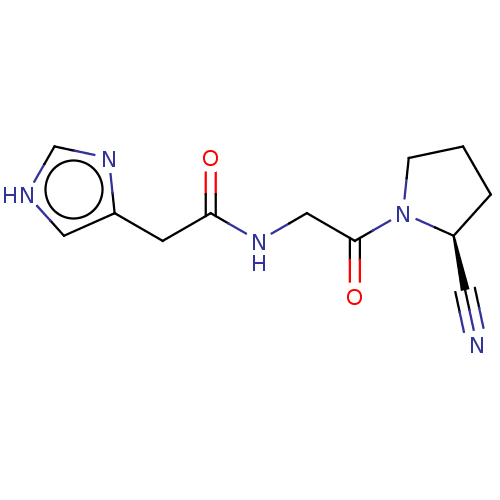
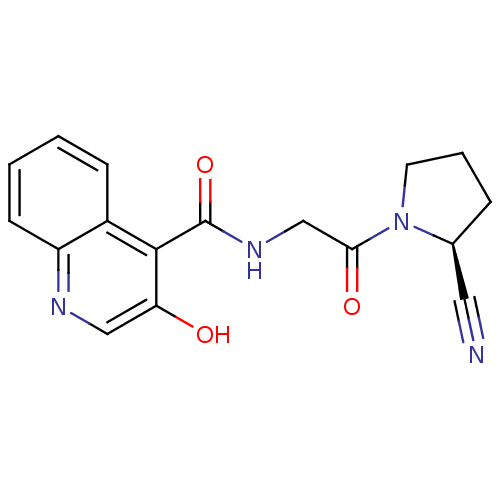
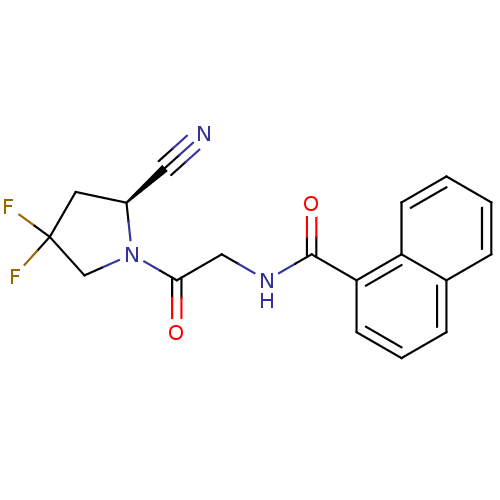
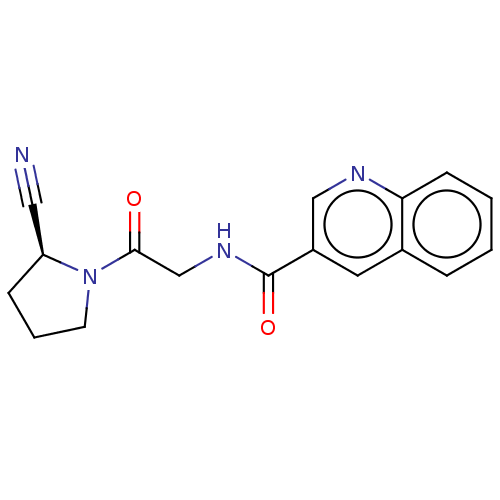
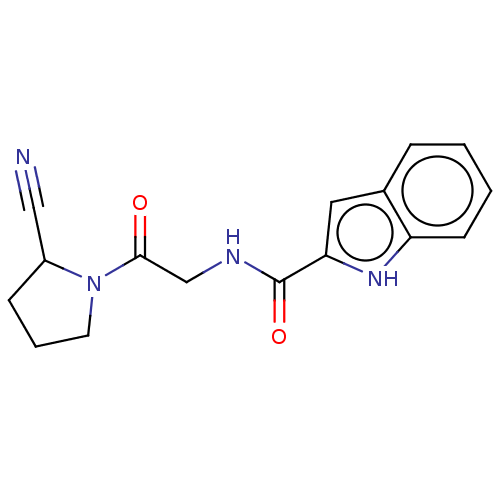
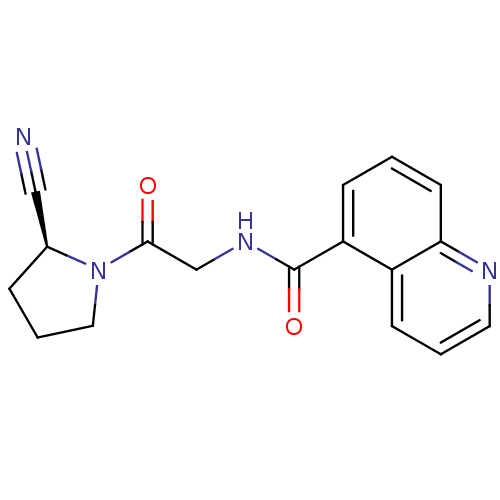
Affinity DataIC50: 4.30E+4nMpH: 2.5Assay Description:Inhibitory concentration oagainst DPP9; Compound dissolved in DMSO solution at a pH of 2.5More data for this Ligand-Target Pair
Affinity DataKi: 73nM ΔG°: -40.3kJ/molepH: 7.5 T: 2°CAssay Description:The DPP activity resulted in the formation of the fluorescent product amidomethylcoumarin (AMC), which was monitored by excitation at 355 nm and meas...More data for this Ligand-Target Pair
Affinity DataKi: 430nM ΔG°: -36.0kJ/molepH: 7.5 T: 2°CAssay Description:The DPP activity resulted in the formation of the fluorescent product amidomethylcoumarin (AMC), which was monitored by excitation at 355 nm and meas...More data for this Ligand-Target Pair
Affinity DataKi: 500nM ΔG°: -35.6kJ/molepH: 7.5 T: 2°CAssay Description:The DPP activity resulted in the formation of the fluorescent product amidomethylcoumarin (AMC), which was monitored by excitation at 355 nm and meas...More data for this Ligand-Target Pair
Affinity DataKi: >3.00E+3nM ΔG°: >-31.2kJ/molepH: 7.5 T: 2°CAssay Description:The DPP activity resulted in the formation of the fluorescent product amidomethylcoumarin (AMC), which was monitored by excitation at 355 nm and meas...More data for this Ligand-Target Pair
Affinity DataKi: >3.00E+3nM ΔG°: >-31.2kJ/molepH: 7.5 T: 2°CAssay Description:The DPP activity resulted in the formation of the fluorescent product amidomethylcoumarin (AMC), which was monitored by excitation at 355 nm and meas...More data for this Ligand-Target Pair
Affinity DataKi: 4.00E+3nM ΔG°: -30.5kJ/molepH: 7.5 T: 2°CAssay Description:The DPP activity resulted in the formation of the fluorescent product amidomethylcoumarin (AMC), which was monitored by excitation at 355 nm and meas...More data for this Ligand-Target Pair
Affinity DataKi: 1.40E+4nM ΔG°: -27.4kJ/molepH: 7.5 T: 2°CAssay Description:The DPP activity resulted in the formation of the fluorescent product amidomethylcoumarin (AMC), which was monitored by excitation at 355 nm and meas...More data for this Ligand-Target Pair
Affinity DataIC50: >4.00E+5nMpH: 7.5 T: 2°CAssay Description:DPP inhibitory activity using fluoresence assay. Spectra MAX Gemini XS fluorescence spectrometer was used as the fluorescence spectrometer and the e...More data for this Ligand-Target Pair
Affinity DataIC50: >4.00E+5nMpH: 7.5 T: 2°CAssay Description:DPP inhibitory activity using fluoresence assay. Spectra MAX Gemini XS fluorescence spectrometer was used as the fluorescence spectrometer and the e...More data for this Ligand-Target Pair
Affinity DataIC50: >1.00E+5nMpH: 7.5 T: 2°CAssay Description:DPP inhibitory activity using fluoresence assay. Spectra MAX Gemini XS fluorescence spectrometer was used as the fluorescence spectrometer and the e...More data for this Ligand-Target Pair
