Reaction Details Report a problem with these data
Report a problem with these data
 Report a problem with these data
Report a problem with these dataTarget
HIV-1 protease
Ligand
BDBM161
Substrate
n/a
Meas. Tech.
Protease Inhibtion Assay
pH
5.5±0
Ki
0.018±0.0 nM
Citation
 Rodgers, JD; Lam, PY; Johnson, BL; Wang, H; Li, R; Ru, Y; Ko, SS; Seitz, SP; Trainor, GL; Anderson, PS; Klabe, RM; Bacheler, LT; Cordova, B; Garber, S; Reid, C; Wright, MR; Chang, CH; Erickson-Viitanen, S Design and selection of DMP 850 and DMP 851: the next generation of cyclic urea HIV protease inhibitors. Chem Biol 5:597-608 (1998) [PubMed] Article
Rodgers, JD; Lam, PY; Johnson, BL; Wang, H; Li, R; Ru, Y; Ko, SS; Seitz, SP; Trainor, GL; Anderson, PS; Klabe, RM; Bacheler, LT; Cordova, B; Garber, S; Reid, C; Wright, MR; Chang, CH; Erickson-Viitanen, S Design and selection of DMP 850 and DMP 851: the next generation of cyclic urea HIV protease inhibitors. Chem Biol 5:597-608 (1998) [PubMed] Article More Info.:
Target
Name:
HIV-1 protease
Synonyms:
HIV-1 protease wild type
Type:
Protein
Mol. Mass.:
10757.68
Organism:
Human immunodeficiency virus
Description:
O90785
Residue:
99
Sequence:
PQITLWQRPLVTVKIGGQLREALLDTGADDTVLEDINLPGKWKPKMIGGIGGFIKVKQYEQVLIEICGKKAIGTVLVGPTPVNIIGRNMLTQIGCTLNF
Inhibitor
Name:
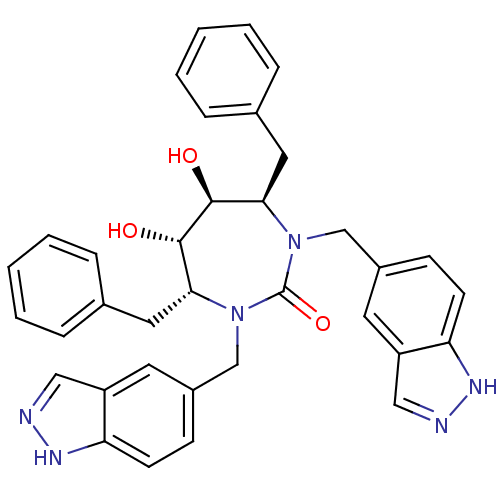
BDBM161
Synonyms:
(4R,5S,6S,7R)-4,7-dibenzyl-5,6-dihydroxy-1,3-bis(1H-indazol-5-ylmethyl)-1,3-diazepan-2-one | CHEMBL78671 | Cyclic Urea | Indazole, 1
Type:
n/a
Emp. Form.:
C35H34N6O3
Mol. Mass.:
586.6829
SMILES:
O[C@@H]1[C@@H](O)[C@@H](Cc2ccccc2)N(Cc2ccc3[nH]ncc3c2)C(=O)N(Cc2ccc3[nH]ncc3c2)[C@@H]1Cc1ccccc1