Reaction Details Report a problem with these data
Report a problem with these data
 Report a problem with these data
Report a problem with these dataTarget
Adenosine receptor A3
Ligand
BDBM50106537
Substrate
n/a
Meas. Tech.
ChEMBL_31703 (CHEMBL645784)
Ki
2.1±n/a nM
Citation
 Gao, ZG; Kim, SK; Biadatti, T; Chen, W; Lee, K; Barak, D; Kim, SG; Johnson, CR; Jacobson, KA Structural determinants of A(3) adenosine receptor activation: nucleoside ligands at the agonist/antagonist boundary. J Med Chem 45:4471-84 (2002) [PubMed] Article
Gao, ZG; Kim, SK; Biadatti, T; Chen, W; Lee, K; Barak, D; Kim, SG; Johnson, CR; Jacobson, KA Structural determinants of A(3) adenosine receptor activation: nucleoside ligands at the agonist/antagonist boundary. J Med Chem 45:4471-84 (2002) [PubMed] Article More Info.:
Target
Name:
Adenosine receptor A3
Synonyms:
A3 adenosine receptor (hA3) | AA3R_HUMAN | ADORA3 | Adenosine A3 receptor (A3AR)
Type:
G Protein-Coupled Receptor (GPCR)
Mol. Mass.:
36197.32
Organism:
Homo sapiens (Human)
Description:
P0DMS8
Residue:
318
Sequence:
MPNNSTALSLANVTYITMEIFIGLCAIVGNVLVICVVKLNPSLQTTTFYFIVSLALADIAVGVLVMPLAIVVSLGITIHFYSCLFMTCLLLIFTHASIMSLLAIAVDRYLRVKLTVRYKRVTTHRRIWLALGLCWLVSFLVGLTPMFGWNMKLTSEYHRNVTFLSCQFVSVMRMDYMVYFSFLTWIFIPLVVMCAIYLDIFYIIRNKLSLNLSNSKETGAFYGREFKTAKSLFLVLFLFALSWLPLSIINCIIYFNGEVPQLVLYMGILLSHANSMMNPIVYAYKIKKFKETYLLILKACVVCHPSDSLDTSIEKNSE
Inhibitor
Name:
BDBM50106537
Synonyms:
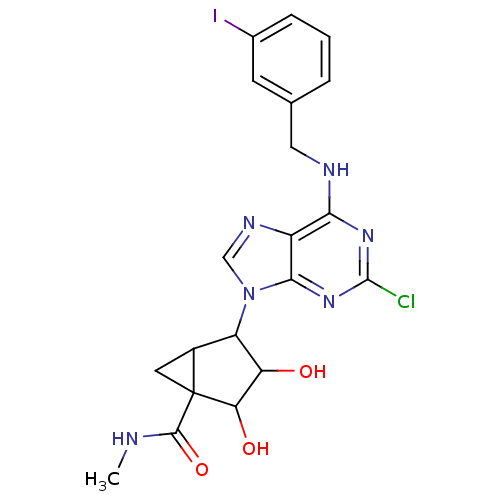
4-[2-Chloro-6-(3-iodo-benzylamino)-purin-9-yl]-2,3-dihydroxy-bicyclo[3.1.0]hexane-1-carboxylic acid methylamide | CHEMBL133387
Type:
Small organic molecule
Emp. Form.:
C20H20ClIN6O3
Mol. Mass.:
554.769
SMILES:
CNC(=O)C12CC1C(C(O)C2O)n1cnc2c(NCc3cccc(I)c3)nc(Cl)nc12