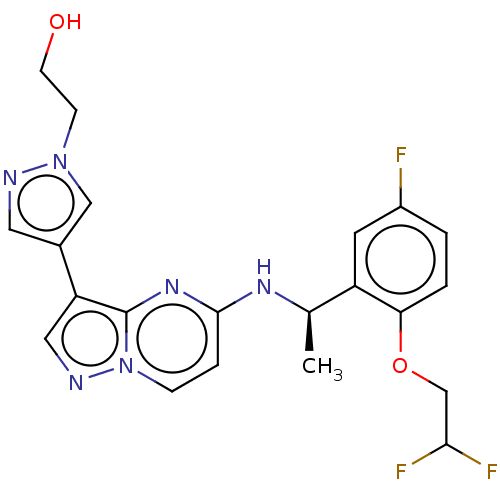
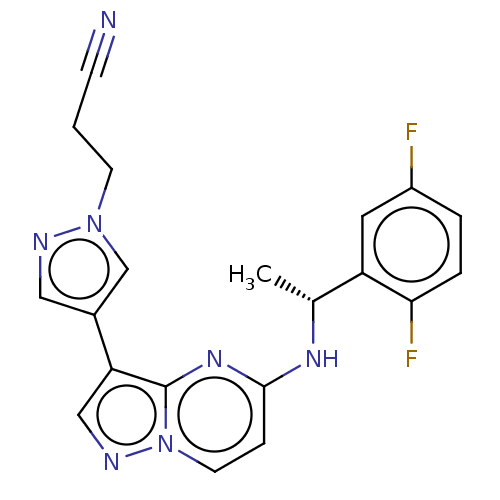
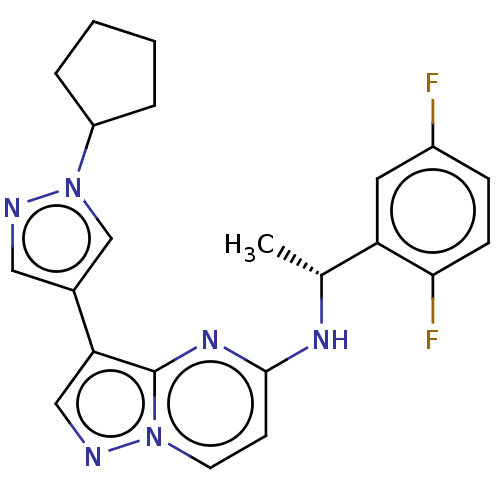
Affinity DataKi: 4.90nMAssay Description:To evaluate the potency of synthesized compounds against Mpro, the proteolytic activity of 50 nM Mpro -His and Mpro was first measured in the presenc...More data for this Ligand-Target Pair
Affinity DataKi: 5.60nMAssay Description:To evaluate the potency of synthesized compounds against Mpro, the proteolytic activity of 50 nM Mpro -His and Mpro was first measured in the presenc...More data for this Ligand-Target Pair
Affinity DataKi: 8.30nMAssay Description:Binding affinity to DCN1 (unknown origin) assessed as inhibitory constantMore data for this Ligand-Target Pair
Affinity DataKi: 12nMAssay Description:To evaluate the potency of synthesized compounds against Mpro, the proteolytic activity of 50 nM Mpro -His and Mpro was first measured in the presenc...More data for this Ligand-Target Pair
Affinity DataKi: 14nMAssay Description:Binding affinity to DCN1 (unknown origin) assessed as inhibitory constantMore data for this Ligand-Target Pair
Affinity DataKi: 14nMAssay Description:To evaluate the potency of synthesized compounds against Mpro, the proteolytic activity of 50 nM Mpro -His and Mpro was first measured in the presenc...More data for this Ligand-Target Pair
Affinity DataKi: 20nMAssay Description:To evaluate the potency of synthesized compounds against Mpro, the proteolytic activity of 50 nM Mpro -His and Mpro was first measured in the presenc...More data for this Ligand-Target Pair
Affinity DataKi: 24nMAssay Description:To evaluate the potency of synthesized compounds against Mpro, the proteolytic activity of 50 nM Mpro -His and Mpro was first measured in the presenc...More data for this Ligand-Target Pair
Affinity DataKi: 29nMAssay Description:To evaluate the potency of synthesized compounds against Mpro, the proteolytic activity of 50 nM Mpro -His and Mpro was first measured in the presenc...More data for this Ligand-Target Pair
Affinity DataKi: 37nMAssay Description:To evaluate the potency of synthesized compounds against Mpro, the proteolytic activity of 50 nM Mpro -His and Mpro was first measured in the presenc...More data for this Ligand-Target Pair
Affinity DataKi: 42nMAssay Description:To evaluate the potency of synthesized compounds against Mpro, the proteolytic activity of 50 nM Mpro -His and Mpro was first measured in the presenc...More data for this Ligand-Target Pair
Affinity DataKi: 45nMAssay Description:To evaluate the potency of synthesized compounds against Mpro, the proteolytic activity of 50 nM Mpro -His and Mpro was first measured in the presenc...More data for this Ligand-Target Pair
Affinity DataKi: 46nMAssay Description:To evaluate the potency of synthesized compounds against Mpro, the proteolytic activity of 50 nM Mpro -His and Mpro was first measured in the presenc...More data for this Ligand-Target Pair
Affinity DataKi: 65nMAssay Description:To evaluate the potency of synthesized compounds against Mpro, the proteolytic activity of 50 nM Mpro -His and Mpro was first measured in the presenc...More data for this Ligand-Target Pair
Affinity DataKi: 810nMAssay Description:Binding affinity to DCN4 (unknown origin) assessed as inhibitory constantMore data for this Ligand-Target Pair
Affinity DataKi: 2.14E+3nMAssay Description:Binding affinity to DCN5 (unknown origin) assessed as inhibitory constantMore data for this Ligand-Target Pair
Affinity DataKi: 1.57E+4nMAssay Description:To evaluate the potency of synthesized compounds against Mpro, the proteolytic activity of 50 nM Mpro -His and Mpro was first measured in the presenc...More data for this Ligand-Target Pair
