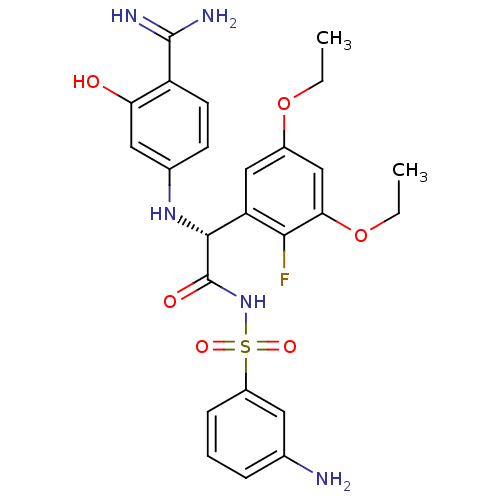
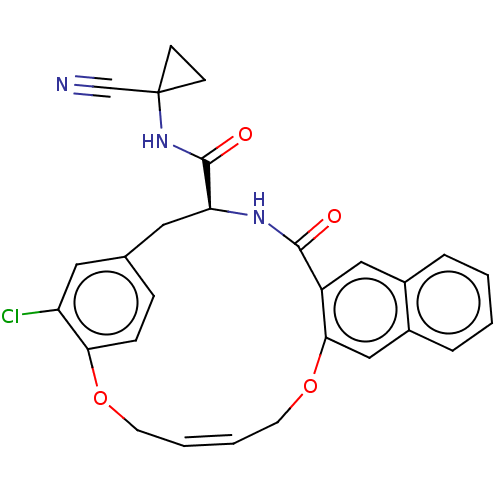
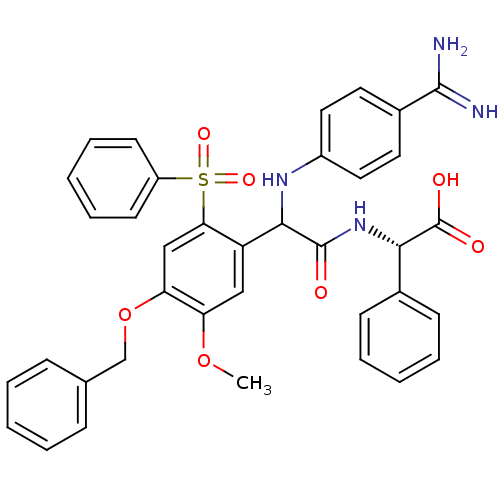
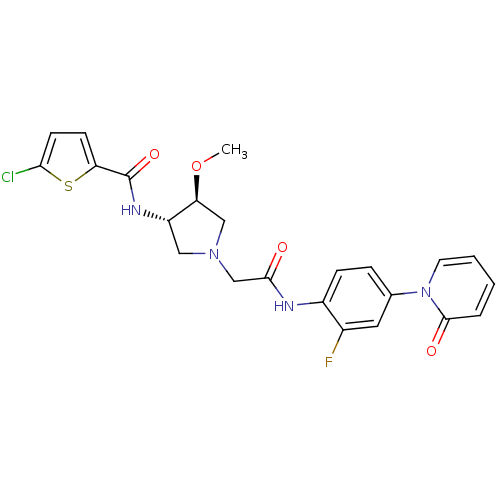
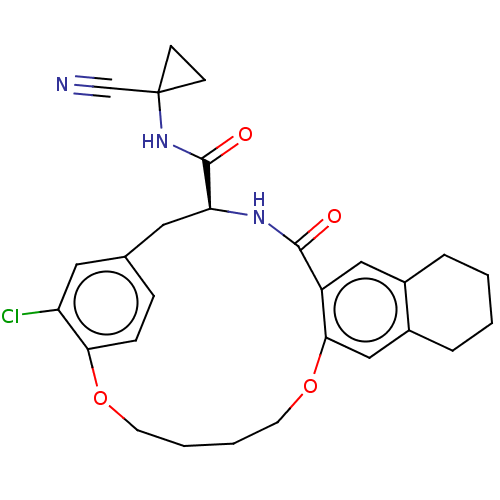
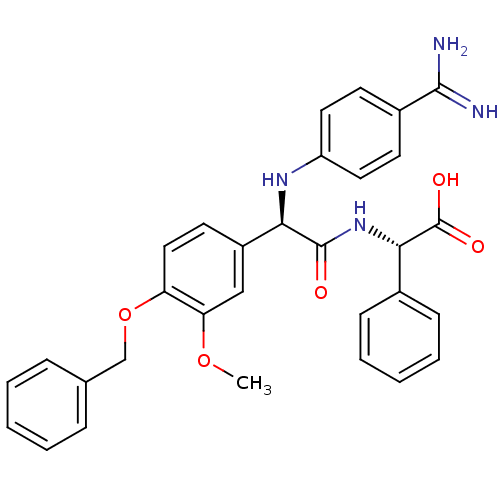
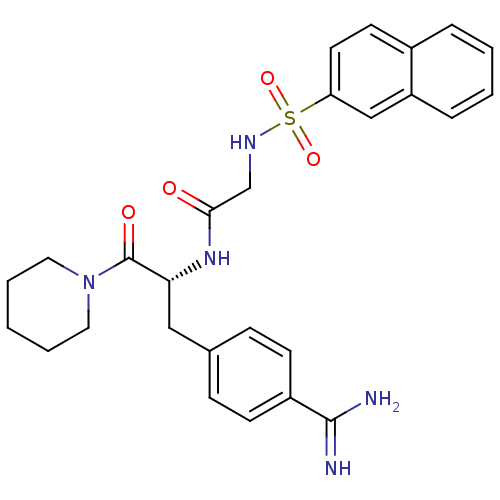
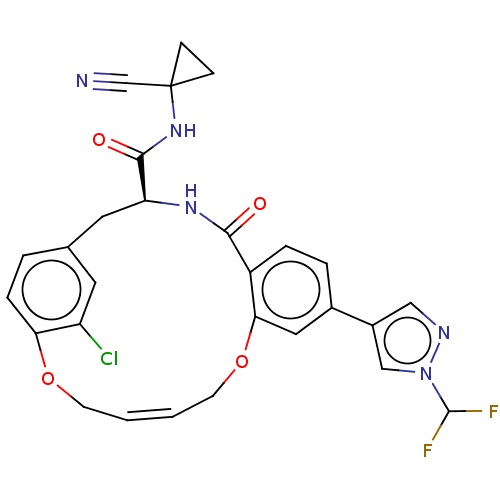
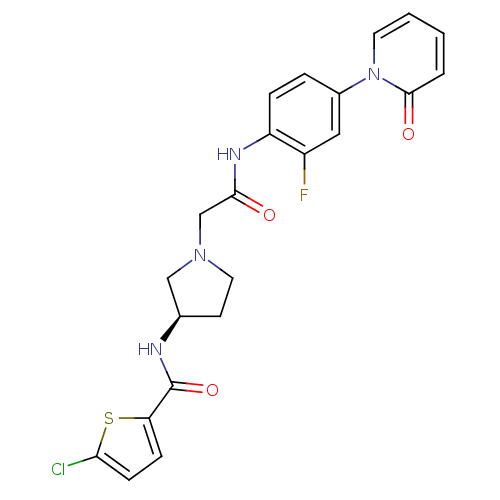
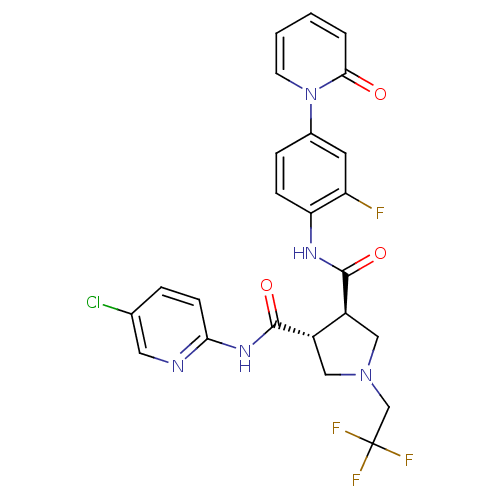
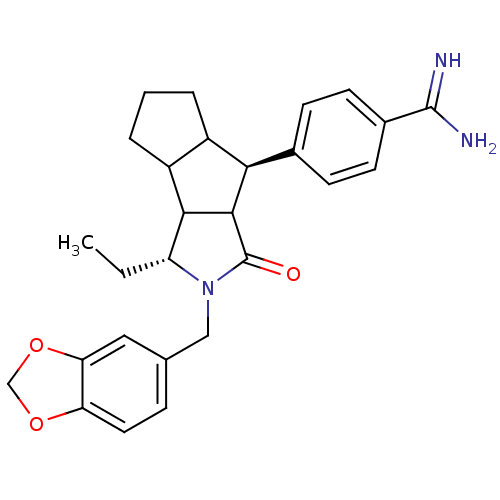
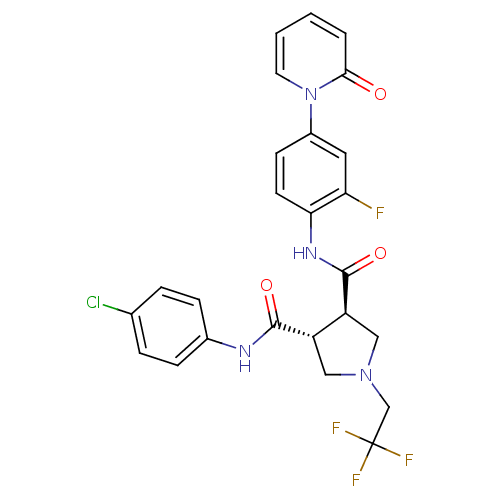
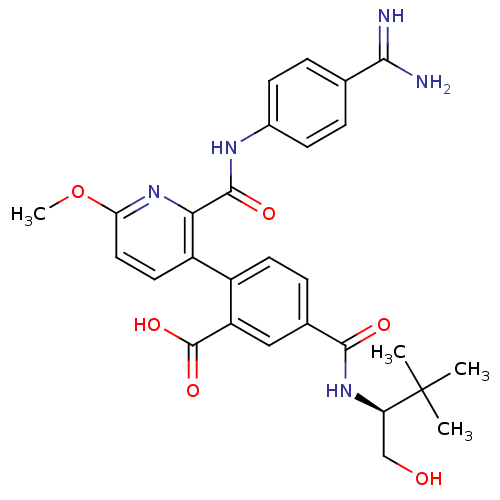
Affinity DataKi: 0.270nMAssay Description:In vitro binding affinity by measuring the inhibition of human thrombinMore data for this Ligand-Target Pair
Affinity DataKi: 0.350nM ΔG°: -53.4kJ/molepH: 7.8 T: 2°CAssay Description:The enzyme reactions were initiated by the addition of substrate, and the color developed from the release of p-nitroanilide from each chromogenic su...More data for this Ligand-Target Pair
Affinity DataKi: 0.680nMAssay Description:In vitro binding affinity by measuring the inhibition of human thrombinMore data for this Ligand-Target Pair
Affinity DataKi: 0.710nMAssay Description:In vitro binding affinity by measuring the inhibition of human thrombinMore data for this Ligand-Target Pair
Affinity DataKi: 0.860nMAssay Description:In vitro binding affinity by measuring the inhibition of human thrombinMore data for this Ligand-Target Pair
Affinity DataKi: 1nMAssay Description:Inhibition of Trypanosoma brucei rhodesiense rhodesain using Cbz-Phe-Arg-AMC as substrate by fluorimetric methodMore data for this Ligand-Target Pair
Affinity DataKi: 1.20nMAssay Description:Inhibition of human CatL using Cbz-Phe-Arg-AMC as substrate measured over 30 mins by fluorimetric methodMore data for this Ligand-Target Pair
