Reaction Details Report a problem with these data
Report a problem with these data
 Report a problem with these data
Report a problem with these dataTarget
E3 ubiquitin-protein ligase TRIM33
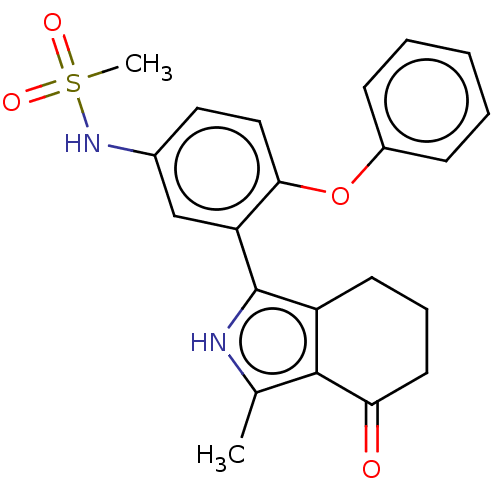
Ligand
BDBM342597
Substrate
n/a
Meas. Tech.
ChEBML_1687850
IC50
>10000±n/a nM
Citation
 Hasvold, LA; Sheppard, GS; Wang, L; Fidanze, SD; Liu, D; Pratt, JK; Mantei, RA; Wada, CK; Hubbard, R; Shen, Y; Lin, X; Huang, X; Warder, SE; Wilcox, D; Li, L; Buchanan, FG; Smithee, L; Albert, DH; Magoc, TJ; Park, CH; Petros, AM; Panchal, SC; Sun, C; Kovar, P; Soni, NB; Elmore, SW; Kati, WM; McDaniel, KF Methylpyrrole inhibitors of BET bromodomains. Bioorg Med Chem Lett 27:2225-2233 (2017) [PubMed] Article
Hasvold, LA; Sheppard, GS; Wang, L; Fidanze, SD; Liu, D; Pratt, JK; Mantei, RA; Wada, CK; Hubbard, R; Shen, Y; Lin, X; Huang, X; Warder, SE; Wilcox, D; Li, L; Buchanan, FG; Smithee, L; Albert, DH; Magoc, TJ; Park, CH; Petros, AM; Panchal, SC; Sun, C; Kovar, P; Soni, NB; Elmore, SW; Kati, WM; McDaniel, KF Methylpyrrole inhibitors of BET bromodomains. Bioorg Med Chem Lett 27:2225-2233 (2017) [PubMed] Article More Info.:
Target
Name:
E3 ubiquitin-protein ligase TRIM33
Synonyms:
6.3.2.- | E3 ubiquitin-protein ligase TRIM33 | Ectodermin homolog | KIAA1113 | KIAA1113 | Protein Rfg7 | RET-fused gene 7 protein | RFG7 | TIF1-gamma | TIF1G | TRI33_HUMAN | TRIM33 | Transcription intermediary factor 1-gamma | Tripartite motif-containing protein 33
Type:
PROTEIN
Mol. Mass.:
122534.89
Organism:
Homo sapiens (Human)
Description:
ChEMBL_105347
Residue:
1127
Sequence:
MAENKGGGEAESGGGGSGSAPVTAGAAGPAAQEAEPPLTAVLVEEEEEEGGRAGAEGGAAGPDDGGVAAASSGSAQAASSPAASVGTGVAGGAVSTPAPAPASAPAPGPSAGPPPGPPASLLDTCAVCQQSLQSRREAEPKLLPCLHSFCLRCLPEPERQLSVPIPGGSNGDIQQVGVIRCPVCRQECRQIDLVDNYFVKDTSEAPSSSDEKSEQVCTSCEDNASAVGFCVECGEWLCKTCIEAHQRVKFTKDHLIRKKEDVSESVGASGQRPVFCPVHKQEQLKLFCETCDRLTCRDCQLLEHKEHRYQFLEEAFQNQKGAIENLLAKLLEKKNYVHFAATQVQNRIKEVNETNKRVEQEIKVAIFTLINEINKKGKSLLQQLENVTKERQMKLLQQQNDITGLSRQVKHVMNFTNWAIASGSSTALLYSKRLITFQLRHILKARCDPVPAANGAIRFHCDPTFWAKNVVNLGNLVIESKPAPGYTPNVVVGQVPPGTNHISKTPGQINLAQLRLQHMQQQVYAQKHQQLQQMRMQQPPAPVPTTTTTTQQHPRQAAPQMLQQQPPRLISVQTMQRGNMNCGAFQAHQMRLAQNAARIPGIPRHSGPQYSMMQPHLQRQHSNPGHAGPFPVVSVHNTTINPTSPTTATMANANRGPTSPSVTAIELIPSVTNPENLPSLPDIPPIQLEDAGSSSLDNLLSRYISGSHLPPQPTSTMNPSPGPSALSPGSSGLSNSHTPVRPPSTSSTGSRGSCGSSGRTAEKTSLSFKSDQVKVKQEPGTEDEICSFSGGVKQEKTEDGRRSACMLSSPESSLTPPLSTNLHLESELDALASLENHVKIEPADMNESCKQSGLSSLVNGKSPIRSLMHRSARIGGDGNNKDDDPNEDWCAVCQNGGDLLCCEKCPKVFHLTCHVPTLLSFPSGDWICTFCRDIGKPEVEYDCDNLQHSKKGKTAQGLSPVDQRKCERLLLYLYCHELSIEFQEPVPASIPNYYKIIKKPMDLSTVKKKLQKKHSQHYQIPDDFVADVRLIFKNCERFNEMMKVVQVYADTQEINLKADSEVAQAGKAVALYFEDKLTEIYSDRTFAPLPEFEQEEDDGEVTEDSDEDFIQPRRKRLKSDERPVHIK