Affinity DataKi: 1.80E+4nM ΔG°: -28.2kJ/molepH: 7.2 T: 2°CAssay Description:The Km and Kappm for the amino acid substrate were first calculated from Lineweaver-Burk plots. The Ki values were calculated from the Kappm vs [I] p...More data for this Ligand-Target Pair
Affinity DataKi: 1.88E+4nMAssay Description:Binding affinity to DCN2 (unknown origin) assessed as inhibitory constantMore data for this Ligand-Target Pair
Affinity DataKi: 2.12E+4nMAssay Description:To evaluate the potency of synthesized compounds against Mpro, the proteolytic activity of 50 nM Mpro -His and Mpro was first measured in the presenc...More data for this Ligand-Target Pair
Affinity DataKi: 2.24E+4nMAssay Description:Binding affinity to DCN4 (unknown origin) assessed as inhibitory constantMore data for this Ligand-Target Pair
Affinity DataKi: 2.89E+4nMAssay Description:Binding affinity to DCN5 (unknown origin) assessed as inhibitory constantMore data for this Ligand-Target Pair
Affinity DataKi: 6.62E+4nMAssay Description:Binding affinity to DCN2 (unknown origin) assessed as inhibitory constantMore data for this Ligand-Target Pair
Affinity DataKi: 3.85E+5nMAssay Description:Binding affinity to DCN3 (unknown origin) assessed as inhibitory constantMore data for this Ligand-Target Pair
Affinity DataKi: 5.92E+5nMAssay Description:Binding affinity to DCN3 (unknown origin) assessed as inhibitory constantMore data for this Ligand-Target Pair
Affinity DataKi: 6.50E+5nM ΔG°: -18.9kJ/molepH: 7.2 T: 2°CAssay Description:The Km and Kappm for the amino acid substrate were first calculated from Lineweaver-Burk plots. The Ki values were calculated from the Kappm vs [I] p...More data for this Ligand-Target Pair
Affinity DataKi: 2.60E+6nM ΔG°: -15.3kJ/molepH: 7.2 T: 2°CAssay Description:The Km and Kappm for the amino acid substrate were first calculated from Lineweaver-Burk plots. The Ki values were calculated from the Kappm vs [I] p...More data for this Ligand-Target Pair
Affinity DataKi: 2.80E+6nM ΔG°: -15.2kJ/molepH: 7.2 T: 2°CAssay Description:The Km and Kappm for the amino acid substrate were first calculated from Lineweaver-Burk plots. The Ki values were calculated from the Kappm vs [I] p...More data for this Ligand-Target Pair
Affinity DataKi: 2.90E+6nM ΔG°: -15.1kJ/molepH: 7.2 T: 2°CAssay Description:The Km and Kappm for the amino acid substrate were first calculated from Lineweaver-Burk plots. The Ki values were calculated from the Kappm vs [I] p...More data for this Ligand-Target Pair
TargetHigh affinity nerve growth factor receptor(Homo sapiens (Human))
Central China Normal University
Curated by ChEMBL
Central China Normal University
Curated by ChEMBL
Affinity DataIC50: 0.100nMAssay Description:Inhibition of TRKA F589L mutant (unknown origin) using TK-sub-biotin peptide as substrate incubated for 30 mins followed by treated with ATP and subs...More data for this Ligand-Target Pair
TargetHigh affinity nerve growth factor receptor(Homo sapiens (Human))
Central China Normal University
Curated by ChEMBL
Central China Normal University
Curated by ChEMBL
Affinity DataIC50: 0.100nMAssay Description:Inhibition of TRKA F589L mutant (unknown origin) using TK-sub-biotin peptide as substrate incubated for 30 mins followed by treated with ATP and subs...More data for this Ligand-Target Pair
TargetHigh affinity nerve growth factor receptor(Homo sapiens (Human))
Central China Normal University
Curated by ChEMBL
Central China Normal University
Curated by ChEMBL
