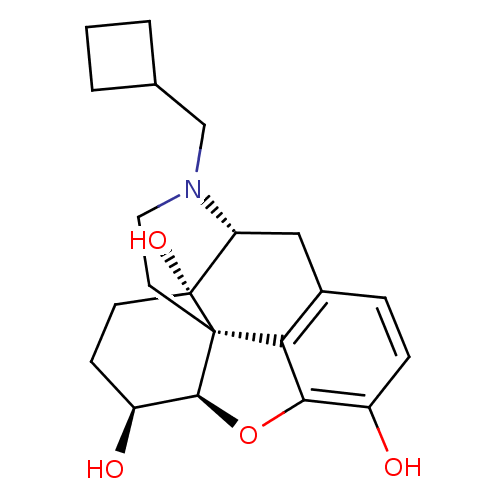
BDBM50105085 17-cyclobutylmethyl-4,5alpha-epoxymorphinan-3,6alpha,14-triol::CHEMBL895::N-cyclobutylmethyl-4,5alpha-epoxy-3,6alpha,14-morphinantriol::NALBUPHINE::US10231963, Table B.1::US10512644, Compound Nalbuphine::US11534436, Compound Table B.1::US9233167, Nalbuphine::US9656961, Example 00118::USRE49340, Rank 9
SMILES O[C@H]1CC[C@@]2(O)[C@H]3Cc4ccc(O)c5O[C@@H]1[C@]2(CCN3CC1CCC1)c45
InChI Key InChIKey=NETZHAKZCGBWSS-CEDHKZHLSA-N
Activity Spreadsheet -- Enzyme Inhibition Constant Data from BindingDB
 Found 45 hits for monomerid = 50105085
Found 45 hits for monomerid = 50105085
Affinity DataKi: 0.890nMAssay Description:Displacement of [3H]DAMGO from human opioid gamma receptor expressed in CHO cellsMore data for this Ligand-Target Pair
Affinity DataKi: 0.980nMAssay Description:The Ki (binding affinity) for μ opioid receptors was determined using a competitive displacement assay as previously described in Neumeyer (Jour...More data for this Ligand-Target Pair
Affinity DataKi: 0.980nMAssay Description:The Ki (binding affinity) for opioid receptors was determined using a competitive displacement assay as previously described in Neumeyer (Journal of ...More data for this Ligand-Target Pair
Affinity DataKi: 0.980nMAssay Description:The Ki (binding affinity) for opioid receptors was determined using a competitive displacement assay as previously described in Neumeyer (Journal of ...More data for this Ligand-Target Pair
Affinity DataKi: 1.30nMAssay Description:The Ki (binding affinity) for μ opioid receptors was determined using a competitive displacement assay as previously described in Neumeyer (Jour...More data for this Ligand-Target Pair
Affinity DataKi: 1.60nMAssay Description:Displacement of [3H]DAMGO from human mu opioid receptor expressed in CHO cells after 60 mins by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 2.20nMAssay Description:Displacement of [3H]U-69593 from human opioid kappa receptor expressed in CHO cellsMore data for this Ligand-Target Pair
Affinity DataKi: 3nMAssay Description:Displacement of [3H]U69,593 from human kappa opioid receptor expressed in CHO cells after 60 mins by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 14.3nMAssay Description:Briefly, the receptor binding affinity of the nalbuphine and PEG-nalbuphine conjugates was measured using radioligand binding assays in CHO cells tha...More data for this Ligand-Target Pair
Affinity DataKi: 14.3nMAssay Description:Specific binding is determined by subtraction of the cpm bound in the presence of 50-100� excess of cold ligand. Binding data assays were analyzed us...More data for this Ligand-Target Pair
Affinity DataKi: 25.9nMAssay Description:Briefly, the receptor binding affinity of the nalbuphine and PEG-nalbuphine conjugates was measured using radioligand binding assays in CHO cells tha...More data for this Ligand-Target Pair
Affinity DataKi: 29.9nMAssay Description:Specific binding is determined by subtraction of the cpm bound in the presence of 50-100� excess of cold ligand. Binding data assays were analyzed us...More data for this Ligand-Target Pair
Affinity DataKi: 240nMAssay Description:Displacement of [3H]naltrindole from human opioid delta receptor expressed in CHO cellsMore data for this Ligand-Target Pair
Affinity DataKi: 319nMAssay Description:Specific binding is determined by subtraction of the cpm bound in the presence of 50-100� excess of cold ligand. Binding data assays were analyzed us...More data for this Ligand-Target Pair
Affinity DataKi: 319nMAssay Description:Briefly, the receptor binding affinity of the nalbuphine and PEG-nalbuphine conjugates was measured using radioligand binding assays in CHO cells tha...More data for this Ligand-Target Pair
Affinity DataKi: 580nMAssay Description:Displacement of [3H]naltrindole from human delta opioid receptor expressed in CHO cells after 3 hrs by scintillation countingMore data for this Ligand-Target Pair
TargetThiosulfate sulfurtransferase(Homo sapiens)
Indiana University School of Medicine
Curated by ChEMBL
Indiana University School of Medicine
Curated by ChEMBL
Affinity DataIC50: >1.00E+5nMAssay Description:Inhibition of native rhodanese (unknown origin) assessed as reduction in rhodanese enzyme activity after 45 mins by Fe(SCN)3 dye based spectrometric ...More data for this Ligand-Target Pair
Affinity DataIC50: >2.50E+5nMAssay Description:Inhibition of Escherichia coli GroEL expressed in Escherichia coli DH5alpha/Escherichia coli GroES expressed in Escherichia coli BL21 (DE3) assessed ...More data for this Ligand-Target Pair
Affinity DataIC50: >2.50E+5nMAssay Description:Inhibition of ATPase activity of Escherichia coli GroEL expressed in Escherichia coliDH5alpha incubated for 60 mins using ATP by spectrometric analys...More data for this Ligand-Target Pair
Affinity DataIC50: >1.00E+5nMAssay Description:Inhibition of Escherichia coli GroEL expressed in Escherichia coliDH5alpha/Escherichia coli GroES expressed in Escherichia coli BL21 (DE3) assessed a...More data for this Ligand-Target Pair
