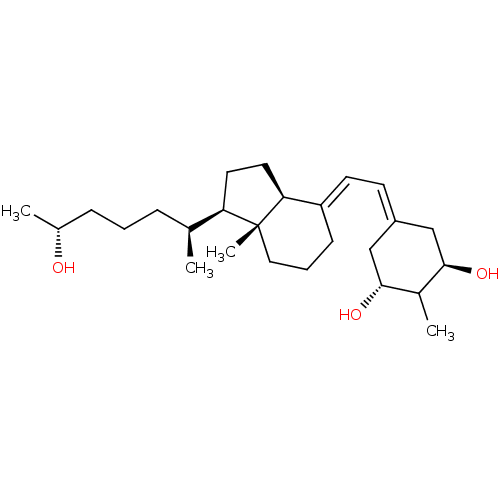
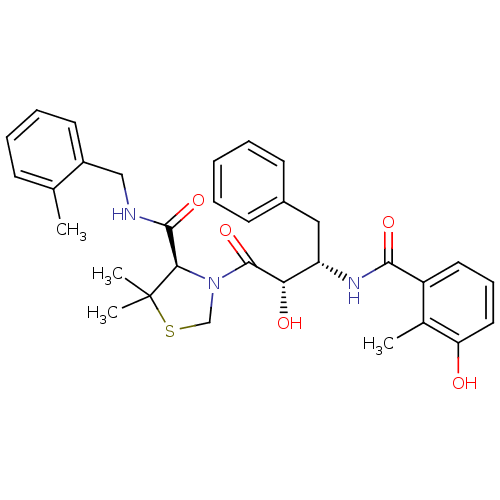
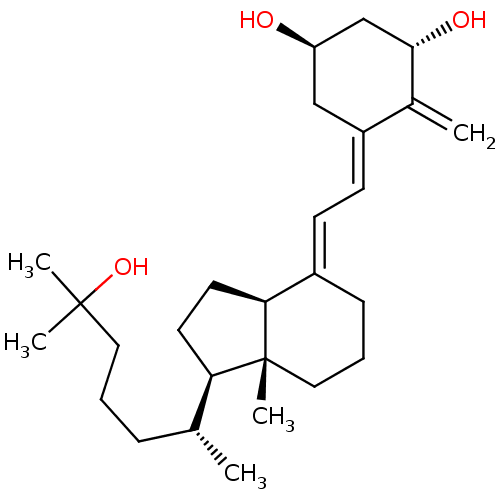
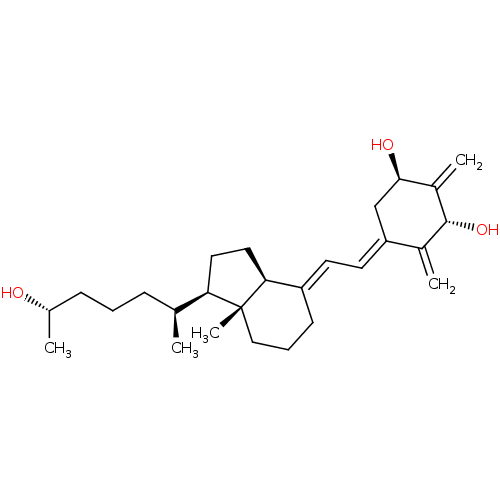
Affinity DataKi: 0.00700nMAssay Description:Activity at rat recombinant full length VDR expressed in rat ROS 17/2.8 cells transfected with 24-hydroxylase gene promoter assessed as transcription...More data for this Ligand-Target Pair
Affinity DataKi: 0.0100nMAssay Description:Activity at rat recombinant full length VDR expressed in rat ROS 17/2.8 cells transfected with 24-hydroxylase gene promoter assessed as transcription...More data for this Ligand-Target Pair
Affinity DataKi: 0.0200nMAssay Description:Displacement of [3H]1alpha,25-(OH)2D3 from full-length recombinant rat VDR by scintillation counterMore data for this Ligand-Target Pair
Affinity DataKi: 0.0200nMAssay Description:Displacement of radiolabeled 1alpha,25-(OH)2D3 from rat recombinant full length VDRMore data for this Ligand-Target Pair
Affinity DataKi: 0.0200nMAssay Description:Displacement of [3H]-1alpha,25(OH)2D3 from recombinant rat VDR after overnight incubation by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 0.0200nMAssay Description:Activity at rat recombinant full length VDR expressed in rat ROS 17/2.8 cells transfected with 24-hydroxylase gene promoter assessed as transcription...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [501-599,D531N,A572V](Human immunodeficiency virus type 1)
University of Florida College of Medicine
University of Florida College of Medicine
Affinity DataKi: <0.0210nM ΔG°: <-63.4kJ/molepH: 4.7 T: 2°CAssay Description:The inhibition constant, Ki was determined by monitoring the competitive inhibition of the hydrolysis of the chromogenic substrate.More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [501-599](Human immunodeficiency virus type 1)
University of Florida College of Medicine
University of Florida College of Medicine
Affinity DataKi: 0.0210nM ΔG°: -63.4kJ/molepH: 4.7 T: 2°CAssay Description:The inhibition constant, Ki was determined by monitoring the competitive inhibition of the hydrolysis of the chromogenic substrate.More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [501-599,A572V](Human immunodeficiency virus type 1)
University of Florida College of Medicine
University of Florida College of Medicine
Affinity DataKi: <0.0210nM ΔG°: <-63.4kJ/molepH: 4.7 T: 2°CAssay Description:The inhibition constant, Ki was determined by monitoring the competitive inhibition of the hydrolysis of the chromogenic substrate.More data for this Ligand-Target Pair
Affinity DataKi: 0.0300nMAssay Description:Displacement of [3H]1alpha,25-(OH)2D3 from full-length recombinant rat VDR by scintillation counterMore data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [501-599,M537I,A572V](Human immunodeficiency virus type 1)
University of Florida College of Medicine
University of Florida College of Medicine
Affinity DataKi: 0.0300nM ΔG°: -62.5kJ/molepH: 4.7 T: 2°CAssay Description:The inhibition constant, Ki was determined by monitoring the competitive inhibition of the hydrolysis of the chromogenic substrate.More data for this Ligand-Target Pair
Affinity DataKi: 0.0300nMAssay Description:Displacement of [3H]1alpha,25-(OH)2D3 from full-length recombinant rat VDR by scintillation counterMore data for this Ligand-Target Pair
Affinity DataKi: 0.0300nMAssay Description:Displacement of [3H]1alpha,25-(OH)2D3 from full-length recombinant rat VDR by scintillation counterMore data for this Ligand-Target Pair
Affinity DataKi: 0.0400nMAssay Description:Displacement of [3H]1alpha,25-(OH)2D3 from full-length recombinant rat VDR by scintillation counterMore data for this Ligand-Target Pair