Affinity DataIC50: >1.30E+5nMpH: 7.5 T: 2°CAssay Description:DPP inhibitory activity using fluoresence assay. Spectra MAX Gemini XS fluorescence spectrometer was used as the fluorescence spectrometer and the e...More data for this Ligand-Target Pair
Affinity DataIC50: >2.10E+5nMpH: 7.5 T: 2°CAssay Description:DPP inhibitory activity using fluoresence assay. Spectra MAX Gemini XS fluorescence spectrometer was used as the fluorescence spectrometer and the e...More data for this Ligand-Target Pair
Affinity DataIC50: >1.00E+5nMpH: 7.5 T: 2°CAssay Description:The enzyme activity resulted in the formation of the fluorescent product amidomethylcoumarin (AMC), which was monitored by excitation at 360 nm and m...More data for this Ligand-Target Pair
Affinity DataIC50: >1.00E+5nMpH: 7.5 T: 2°CAssay Description:The enzyme activity resulted in the formation of the fluorescent product amidomethylcoumarin (AMC), which was monitored by excitation at 360 nm and m...More data for this Ligand-Target Pair
Affinity DataIC50: >1.00E+5nMpH: 7.5 T: 2°CAssay Description:The enzyme activity resulted in the formation of the fluorescent product amidomethylcoumarin (AMC), which was monitored by excitation at 360 nm and m...More data for this Ligand-Target Pair
Affinity DataIC50: >1.00E+5nMpH: 7.5 T: 2°CAssay Description:The enzyme activity resulted in the formation of the fluorescent product amidomethylcoumarin (AMC), which was monitored by excitation at 360 nm and m...More data for this Ligand-Target Pair
Affinity DataIC50: >1.00E+5nMpH: 7.5 T: 2°CAssay Description:The enzyme activity resulted in the formation of the fluorescent product amidomethylcoumarin (AMC), which was monitored by excitation at 360 nm and m...More data for this Ligand-Target Pair
Affinity DataIC50: >1.00E+5nMpH: 7.5 T: 2°CAssay Description:The enzyme activity resulted in the formation of the fluorescent product amidomethylcoumarin (AMC), which was monitored by excitation at 360 nm and m...More data for this Ligand-Target Pair
Affinity DataIC50: >1.00E+5nMpH: 7.5 T: 2°CAssay Description:The enzyme activity resulted in the formation of the fluorescent product amidomethylcoumarin (AMC), which was monitored by excitation at 360 nm and m...More data for this Ligand-Target Pair
Affinity DataIC50: >1.10E+5nMpH: 7.5 T: 2°CAssay Description:DPP inhibitory activity using fluoresence assay. Spectra MAX Gemini XS fluorescence spectrometer was used as the fluorescence spectrometer and the e...More data for this Ligand-Target Pair
Affinity DataIC50: >1.00E+5nMpH: 7.5 T: 2°CAssay Description:The enzyme activity resulted in the formation of the fluorescent product amidomethylcoumarin (AMC), which was monitored by excitation at 360 nm and m...More data for this Ligand-Target Pair
Affinity DataIC50: >1.00E+5nMpH: 8.3 T: 2°CAssay Description:Enzyme activities were determined kinetically in a final volume of 200 μl for 10 minutes at 37° C. by measuring the initial velocities of pNA ...More data for this Ligand-Target Pair
Affinity DataIC50: >1.00E+5nMpH: 8.3 T: 2°CAssay Description:Enzyme activities were determined kinetically in a final volume of 200 μl for 10 minutes at 37° C. by measuring the initial velocities of pNA ...More data for this Ligand-Target Pair
