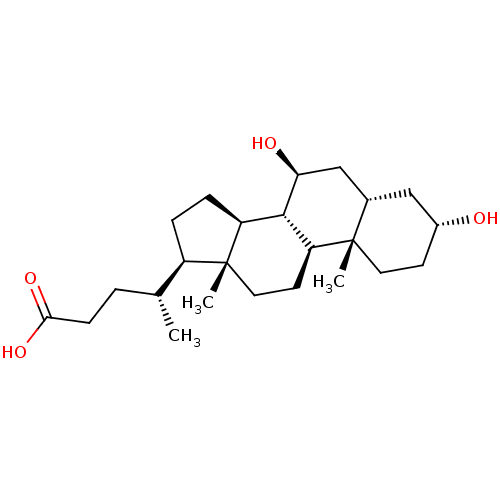
Affinity DataEC50: 361nMpH: 7.2 T: 37°CAssay Description:To evaluate FXR activity of compounds, COS1 (ATCC) cells were transfected with hFXR expression plasmid, mouse retinoid X receptor expression plasmid,...More data for this Ligand-Target Pair
Affinity DataEC50: 85nMAssay Description:Agonist activity at full length FXR in human Huh-7 cellsMore data for this Ligand-Target Pair
Affinity DataEC50: 98nMAssay Description:Binding affinity to FXR assessed as ligand-dependent SRC1 recruitment by FRET based co-activator assayMore data for this Ligand-Target Pair
Affinity DataEC50: 85nMAssay Description:Agonist activity at FXR (unknown origin) by reporter gene assayMore data for this Ligand-Target Pair
Affinity DataEC50: 9.90E+4nMAssay Description:Agonist activity at FXR (unknown origin) by coactivator recruitment assayMore data for this Ligand-Target Pair
Affinity DataEC50: 160nMAssay Description:Agonist activity at human FXR expressed in human HeLa cells assessed as receptor activation by BSEP promoter-driven firefly luciferase reporter gene ...More data for this Ligand-Target Pair
Affinity DataEC50: 160nMAssay Description:Agonist activity at human full length FXR expressed in HeLa cells cotransfected with pSG5-human RXR after 24 hrs by Dual-Glo luciferase reporter gene...More data for this Ligand-Target Pair
Affinity DataEC50: 200nMAssay Description:Agonist activity at human GST-fused FXR LBD assessed as coactivator interaction with receptor ligand binding domain by Alphascreen assayMore data for this Ligand-Target Pair
Affinity DataEC50: >5.00E+4nMAssay Description:Agonist activity at human FXR expressed in COS1 cells by luciferase assayMore data for this Ligand-Target Pair
Affinity DataEC50: 99nMAssay Description:Agonist activity at FXR expressed in COS1 cells by cell-based bioluminescence assayMore data for this Ligand-Target Pair
Affinity DataEC50: 99nMAssay Description:Effective concentration against Farnesoid X receptor (FXR)More data for this Ligand-Target Pair
Affinity DataEC50: 90nMAssay Description:Binding affinity for human Farnesoid X receptor in FRET assayMore data for this Ligand-Target Pair
Affinity DataEC50: 99nMAssay Description:Activation of human farnesoid X receptorMore data for this Ligand-Target Pair
Affinity DataEC50: 550nMAssay Description:The assay utilizes non-human mammalian cells, CHO (Chinese hamster ovary) cells engineered to express human NR1H4 protein (referred to as FXR). Repor...More data for this Ligand-Target Pair
Affinity DataEC50: 42nMAssay Description:Agonist activity at human FXR expressed in HEK293T cells assessed as BSEP promoter driven cellular transcriptional activity after 24 hrs by luciferas...More data for this Ligand-Target Pair
Affinity DataEC50: 550nMAssay Description:The potency and efficacy of the compounds of the invention on TGR5 receptor was evaluated using in vitro assays which carried out using the express k...More data for this Ligand-Target Pair
Affinity DataEC50: 100nMAssay Description:Agonist activity at FXR (unknown origin) assessed as recruitment of SRC1 peptide to FXR by FRET assayMore data for this Ligand-Target Pair