Affinity DataKi: 2nM ΔG°: -49.2kJ/molepH: 7.8 T: 2°CAssay Description:The enzyme reactions were initiated by the addition of substrate, and the color developed from the release of p-nitroanilide from each chromogenic su...More data for this Ligand-Target Pair
Affinity DataKi: 2nM ΔG°: -49.2kJ/molepH: 7.8 T: 2°CAssay Description:The enzyme reactions were initiated by the addition of substrate, and the color developed from the release of p-nitroanilide from each chromogenic su...More data for this Ligand-Target Pair
Affinity DataKi: 2nMAssay Description:Inhibition of Trypanosoma brucei rhodesiense rhodesain using Cbz-Phe-Arg-AMC as substrate by fluorimetric methodMore data for this Ligand-Target Pair
Affinity DataKi: 2.30nMAssay Description:Inhibition of human CatL using Cbz-Phe-Arg-AMC as substrate measured over 30 mins by fluorimetric methodMore data for this Ligand-Target Pair
Affinity DataKi: 2.40nMAssay Description:Inhibition of human CatL using Cbz-Phe-Arg-AMC as substrate measured over 30 mins by fluorimetric methodMore data for this Ligand-Target Pair
Affinity DataKi: 2.5nMAssay Description:Inhibition of human CatL using Cbz-Phe-Arg-AMC as substrate measured over 30 mins by fluorimetric methodMore data for this Ligand-Target Pair
Affinity DataKi: 2.5nMAssay Description:Inhibition of Trypanosoma brucei rhodesiense rhodesain using Cbz-Phe-Arg-AMC as substrate by fluorimetric methodMore data for this Ligand-Target Pair
Affinity DataKi: 3nMAssay Description:Binding affinity to factor 10aMore data for this Ligand-Target Pair
Affinity DataKi: 3nMAssay Description:Inhibition of human CatL using Cbz-Phe-Arg-AMC as substrate measured over 30 mins by fluorimetric methodMore data for this Ligand-Target Pair
Affinity DataKi: 3nMAssay Description:Inhibition of Trypanosoma brucei rhodesiense rhodesain using Cbz-Phe-Arg-AMC as substrate by fluorimetric methodMore data for this Ligand-Target Pair
