Affinity DataKi: 1.10nM ΔG°: -53.2kJ/molepH: 4.5 T: 2°CAssay Description:Enzyme activities were assayed by monitoring the hydrolysis of substrate in the presence or absence of inhibitor compounds. The hydrolysis was record...More data for this Ligand-Target Pair

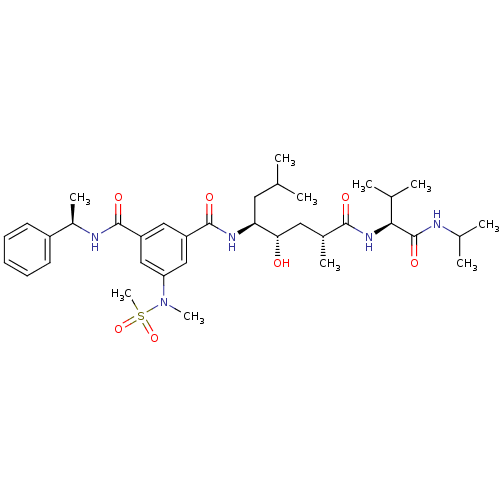
 3D Structure (crystal)
3D Structure (crystal)