Target60 kDa heat shock protein, mitochondrial(Homo sapiens)
Indiana University School of Medicine
Curated by ChEMBL
Indiana University School of Medicine
Curated by ChEMBL
Affinity DataIC50: >1.00E+5nMAssay Description:Inhibition of human N-terminal octa-His-tagged HSP60 expressed in Escherichia coli Rosetta(DE3) pLysS/human HSP10 expressed in Escherichia coli Roset...More data for this Ligand-Target Pair
Affinity DataIC50: 87nMAssay Description:The Ki (binding affinity) for μ opioid receptors was determined using a competitive displacement assay as previously described in Neumeyer (Jour...More data for this Ligand-Target Pair
Affinity DataIC50: 87nMAssay Description:The Ki (binding affinity) for opioid receptors was determined using a competitive displacement assay as previously described in Neumeyer (Journal of ...More data for this Ligand-Target Pair
Affinity DataEC50: 34nMAssay Description:The Ki (binding affinity) for opioid receptors was determined using a competitive displacement assay as previously described in Neumeyer (Journal of ...More data for this Ligand-Target Pair
Affinity DataIC50: 88nMAssay Description:The Ki (binding affinity) for μ opioid receptors was determined using a competitive displacement assay as previously described in Neumeyer (Jour...More data for this Ligand-Target Pair
Affinity DataEC50: 21nMAssay Description:The EC50 and Imax for μ opioid receptors was determined using a [35S]GTPγS binding assay. This assay measures the functional properties of ...More data for this Ligand-Target Pair
Affinity DataIC50: 87nMAssay Description:The Ki (binding affinity) for opioid receptors was determined using a competitive displacement assay as previously described in Neumeyer (Journal of ...More data for this Ligand-Target Pair
Affinity DataEC50: 34nMAssay Description:The Ki (binding affinity) for opioid receptors was determined using a competitive displacement assay as previously described in Neumeyer (Journal of ...More data for this Ligand-Target Pair
Affinity DataEC50: 46nMAssay Description:Agonist activity at human mu opioid receptor expressed in CHO cells assessed as stimulation of [35S]GTPgammaS bindingMore data for this Ligand-Target Pair
Affinity DataIC50: 1.21E+5nMAssay Description:The DAAO enzymatic activity assay was modified according to the report of Oguri et al (Oguri, S., Screening of d-amino acid oxidase inhibitor by a ne...More data for this Ligand-Target Pair
Affinity DataEC50: 27nMAssay Description:Agonist activity at human opioid kappa receptor expressed in CHO cells assessed as stimulation of [35S]GTPgammaS bindingMore data for this Ligand-Target Pair
Affinity DataIC50: 83nMAssay Description:Binding affinity against opioid receptor kappa 1 using [3H]- U-69,593 radioligandMore data for this Ligand-Target Pair
Affinity DataIC50: 1nMAssay Description:Binding affinity against mu-opiate receptor (human) using [3H]DAMGO radioligandMore data for this Ligand-Target Pair
Affinity DataEC50: 250nMAssay Description:Agonist activity at human KOPR expressed in U2OS cells assessed as beta-arrestin2 recruitment after 90 mins by DiscoveRx PathHunter assayMore data for this Ligand-Target Pair
Affinity DataEC50: 65nMAssay Description:Agonist activity at human KOPR expressed in CHO cells assessed as [35S]GTPgammaS binding after 60 mins by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataIC50: 96nMAssay Description:Antagonist activity against human mu opioid receptor expressed in CHO cells assessed as inhibition of DAMGO-stimulated [35S]GTPgammaS bindingMore data for this Ligand-Target Pair
Affinity DataEC50: 56nMAssay Description:Agonist activity at human kappa opioid receptor expressed in CHO cells assessed as stimulation of [35S]GTPgammaS bindingMore data for this Ligand-Target Pair
Affinity DataIC50: 353nMAssay Description:Binding affinity against delta-opiate receptor (human) using [3H]-DPDPE radioligandMore data for this Ligand-Target Pair
Affinity DataIC50: 110nMAssay Description:Agonist activity at human opioid gamma receptor expressed in CHO cells assessed as inhibition of DAGO-stimulated [35S]GTPgammaS bindingMore data for this Ligand-Target Pair
Affinity DataIC50: 752nMpH: 7.4 T: 2°CAssay Description:Test compound and/or vehicle was preincubated with the cell membranes and 3 uM GDP in modified HEPES buffer (pH 7.4) for 20 minutes, followed by addi...More data for this Ligand-Target Pair
Affinity DataEC50: 25.1nMpH: 7.4 T: 2°CAssay Description:Test compound and/or vehicle was preincubated with the cell membranes and 3 uM GDP in modified HEPES buffer (pH 7.4) for 20 minutes, followed by addi...More data for this Ligand-Target Pair
Affinity DataEC50: 14nMAssay Description:Agonist activity at human opioid gamma receptor expressed in CHO cells assessed as stimulation of [35S]GTP-gamma-S bindingMore data for this Ligand-Target Pair