Affinity DataKi: 0.0400nMAssay Description:Displacement of [3H]-1alpha,25(OH)2D3 from recombinant rat VDR after overnight incubation by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 0.0400nMAssay Description:Displacement of [3H]1alpha,25-(OH)2D3 from full-length recombinant rat VDR by scintillation counterMore data for this Ligand-Target Pair
Affinity DataKi: 0.0500nM ΔG°: -58.8kJ/mole IC50: 58nMpH: 7.5 T: 2°CAssay Description:The Ki (binding affinity) for u opioid receptors was determined using a competitive displacement assay as previously described in Neumeyer (Journal o...More data for this Ligand-Target Pair
Affinity DataKi: 0.0500nMAssay Description:The Ki (binding affinity) for μ opioid receptors was determined using a competitive displacement assay as previously described in Neumeyer (Jour...More data for this Ligand-Target Pair
Affinity DataKi: 0.0500nMAssay Description:The Ki (binding affinity) for μ opioid receptors was determined using a competitive displacement assay as previously described in Neumeyer (Jour...More data for this Ligand-Target Pair
Affinity DataKi: 0.0500nMAssay Description:The Ki (binding affinity) for opioid receptors was determined using a competitive displacement assay as previously described in Neumeyer (Journal of ...More data for this Ligand-Target Pair
Affinity DataKi: 0.0500nMAssay Description:The Ki (binding affinity) for opioid receptors was determined using a competitive displacement assay as previously described in Neumeyer (Journal of ...More data for this Ligand-Target Pair
Affinity DataKi: 0.0500nMAssay Description:Displacement of [3H]1alpha,25-(OH)2D3 from rat recombinant full length VDRMore data for this Ligand-Target Pair
Affinity DataKi: 0.0500nMAssay Description:The Ki (binding affinity) for μ opioid receptors was determined using a competitive displacement assay as previously described in Neumeyer (Jour...More data for this Ligand-Target Pair
Affinity DataKi: 0.0500nMAssay Description:The Ki (binding affinity) for μ opioid receptors was determined using a competitive displacement assay as previously described in Neumeyer (Jour...More data for this Ligand-Target Pair
Affinity DataKi: 0.0600nMAssay Description:Displacement of [3H]1alpha,25-(OH)2D3 from rat recombinant full length VDRMore data for this Ligand-Target Pair
Affinity DataKi: 0.0600nMAssay Description:Activity at rat recombinant full length VDR expressed in rat ROS 17/2.8 cells transfected with 24-hydroxylase gene promoter assessed as transcription...More data for this Ligand-Target Pair
Affinity DataKi: 0.0600nMAssay Description:Displacement of [3H]-1alpha,25(OH)2D3 from recombinant rat VDR after overnight incubation by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 0.0610nMAssay Description:The Ki (binding affinity) for μ opioid receptors was determined using a competitive displacement assay as previously described in Neumeyer (Jour...More data for this Ligand-Target Pair
Affinity DataKi: 0.0610nMAssay Description:The Ki (binding affinity) for opioid receptors was determined using a competitive displacement assay as previously described in Neumeyer (Journal of ...More data for this Ligand-Target Pair
Affinity DataKi: 0.0610nMAssay Description:The Ki (binding affinity) for opioid receptors was determined using a competitive displacement assay as previously described in Neumeyer (Journal of ...More data for this Ligand-Target Pair
Affinity DataKi: 0.0610nMAssay Description:The Ki (binding affinity) for opioid receptors was determined using a competitive displacement assay as previously described in Neumeyer (Journal of ...More data for this Ligand-Target Pair
Affinity DataKi: 0.0610nMAssay Description:The Ki (binding affinity) for μ opioid receptors was determined using a competitive displacement assay as previously described in Neumeyer (Jour...More data for this Ligand-Target Pair
Affinity DataKi: 0.0610nMAssay Description:The Ki (binding affinity) for μ opioid receptors was determined using a competitive displacement assay as previously described in Neumeyer (Jour...More data for this Ligand-Target Pair