Affinity DataKi: 3.10nMAssay Description:In vitro binding affinity by measuring the inhibition of human thrombinMore data for this Ligand-Target Pair
Affinity DataKi: 3.30nMAssay Description:Binding affinity to rabbit factor 10aMore data for this Ligand-Target Pair
Affinity DataKi: 3.70nMAssay Description:Inhibition of human CatL using Cbz-Phe-Arg-AMC as substrate measured over 30 mins by fluorimetric methodMore data for this Ligand-Target Pair
Affinity DataKi: 4nM ΔG°: -47.5kJ/molepH: 7.8 T: 2°CAssay Description:The enzyme reactions were initiated by the addition of substrate, and the color developed from the release of p-nitroanilide from each chromogenic su...More data for this Ligand-Target Pair
Affinity DataKi: 4nM ΔG°: -47.5kJ/molepH: 7.8 T: 2°CAssay Description:The enzyme reactions were initiated by the addition of substrate, and the color developed from the release of p-nitroanilide from each chromogenic su...More data for this Ligand-Target Pair
Affinity DataKi: 4nMAssay Description:In vitro binding affinity by measuring the inhibition of human thrombinMore data for this Ligand-Target Pair
Affinity DataKi: 4.60nMAssay Description:Inhibition of human CatL using Cbz-Phe-Arg-AMC as substrate measured over 30 mins by fluorimetric methodMore data for this Ligand-Target Pair
Affinity DataKi: 4.80nMAssay Description:In vitro binding affinity by measuring the inhibition of human thrombinMore data for this Ligand-Target Pair
Affinity DataKi: 4.80nMAssay Description:Agonist activity at human SP/Myc epitope-tagged muscarinic M1 receptor expressed in HEK293T cells assessed as RLuc8-fused Galphaq activation after 2 ...More data for this Ligand-Target Pair
Affinity DataKi: 4.80nMAssay Description:Inhibition of Trypanosoma brucei rhodesiense rhodesain using Cbz-Phe-Arg-AMC as substrate by fluorimetric methodMore data for this Ligand-Target Pair