Affinity DataIC50: 0.100nMAssay Description:Inhibition of TRKA F589L mutant (unknown origin) using TK-sub-biotin peptide as substrate incubated for 30 mins followed by treated with ATP and subs...More data for this Ligand-Target Pair
TargetHigh affinity nerve growth factor receptor(Homo sapiens (Human))
Central China Normal University
Curated by ChEMBL
Central China Normal University
Curated by ChEMBL
Affinity DataIC50: 0.200nMAssay Description:Inhibition of TRKA GS95R mutant (unknown origin) preincubated for 30 mins followed by biotinylated TK-peptide substrate addition and measured after 4...More data for this Ligand-Target Pair
TargetHigh affinity nerve growth factor receptor(Homo sapiens (Human))
Central China Normal University
Curated by ChEMBL
Central China Normal University
Curated by ChEMBL
Affinity DataIC50: 0.200nMAssay Description:Inhibition of recombinant human His-tagged TRKA cytoplasmic domain (441 to 796 residues) expressed in insect cells using TK-sub biotin peptide substr...More data for this Ligand-Target Pair
TargetHigh affinity nerve growth factor receptor(Homo sapiens (Human))
Central China Normal University
Curated by ChEMBL
Central China Normal University
Curated by ChEMBL
Affinity DataIC50: 0.200nMAssay Description:Inhibition of wild type TRKA (unknown origin) using TK-sub-biotin peptide as substrate incubated for 30 mins followed by treated with ATP and substra...More data for this Ligand-Target Pair
TargetHigh affinity nerve growth factor receptor(Homo sapiens (Human))
Central China Normal University
Curated by ChEMBL
Central China Normal University
Curated by ChEMBL
Affinity DataIC50: 0.200nMAssay Description:Inhibition of TRKA GS95R mutant (unknown origin) preincubated for 30 mins followed by biotinylated TK-peptide substrate addition and measured after 4...More data for this Ligand-Target Pair
TargetHigh affinity nerve growth factor receptor(Homo sapiens (Human))
Central China Normal University
Curated by ChEMBL
Central China Normal University
Curated by ChEMBL
Affinity DataIC50: 0.200nMAssay Description:Inhibition of wild type TRKA (unknown origin) using TK-sub-biotin peptide as substrate incubated for 30 mins followed by treated with ATP and substra...More data for this Ligand-Target Pair
TargetHigh affinity nerve growth factor receptor(Homo sapiens (Human))
Central China Normal University
Curated by ChEMBL
Central China Normal University
Curated by ChEMBL
Affinity DataIC50: 0.200nMAssay Description:Inhibition of wild type TRKA (unknown origin) using TK-sub-biotin peptide as substrate incubated for 30 mins followed by treated with ATP and substra...More data for this Ligand-Target Pair
TargetHigh affinity nerve growth factor receptor(Homo sapiens (Human))
Central China Normal University
Curated by ChEMBL
Central China Normal University
Curated by ChEMBL
Affinity DataIC50: 0.200nMAssay Description:Inhibition of wild type TRKA (unknown origin) using TK-sub-biotin peptide as substrate incubated for 30 mins followed by treated with ATP and substra...More data for this Ligand-Target Pair
TargetHigh affinity nerve growth factor receptor(Homo sapiens (Human))
Central China Normal University
Curated by ChEMBL
Central China Normal University
Curated by ChEMBL
Affinity DataIC50: 0.200nMAssay Description:Inhibition of wild type TRKA (unknown origin) using TK-sub-biotin peptide as substrate incubated for 30 mins followed by treated with ATP and substra...More data for this Ligand-Target Pair
TargetHigh affinity nerve growth factor receptor(Homo sapiens (Human))
Central China Normal University
Curated by ChEMBL
Central China Normal University
