Reaction Details Report a problem with these data
Report a problem with these data
 Report a problem with these data
Report a problem with these dataTarget
Gag-Pol polyprotein [489-587]
Ligand
BDBM164
Substrate
n/a
Meas. Tech.
ChEMBL_157733 (CHEMBL765171)
Ki
0.023±n/a nM
Citation
 De Lucca, GV; Kim, UT; Liang, J; Cordova, B; Klabe, RM; Garber, S; Bacheler, LT; Lam, GN; Wright, MR; Logue, KA; Erickson-Viitanen, S; Ko, SS; Trainor, GL Nonsymmetric P2/P2' cyclic urea HIV protease inhibitors. Structure-activity relationship, bioavailability, and resistance profile of monoindazole-substituted P2 analogues. J Med Chem 41:2411-23 (1998) [PubMed] Article
De Lucca, GV; Kim, UT; Liang, J; Cordova, B; Klabe, RM; Garber, S; Bacheler, LT; Lam, GN; Wright, MR; Logue, KA; Erickson-Viitanen, S; Ko, SS; Trainor, GL Nonsymmetric P2/P2' cyclic urea HIV protease inhibitors. Structure-activity relationship, bioavailability, and resistance profile of monoindazole-substituted P2 analogues. J Med Chem 41:2411-23 (1998) [PubMed] Article More Info.:
Target
Name:
Gag-Pol polyprotein [489-587]
Synonyms:
Human immunodeficiency virus type 1 protease | POL_HV1H2 | Pol polyprotein | gag-pol
Type:
Enzyme Subunit
Mol. Mass.:
10781.16
Organism:
Human immunodeficiency virus type 1
Description:
P04585[489-587]
Residue:
99
Sequence:
PQVTLWQRPLVTIKIGGQLKEALLDTGADDTVLEEMSLPGRWKPKMIGGIGGFIKVRQYDQILIEICGHKAIGTVLVGPTPVNIIGRNLLTQIGCTLNF
Inhibitor
Name:
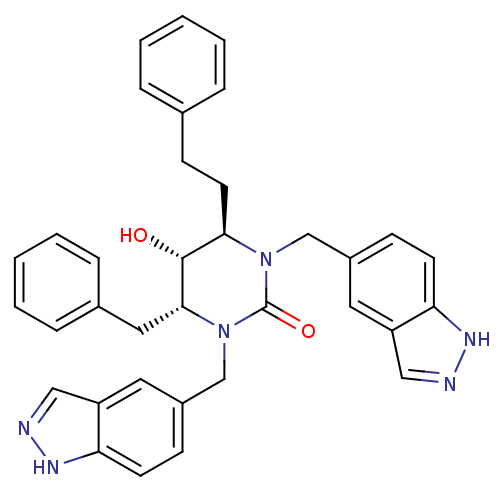
BDBM164
Synonyms:
(4R,5R,6R)-4-benzyl-5-hydroxy-1,3-bis(1H-indazol-5-ylmethyl)-6-(2-phenylethyl)-1,3-diazinan-2-one | CHEMBL288254 | Cyclic Urea | [4R-(4,5,6)]-Tetrahydro-5-hydroxy-1,3-bis(1H-indazol-5-ylmethyl)-4-(2-phenylethyl)-6-(phenylmethyl)-2(1H)-pyrimidinone
Type:
n/a
Emp. Form.:
C35H34N6O2
Mol. Mass.:
570.6835
SMILES:
O[C@@H]1[C@@H](CCc2ccccc2)N(Cc2ccc3[nH]ncc3c2)C(=O)N(Cc2ccc3[nH]ncc3c2)[C@@H]1Cc1ccccc1