Affinity DataKi: 5.20nMAssay Description:Inhibition of Trypanosoma brucei rhodesiense rhodesain using Cbz-Phe-Arg-AMC as substrate by fluorimetric methodMore data for this Ligand-Target Pair
Affinity DataKi: 5.40nMAssay Description:Inhibition of Trypanosoma brucei rhodesiense rhodesain using Cbz-Phe-Arg-AMC as substrate by fluorimetric methodMore data for this Ligand-Target Pair
Affinity DataKi: 5.60nMAssay Description:Inhibition of human CatL using Cbz-Phe-Arg-AMC as substrate measured over 30 mins by fluorimetric methodMore data for this Ligand-Target Pair
Affinity DataKi: 5.80nMAssay Description:Inhibition of human CatL using Cbz-Phe-Arg-AMC as substrate measured over 30 mins by fluorimetric methodMore data for this Ligand-Target Pair
Affinity DataKi: 6nMAssay Description:Binding affinity to factor 10aMore data for this Ligand-Target Pair
Affinity DataKi: 6nMAssay Description:Binding affinity to factor 10aMore data for this Ligand-Target Pair
Affinity DataKi: 6.10nMAssay Description:Inhibition of Trypanosoma brucei rhodesiense rhodesain using Cbz-Phe-Arg-AMC as substrate by fluorimetric methodMore data for this Ligand-Target Pair
Affinity DataKi: 6.70nMAssay Description:In vitro binding affinity by measuring the inhibition of human thrombinMore data for this Ligand-Target Pair
Affinity DataKi: 7nM ΔG°: -46.1kJ/molepH: 7.8 T: 2°CAssay Description:The enzyme reactions were initiated by the addition of substrate, and the color developed from the release of p-nitroanilide from each chromogenic su...More data for this Ligand-Target Pair
Affinity DataKi: 7nM ΔG°: -46.1kJ/molepH: 7.8 T: 2°CAssay Description:The enzyme reactions were initiated by the addition of substrate, and the color developed from the release of p-nitroanilide from each chromogenic su...More data for this Ligand-Target Pair