Affinity DataIC50: >1.00E+5nMpH: 8.3 T: 2°CAssay Description:Enzyme activities were determined kinetically in a final volume of 200 μl for 10 minutes at 37° C. by measuring the initial velocities of pNA ...More data for this Ligand-Target Pair
Affinity DataIC50: >5.00E+4nMpH: 8.3 T: 2°CAssay Description:Enzyme activities were determined kinetically in a final volume of 200 μl for 10 minutes at 37° C. by measuring the initial velocities of pNA ...More data for this Ligand-Target Pair
Affinity DataIC50: >1.00E+5nMpH: 8.3 T: 2°CAssay Description:Enzyme activities were determined kinetically in a final volume of 200 μl for 10 minutes at 37° C. by measuring the initial velocities of pNA ...More data for this Ligand-Target Pair
Affinity DataIC50: >1.00E+5nMpH: 8.3 T: 2°CAssay Description:Enzyme activities were determined kinetically in a final volume of 200 μl for 10 minutes at 37° C. by measuring the initial velocities of pNA ...More data for this Ligand-Target Pair
Affinity DataIC50: >1.00E+5nMpH: 8.3 T: 2°CAssay Description:Enzyme activities were determined kinetically in a final volume of 200 μl for 10 minutes at 37° C. by measuring the initial velocities of pNA ...More data for this Ligand-Target Pair
Affinity DataIC50: >1.00E+5nMpH: 8.3 T: 2°CAssay Description:Enzyme activities were determined kinetically in a final volume of 200 μl for 10 minutes at 37° C. by measuring the initial velocities of pNA ...More data for this Ligand-Target Pair
Affinity DataIC50: >1.00E+5nMpH: 8.3 T: 2°CAssay Description:Enzyme activities were determined kinetically in a final volume of 200 μl for 10 minutes at 37° C. by measuring the initial velocities of pNA ...More data for this Ligand-Target Pair
Affinity DataIC50: >1.00E+5nMpH: 8.3 T: 2°CAssay Description:Enzyme activities were determined kinetically in a final volume of 200 μl for 10 minutes at 37° C. by measuring the initial velocities of pNA ...More data for this Ligand-Target Pair
Affinity DataIC50: >1.00E+5nMpH: 8.3 T: 2°CAssay Description:Enzyme activities were determined kinetically in a final volume of 200 μl for 10 minutes at 37° C. by measuring the initial velocities of pNA ...More data for this Ligand-Target Pair
Affinity DataIC50: >2.50E+4nMpH: 8.3 T: 2°CAssay Description:Enzyme activities were determined kinetically in a final volume of 200 μl for 10 minutes at 37° C. by measuring the initial velocities of pNA ...More data for this Ligand-Target Pair
Affinity DataIC50: >1.00E+5nMpH: 8.3 T: 2°CAssay Description:Enzyme activities were determined kinetically in a final volume of 200 μl for 10 minutes at 37° C. by measuring the initial velocities of pNA ...More data for this Ligand-Target Pair
Affinity DataIC50: >1.00E+5nMpH: 8.3 T: 2°CAssay Description:Enzyme activities were determined kinetically in a final volume of 200 μl for 10 minutes at 37° C. by measuring the initial velocities of pNA ...More data for this Ligand-Target Pair
Affinity DataIC50: >1.00E+5nMpH: 8.3 T: 2°CAssay Description:Enzyme activities were determined kinetically in a final volume of 200 μl for 10 minutes at 37° C. by measuring the initial velocities of pNA ...More data for this Ligand-Target Pair
