Reaction Details Report a problem with these data
Report a problem with these data
 Report a problem with these data
Report a problem with these dataTarget
Lysine-specific demethylase 5B
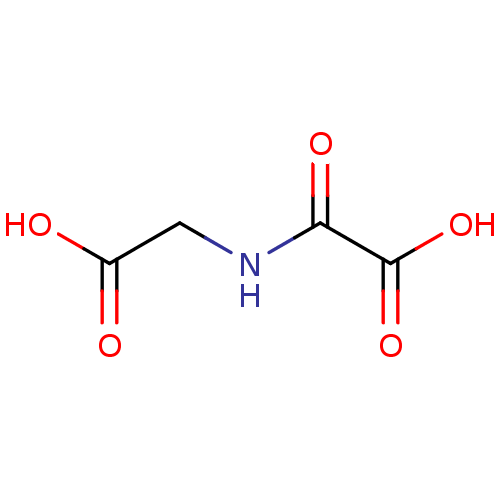
Ligand
BDBM26106
Substrate
n/a
Meas. Tech.
Demethylase AlphaScreen Assay
pH
7.5±n/a
IC50
2.45e+4±n/a nM
Comments
extracted
Citation
 Johansson, C; Velupillai, S; Tumber, A; Szykowska, A; Hookway, ES; Nowak, RP; Strain-Damerell, C; Gileadi, C; Philpott, M; Burgess-Brown, N; Wu, N; Kopec, J; Nuzzi, A; Steuber, H; Egner, U; Badock, V; Munro, S; LaThangue, NB; Westaway, S; Brown, J; Athanasou, N; Prinjha, R; Brennan, PE; Oppermann, U Structural analysis of human KDM5B guides histone demethylase inhibitor development. Nat Chem Biol 12:539-45 (2016) [PubMed] Article
Johansson, C; Velupillai, S; Tumber, A; Szykowska, A; Hookway, ES; Nowak, RP; Strain-Damerell, C; Gileadi, C; Philpott, M; Burgess-Brown, N; Wu, N; Kopec, J; Nuzzi, A; Steuber, H; Egner, U; Badock, V; Munro, S; LaThangue, NB; Westaway, S; Brown, J; Athanasou, N; Prinjha, R; Brennan, PE; Oppermann, U Structural analysis of human KDM5B guides histone demethylase inhibitor development. Nat Chem Biol 12:539-45 (2016) [PubMed] Article More Info.:
Target
Name:
Lysine-specific demethylase 5B
Synonyms:
CT31 | Cancer/testis antigen 31 | Histone demethylase JARID1B | JARID1B | Jumonji/ARID domain-containing protein 1B | KDM5B | KDM5B_HUMAN | Lysine-specific demethylase 5B (KDM5B) | Lysine-specific demethylase 5B (KDM5Flag-5B-FL) | PLU-1 | PLU1 | RBBP2H1 | RBP2-H1 | Retinoblastoma-binding protein 2 homolog 1
Type:
Protein
Mol. Mass.:
175659.67
Organism:
Homo sapiens (Human)
Description:
Q9UGL1
Residue:
1544
Sequence:
MEAATTLHPGPRPALPLGGPGPLGEFLPPPECPVFEPSWEEFADPFAFIHKIRPIAEQTGICKVRPPPDWQPPFACDVDKLHFTPRIQRLNELEAQTRVKLNFLDQIAKYWELQGSTLKIPHVERKILDLFQLNKLVAEEGGFAVVCKDRKWTKIATKMGFAPGKAVGSHIRGHYERILNPYNLFLSGDSLRCLQKPNLTTDTKDKEYKPHDIPQRQSVQPSETCPPARRAKRMRAEAMNIKIEPEETTEARTHNLRRRMGCPTPKCENEKEMKSSIKQEPIERKDYIVENEKEKPKSRSKKATNAVDLYVCLLCGSGNDEDRLLLCDGCDDSYHTFCLIPPLHDVPKGDWRCPKCLAQECSKPQEAFGFEQAARDYTLRTFGEMADAFKSDYFNMPVHMVPTELVEKEFWRLVSTIEEDVTVEYGADIASKEFGSGFPVRDGKIKLSPEEEEYLDSGWNLNNMPVMEQSVLAHITADICGMKLPWLYVGMCFSSFCWHIEDHWSYSINYLHWGEPKTWYGVPGYAAEQLENVMKKLAPELFVSQPDLLHQLVTIMNPNTLMTHEVPVYRTNQCAGEFVITFPRAYHSGFNQGFNFAEAVNFCTVDWLPLGRQCVEHYRLLHRYCVFSHDEMICKMASKADVLDVVVASTVQKDMAIMIEDEKALRETVRKLGVIDSERMDFELLPDDERQCVKCKTTCFMSAISCSCKPGLLVCLHHVKELCSCPPYKYKLRYRYTLDDLYPMMNALKLRAESYNEWALNVNEALEAKINKKKSLVSFKALIEESEMKKFPDNDLLRHLRLVTQDAEKCASVAQQLLNGKRQTRYRSGGGKSQNQLTVNELRQFVTQLYALPCVLSQTPLLKDLLNRVEDFQQHSQKLLSEETPSAAELQDLLDVSFEFDVELPQLAEMRIRLEQARWLEEVQQACLDPSSLTLDDMRRLIDLGVGLAPYSAVEKAMARLQELLTVSEHWDDKAKSLLKARPRHSLNSLATAVKEIEEIPAYLPNGAALKDSVQRARDWLQDVEGLQAGGRVPVLDTLIELVTRGRSIPVHLNSLPRLETLVAEVQAWKECAVNTFLTENSPYSLLEVLCPRCDIGLLGLKRKQRKLKEPLPNGKKKSTKLESLSDLERALTESKETASAMATLGEARLREMEALQSLRLANEGKLLSPLQDVDIKICLCQKAPAAPMIQCELCRDAFHTSCVAVPSISQGLRIWLCPHCRRSEKPPLEKILPLLASLQRIRVRLPEGDALRYMIERTVNWQHRAQQLLSSGNLKFVQDRVGSGLLYSRWQASAGQVSDTNKVSQPPGTTSFSLPDDWDNRTSYLHSPFSTGRSCIPLHGVSPEVNELLMEAQLLQVSLPEIQELYQTLLAKPSPAQQTDRSSPVRPSSEKNDCCRGKRDGINSLERKLKRRLEREGLSSERWERVKKMRTPKKKKIKLSHPKDMNNFKLERERSYELVRSAETHSLPSDTSYSEQEDSEDEDAICPAVSCLQPEGDEVDWVQCDGSCNQWFHQVCVGVSPEMAEKEDYICVRCTVKDAPSRK