TargetGlutamate receptor 4(Homo sapiens (Human))
The Danish University Of Pharmaceutical Sciences
Curated by ChEMBL
The Danish University Of Pharmaceutical Sciences
Curated by ChEMBL
Affinity DataKi: 40nMAssay Description:Binding affinity for ionotropic Glutamate receptor AMPA 4 expressed in Sf9 cellsMore data for this Ligand-Target Pair
TargetGlutamate receptor 4(Homo sapiens (Human))
The Danish University Of Pharmaceutical Sciences
Curated by ChEMBL
The Danish University Of Pharmaceutical Sciences
Curated by ChEMBL
Affinity DataKi: 155nMAssay Description:Displacement of [3H]AMPA from human Ionotropic glutamate receptor AMPA 4 expressed in HEK293 cellsMore data for this Ligand-Target Pair

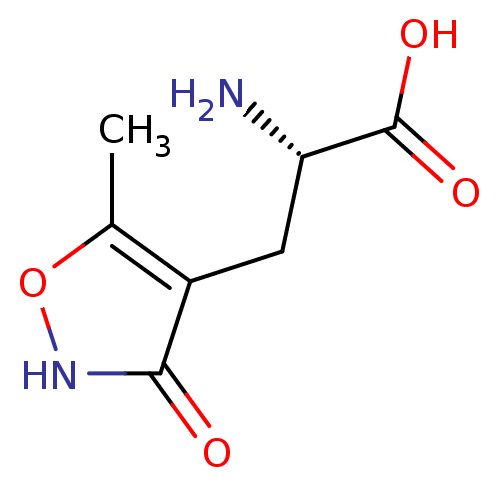
 3D Structure (crystal)
3D Structure (crystal)