Affinity DataIC50: >1.00E+5nMpH: 8.3 T: 2°CAssay Description:Enzyme activities were determined kinetically in a final volume of 200 μl for 10 minutes at 37° C. by measuring the initial velocities of pNA ...More data for this Ligand-Target Pair
Affinity DataIC50: >1.00E+5nMpH: 8.3 T: 2°CAssay Description:Enzyme activities were determined kinetically in a final volume of 200 μl for 10 minutes at 37° C. by measuring the initial velocities of pNA ...More data for this Ligand-Target Pair
Affinity DataIC50: >1.00E+5nMpH: 8.3 T: 2°CAssay Description:Enzyme activities were determined kinetically in a final volume of 200 μl for 10 minutes at 37° C. by measuring the initial velocities of pNA ...More data for this Ligand-Target Pair
Affinity DataIC50: >1.25E+4nMpH: 8.3 T: 2°CAssay Description:Enzyme activities were determined kinetically in a final volume of 200 μl for 10 minutes at 37° C. by measuring the initial velocities of pNA ...More data for this Ligand-Target Pair
Affinity DataIC50: >1.00E+5nMpH: 8.3 T: 2°CAssay Description:Enzyme activities were determined kinetically in a final volume of 200 μl for 10 minutes at 37° C. by measuring the initial velocities of pNA ...More data for this Ligand-Target Pair
Affinity DataIC50: >1.00E+5nMpH: 8.3 T: 2°CAssay Description:Enzyme activities were determined kinetically in a final volume of 200 μl for 10 minutes at 37° C. by measuring the initial velocities of pNA ...More data for this Ligand-Target Pair
Affinity DataIC50: >1.00E+5nMpH: 8.3 T: 2°CAssay Description:Enzyme activities were determined kinetically in a final volume of 200 μl for 10 minutes at 37° C. by measuring the initial velocities of pNA ...More data for this Ligand-Target Pair
Affinity DataIC50: >1.00E+5nMpH: 8.3 T: 2°CAssay Description:Enzyme activities were determined kinetically in a final volume of 200 μl for 10 minutes at 37° C. by measuring the initial velocities of pNA ...More data for this Ligand-Target Pair
Affinity DataIC50: >1.00E+5nMpH: 8.3 T: 2°CAssay Description:Enzyme activities were determined kinetically in a final volume of 200 μl for 10 minutes at 37° C. by measuring the initial velocities of pNA ...More data for this Ligand-Target Pair
Affinity DataIC50: >1.00E+5nMpH: 8.3 T: 2°CAssay Description:Enzyme activities were determined kinetically in a final volume of 200 μl for 10 minutes at 37° C. by measuring the initial velocities of pNA ...More data for this Ligand-Target Pair
Affinity DataIC50: >1.00E+5nMpH: 8.3 T: 2°CAssay Description:Enzyme activities were determined kinetically in a final volume of 200 μl for 10 minutes at 37° C. by measuring the initial velocities of pNA ...More data for this Ligand-Target Pair
Affinity DataIC50: >1.00E+5nMpH: 8.3 T: 2°CAssay Description:Enzyme activities were determined kinetically in a final volume of 200 μl for 10 minutes at 37° C. by measuring the initial velocities of pNA ...More data for this Ligand-Target Pair
Affinity DataIC50: >1.00E+5nMpH: 8.3 T: 2°CAssay Description:Enzyme activities were determined kinetically in a final volume of 200 μl for 10 minutes at 37° C. by measuring the initial velocities of pNA ...More data for this Ligand-Target Pair