Affinity DataKi: 7.30nMAssay Description:Inhibition of human CatL using Cbz-Phe-Arg-AMC as substrate measured over 30 mins by fluorimetric methodMore data for this Ligand-Target Pair
Affinity DataKi: 7.40nMAssay Description:Inhibition of human CatL using Cbz-Phe-Arg-AMC as substrate measured over 30 mins by fluorimetric methodMore data for this Ligand-Target Pair
Affinity DataKi: 8nMAssay Description:Binding affinity to factor 10aMore data for this Ligand-Target Pair
Affinity DataKi: 8nMAssay Description:Binding affinity to factor 10aMore data for this Ligand-Target Pair
Affinity DataKi: 8nMAssay Description:Binding affinity to factor 10aMore data for this Ligand-Target Pair
Affinity DataKi: 8.10nMAssay Description:Enzyme inhibition assay using thrombin, a trypsin-like serine protease.More data for this Ligand-Target Pair
Affinity DataKi: 8.70nMAssay Description:Inhibition of Trypanosoma brucei rhodesiense rhodesain using Cbz-Phe-Arg-AMC as substrate by fluorimetric methodMore data for this Ligand-Target Pair
Affinity DataKi: 9nMAssay Description:Binding affinity to factor 10aMore data for this Ligand-Target Pair
Affinity DataKi: 9nMAssay Description:In vitro binding affinity by measuring the inhibition of human thrombinMore data for this Ligand-Target Pair
Affinity DataKi: 10nM ΔG°: -45.2kJ/molepH: 7.8 T: 2°CAssay Description:The enzyme reactions were initiated by the addition of substrate, and the color developed from the release of p-nitroanilide from each chromogenic su...More data for this Ligand-Target Pair
Affinity DataKi: 10nMAssay Description:Enzyme inhibition assay using thrombin, a trypsin-like serine protease.More data for this Ligand-Target Pair
Affinity DataKi: 10nMAssay Description:Binding affinity to factor 10aMore data for this Ligand-Target Pair
Affinity DataKi: 10nMAssay Description:Binding affinity to factor 10aMore data for this Ligand-Target Pair


 3D Structure (crystal)
3D Structure (crystal)