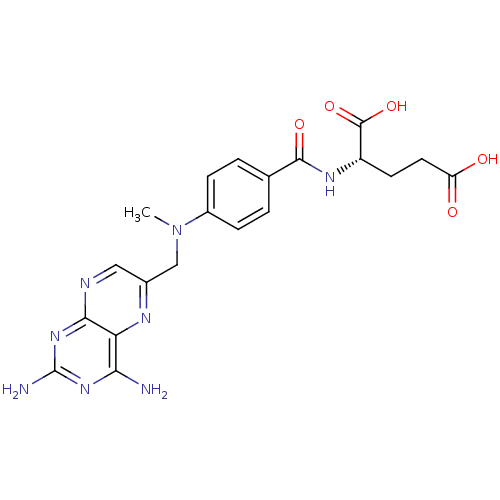
TargetDihydrofolate reductase(Mycobacterium tuberculosis)
Institute Of Chemical Technology
Curated by ChEMBL
Institute Of Chemical Technology
Curated by ChEMBL
Affinity DataIC50: 8.30nMAssay Description:Inhibition of recombinant Mycobacterium tuberculosis DHFR using 45 uM DHF as substrate by spectrophotometryMore data for this Ligand-Target Pair
TargetDihydrofolate reductase(Mycobacterium tuberculosis)
Institute Of Chemical Technology
Curated by ChEMBL
Institute Of Chemical Technology
Curated by ChEMBL
Affinity DataIC50: 35nMAssay Description:Inhibition of Mycobacterium tuberculosis His-tagged DHFR assessed as reduction in consumption of NADPH using DHF as substrate measured every 30 secs ...More data for this Ligand-Target Pair

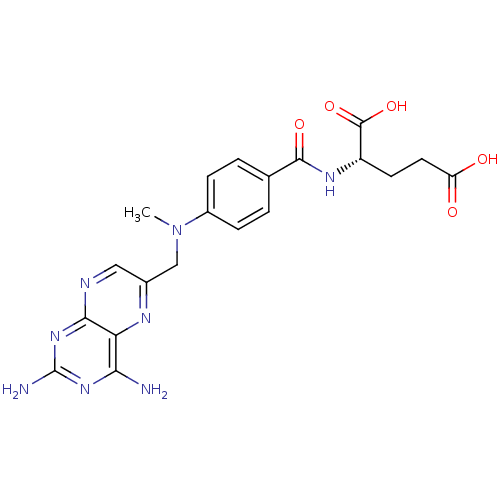
 3D Structure (crystal)
3D Structure (crystal)