Target5'-methylthioadenosine/S-adenosylhomocysteine nucleosidase(Escherichia coli (strain K12))
Albert Einstein College Of Medicine
Albert Einstein College Of Medicine
Affinity DataKi: 0.208nM ΔG°: -55.3kJ/molepH: 7.5 T: 2°CAssay Description:Purified MTAN activity in MTAN enzyme inhibition assayMore data for this Ligand-Target Pair

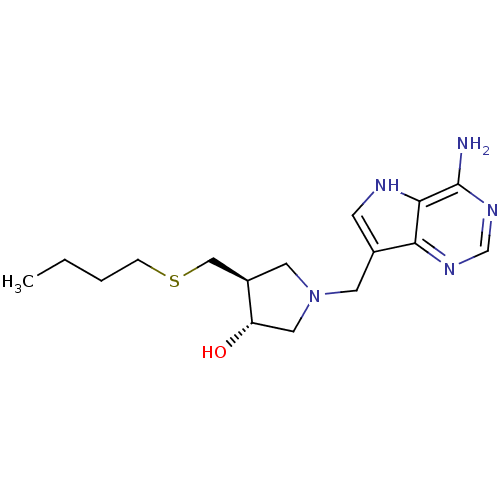
 3D Structure (crystal)
3D Structure (crystal)