Affinity DataEC50: 550nMAssay Description:Determination of a ligand mediated Gal4 promoter driven transactivation to quantify ligand binding mediated activation of FXR. FXR Reporter Assay kit...More data for this Ligand-Target Pair
Affinity DataEC50: 180nMAssay Description:Agonist activity at recombinant human GST-tagged FXR LBD (193 to 472 residues) expressed in baculovirus-infected insect cells assessed as recruitment...More data for this Ligand-Target Pair
Affinity DataEC50: 10nMAssay Description:Agonist activity at human FXR expressed in HEK293T cells assessed as BSEP promoter driven cellular transcriptional activity after 24 hrs by luciferas...More data for this Ligand-Target Pair
Affinity DataEC50: 99nMAssay Description:Agonist activity at FXR (unknown origin)More data for this Ligand-Target Pair
Affinity DataEC50: 85nMAssay Description:Agonist activity at FXR (unknown origin) by cell based assayMore data for this Ligand-Target Pair
Affinity DataEC50: 500nMAssay Description:Transactivation of human FXR (unknown origin) expressed in HepG2 cells co-expressing pSG5RXR/pGL4.70 after 24 hrs post transfection by luciferase rep...More data for this Ligand-Target Pair
Affinity DataEC50: 1.00E+4nMAssay Description:Agonist activity at GST-tagged human FXR-LBD (193 to 472 residues) using biotinylated SRC-1 peptide as substrate incubated for 1 hrs by HTRF assayMore data for this Ligand-Target Pair
Affinity DataEC50: 4.20E+4nMAssay Description:Agonist activity at FXR transfected in human HEK293T cells o-expressing pGL3/hBSEP/luc incubated for 24 hrs by Steady-Glo reagent based luciferase re...More data for this Ligand-Target Pair
Affinity DataEC50: 290nMAssay Description:Agonist activity at GST-tagged human FXR LBD assessed as induction of biotinylated coactivator SRC-1 peptide recruitment measured after 2 hrs by stre...More data for this Ligand-Target Pair
Affinity DataEC50: 710nMAssay Description:Agonist activity at human FXR expressed in HEK293 cells measured after 18 hrs by steady-glo luciferase reporter gene assayMore data for this Ligand-Target Pair
Affinity DataEC50: >1.00E+5nMAssay Description:Agonist activity at glutathione transferase-tagged human FXR-LBD using biotinylated Src-1 peptide incubated for 30 mins by recruitment coactivator as...More data for this Ligand-Target Pair
Affinity DataEC50: 150nMAssay Description:Agonist activity at glutathione transferase-tagged human FXR-LBD using biotinylated Src-1 peptide incubated for 30 mins by recruitment coactivator as...More data for this Ligand-Target Pair
Affinity DataEC50: 75nMAssay Description:Purchasing Invitrogen PV4833 Kit.Step 1, the compound was weighed and dissolved in 100% DMSO, the highest concentration is 3000 μM, and then dil...More data for this Ligand-Target Pair
Affinity DataEC50: 324nMAssay Description:Agonist activity at human FXR expressed in human Huh-7 cells co-transfected with FXRE-Luc reporter plasmid assessed as receptor transactivation incub...More data for this Ligand-Target Pair
Affinity DataEC50: 99nMAssay Description:Agonist activity at FXR (unknown origin) by fluorescence resonance energy tranfer assayMore data for this Ligand-Target Pair
Affinity DataEC50: 99nMAssay Description:Binding affinity for Farnesoid X Receptor (FXR)More data for this Ligand-Target Pair
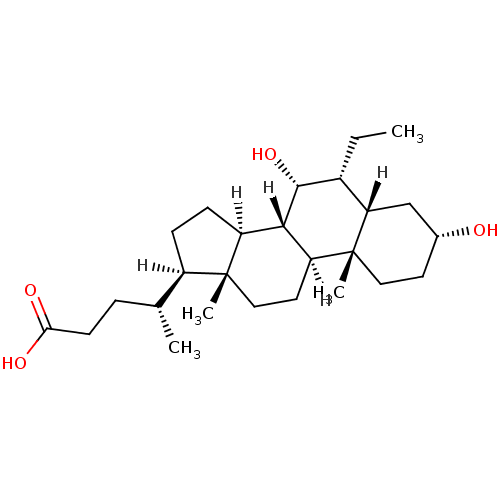
 3D Structure (crystal)
3D Structure (crystal)