Reaction Details Report a problem with these data
Report a problem with these data
 Report a problem with these data
Report a problem with these dataTarget
Inosine-5'-monophosphate dehydrogenase
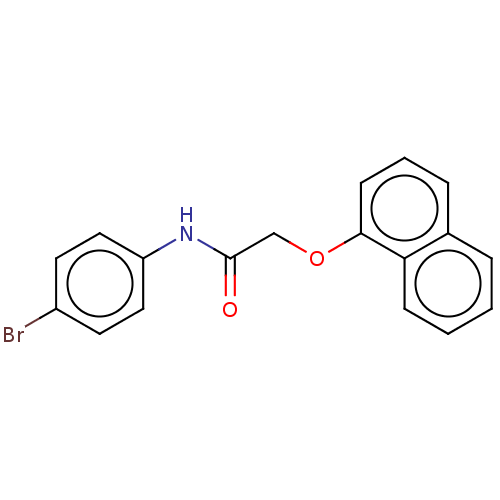
Ligand
BDBM50100896
Substrate
n/a
Meas. Tech.
ChEMBL_1452429 (CHEMBL3362091)
IC50
760±n/a nM
Citation
 Mandapati, K; Gorla, SK; House, AL; McKenney, ES; Zhang, M; Rao, SN; Gollapalli, DR; Mann, BJ; Goldberg, JB; Cuny, GD; Glomski, IJ; Hedstrom, L Repurposing cryptosporidium inosine 5'-monophosphate dehydrogenase inhibitors as potential antibacterial agents. ACS Med Chem Lett 5:846-50 (2014) [PubMed] Article
Mandapati, K; Gorla, SK; House, AL; McKenney, ES; Zhang, M; Rao, SN; Gollapalli, DR; Mann, BJ; Goldberg, JB; Cuny, GD; Glomski, IJ; Hedstrom, L Repurposing cryptosporidium inosine 5'-monophosphate dehydrogenase inhibitors as potential antibacterial agents. ACS Med Chem Lett 5:846-50 (2014) [PubMed] Article More Info.:
Target
Name:
Inosine-5'-monophosphate dehydrogenase
Synonyms:
AB164_07585 | AB165_01610 | AB166_11280 | AB167_18060 | AB168_06335 | AB169_08740 | AB170_00090 | AB171_19425 | AB893_00065 | ABW01_29210 | ADT20_14595 | ADT21_00085 | BF27_2004 | guaB
Type:
PROTEIN
Mol. Mass.:
52375.32
Organism:
Bacillus anthracis
Description:
ChEMBL_109471
Residue:
487
Sequence:
MWESKFVKEGLTFDDVLLVPAKSDVLPREVSVKTVLSESLQLNIPLISAGMDTVTEADMAIAMARQGGLGIIHKNMSIEQQAEQVDKVKRSESGVISDPFFLTPEHQVYDAEHLMGKYRISGVPVVNNLDERKLVGIITNRDMRFIQDYSIKISDVMTKEQLITAPVGTTLSEAEKILQKYKIEKLPLVDNNGVLQGLITIKDIEKVIEFPNSAKDKQGRLLVGAAVGVTADAMTRIDALVKASVDAIVLDTAHGHSQGVIDKVKEVRAKYPSLNIIAGNVATAEATKALIEAGANVVKVGIGPGSICTTRVVAGVGVPQLTAVYDCATEARKHGIPVIADGGIKYSGDMVKALAAGAHVVMLGSMFAGVAESPGETEIYQGRQFKVYRGMGSVGAMEKGSKDRYFQEGNKKLVPEGIEGRVPYKGPLADTVHQLVGGLRAGMGYCGAQDLEFLRENAQFIRMSGAGLLESHPHHVQITKEAPNYSL