Reaction Details Report a problem with these data
Report a problem with these data
 Report a problem with these data
Report a problem with these dataTarget
Misshapen-like kinase 1
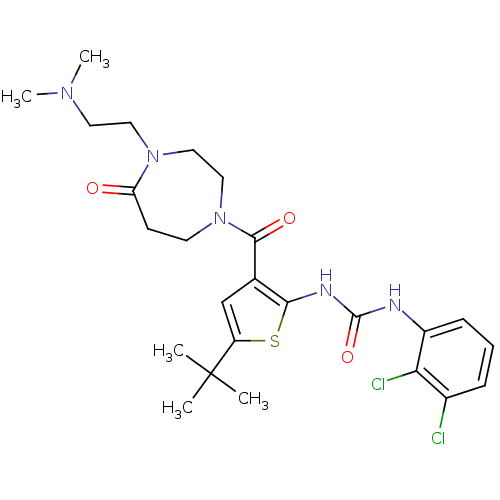
Ligand
BDBM50359359
Substrate
n/a
Meas. Tech.
ChEMBL_793919 (CHEMBL1931511)
IC50
>30000±n/a nM
Citation
 Moffett, K; Konteatis, Z; Nguyen, D; Shetty, R; Ludington, J; Fujimoto, T; Lee, KJ; Chai, X; Namboodiri, H; Karpusas, M; Dorsey, B; Guarnieri, F; Bukhtiyarova, M; Springman, E; Michelotti, E Discovery of a novel class of non-ATP site DFG-out state p38 inhibitors utilizing computationally assisted virtual fragment-based drug design (vFBDD). Bioorg Med Chem Lett 21:7155-65 (2011) [PubMed] Article
Moffett, K; Konteatis, Z; Nguyen, D; Shetty, R; Ludington, J; Fujimoto, T; Lee, KJ; Chai, X; Namboodiri, H; Karpusas, M; Dorsey, B; Guarnieri, F; Bukhtiyarova, M; Springman, E; Michelotti, E Discovery of a novel class of non-ATP site DFG-out state p38 inhibitors utilizing computationally assisted virtual fragment-based drug design (vFBDD). Bioorg Med Chem Lett 21:7155-65 (2011) [PubMed] Article More Info.:
Target
Name:
Misshapen-like kinase 1
Synonyms:
B55 | GCK family kinase MiNK | MAP4K6 | MAPK/ERK kinase kinase kinase 6 | MEK kinase kinase 6 | MEKKK 6 | MINK | MINK1 | MINK1_HUMAN | Misshapen-like kinase 1 (MINK) | Misshapen/NIK-related kinase | Mitogen-activated protein kinase kinase kinase kinase 6 | Voltage-gated potassium channel subunit Kv7.1/Misshapen-like kinase 1 | YSK2 | ZC3
Type:
PROTEIN
Mol. Mass.:
149838.92
Organism:
Homo sapiens (Human)
Description:
ChEMBL_793919
Residue:
1332
Sequence:
MGDPAPARSLDDIDLSALRDPAGIFELVEVVGNGTYGQVYKGRHVKTGQLAAIKVMDVTEDEEEEIKQEINMLKKYSHHRNIATYYGAFIKKSPPGNDDQLWLVMEFCGAGSVTDLVKNTKGNALKEDCIAYICREILRGLAHLHAHKVIHRDIKGQNVLLTENAEVKLVDFGVSAQLDRTVGRRNTFIGTPYWMAPEVIACDENPDATYDYRSDIWSLGITAIEMAEGAPPLCDMHPMRALFLIPRNPPPRLKSKKWSKKFIDFIDTCLIKTYLSRPPTEQLLKFPFIRDQPTERQVRIQLKDHIDRSRKKRGEKEETEYEYSGSEEEDDSHGEEGEPSSIMNVPGESTLRREFLRLQQENKSNSEALKQQQQLQQQQQRDPEAHIKHLLHQRQRRIEEQKEERRRVEEQQRREREQRKLQEKEQQRRLEDMQALRREEERRQAEREQEYKRKQLEEQRQSERLQRQLQQEHAYLKSLQQQQQQQQLQKQQQQQLLPGDRKPLYHYGRGMNPADKPAWAREVEERTRMNKQQNSPLAKSKPGSTGPEPPIPQASPGPPGPLSQTPPMQRPVEPQEGPHKSLVAHRVPLKPYAAPVPRSQSLQDQPTRNLAAFPASHDPDPAIPAPTATPSARGAVIRQNSDPTSEGPGPSPNPPAWVRPDNEAPPKVPQRTSSIATALNTSGAGGSRPAQAVRARPRSNSAWQIYLQRRAERGTPKPPGPPAQPPGPPNASSNPDLRRSDPGWERSDSVLPASHGHLPQAGSLERNRVGVSSKPDSSPVLSPGNKAKPDDHRSRPGRPADFVLLKERTLDEAPRPPKKAMDYSSSSEEVESSEDDEEEGEGGPAEGSRDTPGGRSDGDTDSVSTMVVHDVEEITGTQPPYGGGTMVVQRTPEEERNLLHADSNGYTNLPDVVQPSHSPTENSKGQSPPSKDGSGDYQSRGLVKAPGKSSFTMFVDLGIYQPGGSGDSIPITALVGGEGTRLDQLQYDVRKGSVVNVNPTNTRAHSETPEIRKYKKRFNSEILCAALWGVNLLVGTENGLMLLDRSGQGKVYGLIGRRRFQQMDVLEGLNLLITISGKRNKLRVYYLSWLRNKILHNDPEVEKKQGWTTVGDMEGCGHYRVVKYERIKFLVIALKSSVEVYAWAPKPYHKFMAFKSFADLPHRPLLVDLTVEEGQRLKVIYGSSAGFHAVDVDSGNSYDIYIPVHIQSQITPHAIIFLPNTDGMEMLLCYEDEGVYVNTYGRIIKDVVLQWGEMPTSVAYICSNQIMGWGEKAIEIRSVETGHLDGVFMHKRAQRLKFLCERNDKVFFASVRSGGSSQVYFMTLNRNCIMNW