Reaction Details Report a problem with these data
Report a problem with these data
 Report a problem with these data
Report a problem with these dataTarget
Vascular endothelial growth factor receptor 3
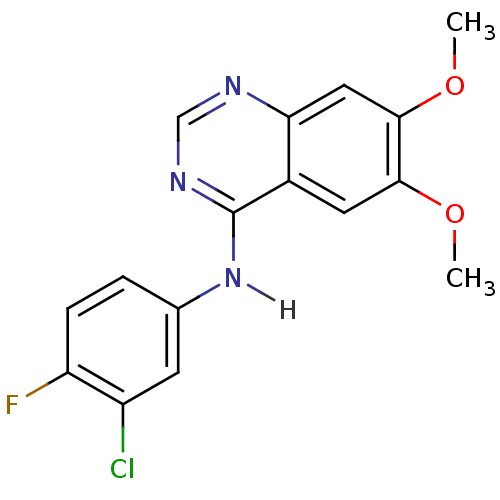
Ligand
BDBM4626
Substrate
n/a
Meas. Tech.
ChEMBL_566610 (CHEMBL964949)
IC50
2400±n/a nM
Citation
 Tasler, S; Müller, O; Wieber, T; Herz, T; Krauss, R; Totzke, F; Kubbutat, MH; Schächtele, C N-substituted 2'-(aminoaryl)benzothiazoles as kinase inhibitors: hit identification and scaffold hopping. Bioorg Med Chem Lett 19:1349-56 (2009) [PubMed] Article
Tasler, S; Müller, O; Wieber, T; Herz, T; Krauss, R; Totzke, F; Kubbutat, MH; Schächtele, C N-substituted 2'-(aminoaryl)benzothiazoles as kinase inhibitors: hit identification and scaffold hopping. Bioorg Med Chem Lett 19:1349-56 (2009) [PubMed] Article More Info.:
Target
Name:
Vascular endothelial growth factor receptor 3
Synonyms:
FLT-4 | FLT4 | Fms-like tyrosine kinase 4 | Focal adhesion kinase 1/vascular endothelial growth factor receptor 3 | VEGFR-3 | VEGFR3 | VGFR3_HUMAN | Vascular endothelial growth factor receptor | Vascular endothelial growth factor receptor 3 (VEGFR-3) | Vascular endothelial growth factor receptor 3 (VEGFR3)
Type:
Protein
Mol. Mass.:
152749.58
Organism:
Homo sapiens (Human)
Description:
P35916-2
Residue:
1363
Sequence:
MQRGAALCLRLWLCLGLLDGLVSGYSMTPPTLNITEESHVIDTGDSLSISCRGQHPLEWAWPGAQEAPATGDKDSEDTGVVRDCEGTDARPYCKVLLLHEVHANDTGSYVCYYKYIKARIEGTTAASSYVFVRDFEQPFINKPDTLLVNRKDAMWVPCLVSIPGLNVTLRSQSSVLWPDGQEVVWDDRRGMLVSTPLLHDALYLQCETTWGDQDFLSNPFLVHITGNELYDIQLLPRKSLELLVGEKLVLNCTVWAEFNSGVTFDWDYPGKQAERGKWVPERRSQQTHTELSSILTIHNVSQHDLGSYVCKANNGIQRFRESTEVIVHENPFISVEWLKGPILEATAGDELVKLPVKLAAYPPPEFQWYKDGKALSGRHSPHALVLKEVTEASTGTYTLALWNSAAGLRRNISLELVVNVPPQIHEKEASSPSIYSRHSRQALTCTAYGVPLPLSIQWHWRPWTPCKMFAQRSLRRRQQQDLMPQCRDWRAVTTQDAVNPIESLDTWTEFVEGKNKTVSKLVIQNANVSAMYKCVVSNKVGQDERLIYFYVTTIPDGFTIESKPSEELLEGQPVLLSCQADSYKYEHLRWYRLNLSTLHDAHGNPLLLDCKNVHLFATPLAASLEEVAPGARHATLSLSIPRVAPEHEGHYVCEVQDRRSHDKHCHKKYLSVQALEAPRLTQNLTDLLVNVSDSLEMQCLVAGAHAPSIVWYKDERLLEEKSGVDLADSNQKLSIQRVREEDAGRYLCSVCNAKGCVNSSASVAVEGSEDKGSMEIVILVGTGVIAVFFWVLLLLIFCNMRRPAHADIKTGYLSIIMDPGEVPLEEQCEYLSYDASQWEFPRERLHLGRVLGYGAFGKVVEASAFGIHKGSSCDTVAVKMLKEGATASEHRALMSELKILIHIGNHLNVVNLLGACTKPQGPLMVIVEFCKYGNLSNFLRAKRDAFSPCAEKSPEQRGRFRAMVELARLDRRRPGSSDRVLFARFSKTEGGARRASPDQEAEDLWLSPLTMEDLVCYSFQVARGMEFLASRKCIHRDLAARNILLSESDVVKICDFGLARDIYKDPDYVRKGSARLPLKWMAPESIFDKVYTTQSDVWSFGVLLWEIFSLGASPYPGVQINEEFCQRLRDGTRMRAPELATPAIRRIMLNCWSGDPKARPAFSELVEILGDLLQGRGLQEEEEVCMAPRSSQSSEEGSFSQVSTMALHIAQADAEDSPPSLQRHSLAARYYNWVSFPGCLARGAETRGSSRMKTFEEFPMTPTTYKGSVDNQTDSGMVLASEEFEQIESRHRQESGFSCKGPGQNVAVTRAHPDSQGRRRRPERGARGGQVFYNSEYGELSEPSEEDHCSPSARVTFFTDNSY