Curated by ChEMBL
Affinity DataIC50: 0.200nMAssay Description:Inhibition of wild type TRKA (unknown origin) using TK-sub-biotin peptide as substrate incubated for 30 mins followed by treated with ATP and substra...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase 2/G1/S-specific cyclin-E1(Homo sapiens (Human))
Incyte
Curated by ChEMBL
Incyte
Curated by ChEMBL
Affinity DataIC50: 0.25nMAssay Description:Inhibition of recombinant human N-terminal GST-tagged full-length CDK2/Flag-tagged cyclinE1 expressed in baculovirus expression system using eIF4E-bi...More data for this Ligand-Target Pair
Affinity DataIC50: 0.25nMAssay Description:Inhibition of GST-tagged DCN1 (unknown origin) using AcUBE2M1-21 incubated for 30 mins by HTRF assayMore data for this Ligand-Target Pair
TargetHigh affinity nerve growth factor receptor(Homo sapiens (Human))
Central China Normal University
Curated by ChEMBL
Central China Normal University
Curated by ChEMBL
Affinity DataIC50: 0.300nMAssay Description:Inhibition of wild type TRKA (unknown origin) using TK-sub-biotin peptide as substrate incubated for 30 mins followed by treated with ATP and substra...More data for this Ligand-Target Pair
TargetHigh affinity nerve growth factor receptor(Homo sapiens (Human))
Central China Normal University
Curated by ChEMBL
Central China Normal University
Curated by ChEMBL
Affinity DataIC50: 0.300nMAssay Description:Inhibition of wild type TRKA (unknown origin) using TK-sub-biotin peptide as substrate incubated for 30 mins followed by treated with ATP and substra...More data for this Ligand-Target Pair
TargetHigh affinity nerve growth factor receptor(Homo sapiens (Human))
Central China Normal University
Curated by ChEMBL
Central China Normal University
Curated by ChEMBL
Affinity DataIC50: 0.300nMAssay Description:Inhibition of TRKA F589L mutant (unknown origin) using TK-sub-biotin peptide as substrate incubated for 30 mins followed by treated with ATP and subs...More data for this Ligand-Target Pair
TargetHigh affinity nerve growth factor receptor(Homo sapiens (Human))
Central China Normal University
Curated by ChEMBL
Central China Normal University
Curated by ChEMBL
Affinity DataIC50: 0.300nMAssay Description:Inhibition of wild type TRKA (unknown origin) using TK-sub-biotin peptide as substrate incubated for 30 mins followed by treated with ATP and substra...More data for this Ligand-Target Pair
TargetHigh affinity nerve growth factor receptor(Homo sapiens (Human))
Central China Normal University
Curated by ChEMBL
Central China Normal University
Curated by ChEMBL
Affinity DataIC50: 0.300nMAssay Description:Inhibition of wild type TRKA (unknown origin) using TK-sub-biotin peptide as substrate incubated for 30 mins followed by treated with ATP and substra...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase 2/G1/S-specific cyclin-E1(Homo sapiens (Human))
Incyte
Curated by ChEMBL
Incyte
Curated by ChEMBL
Affinity DataIC50: 0.320nMAssay Description:Inhibition of recombinant human N-terminal GST-tagged full-length CDK2/Flag-tagged cyclinE1 expressed in baculovirus expression system using eIF4E-bi...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase 2/G1/S-specific cyclin-E1(Homo sapiens (Human))
Incyte
Curated by ChEMBL
Incyte
Curated by ChEMBL
Affinity DataIC50: 0.330nMAssay Description:Inhibition of recombinant human N-terminal GST-tagged full-length CDK2/Flag-tagged cyclinE1 expressed in baculovirus expression system using eIF4E-bi...More data for this Ligand-Target Pair



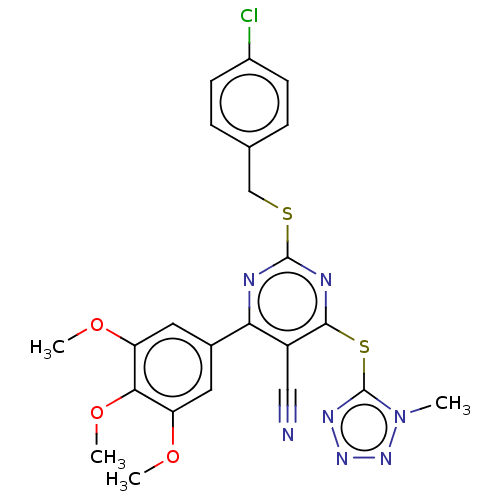





 3D Structure (crystal)
3D Structure (crystal)