Affinity DataKi: 0.0610nMAssay Description:The Ki (binding affinity) for μ opioid receptors was determined using a competitive displacement assay as previously described in Neumeyer (Jour...More data for this Ligand-Target Pair
Affinity DataKi: 0.0610nMAssay Description:The Ki (binding affinity) for opioid receptors was determined using a competitive displacement assay as previously described in Neumeyer (Journal of ...More data for this Ligand-Target Pair
Affinity DataKi: 0.0790nMAssay Description:The Ki (binding affinity) for μ opioid receptors was determined using a competitive displacement assay as previously described in Neumeyer (Jour...More data for this Ligand-Target Pair
Affinity DataKi: 0.0790nMAssay Description:The Ki (binding affinity) for opioid receptors was determined using a competitive displacement assay as previously described in Neumeyer (Journal of ...More data for this Ligand-Target Pair
Affinity DataKi: 0.0790nMAssay Description:The Ki (binding affinity) for opioid receptors was determined using a competitive displacement assay as previously described in Neumeyer (Journal of ...More data for this Ligand-Target Pair
Affinity DataKi: 0.0790nMAssay Description:The Ki (binding affinity) for μ opioid receptors was determined using a competitive displacement assay as previously described in Neumeyer (Jour...More data for this Ligand-Target Pair
Affinity DataKi: 0.0800nMAssay Description:The Ki (binding affinity) for μ opioid receptors was determined using a competitive displacement assay as previously described in Neumeyer (Jour...More data for this Ligand-Target Pair
Affinity DataKi: 0.0800nMAssay Description:Activity at rat recombinant full length VDR expressed in rat ROS 17/2.8 cells transfected with 24-hydroxylase gene promoter assessed as transcription...More data for this Ligand-Target Pair
Affinity DataKi: 0.0800nMAssay Description:Displacement of [3H]1alpha,25-(OH)2D3 from rat recombinant full length VDRMore data for this Ligand-Target Pair
Affinity DataKi: 0.0800nMAssay Description:The Ki (binding affinity) for opioid receptors was determined using a competitive displacement assay as previously described in Neumeyer (Journal of ...More data for this Ligand-Target Pair
Affinity DataKi: 0.0800nMAssay Description:The Ki (binding affinity) for μ opioid receptors was determined using a competitive displacement assay as previously described in Neumeyer (Jour...More data for this Ligand-Target Pair
Affinity DataKi: 0.0800nMAssay Description:The Ki (binding affinity) for opioid receptors was determined using a competitive displacement assay as previously described in Neumeyer (Journal of ...More data for this Ligand-Target Pair
Affinity DataKi: 0.0830nM ΔG°: -57.5kJ/mole IC50: 1.70nMpH: 7.5 T: 2°CAssay Description:The Ki (binding affinity) for u opioid receptors was determined using a competitive displacement assay as previously described in Neumeyer (Journal o...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [501-599,Q508K,L534I,L564I,C568A,C596A,D531N](Human immunodeficiency virus type 1)
University of Florida College of Medicine
University of Florida College of Medicine
Affinity DataKi: 0.0830nM ΔG°: -59.9kJ/molepH: 4.7 T: 2°CAssay Description:The inhibition constant, Ki was determined by monitoring the competitive inhibition of the hydrolysis of the chromogenic substrate.More data for this Ligand-Target Pair
Affinity DataKi: 0.0830nMAssay Description:The Ki (binding affinity) for opioid receptors was determined using a competitive displacement assay as previously described in Neumeyer (Journal of ...More data for this Ligand-Target Pair
Affinity DataKi: 0.0830nMAssay Description:The Ki (binding affinity) for μ opioid receptors was determined using a competitive displacement assay as previously described in Neumeyer (Jour...More data for this Ligand-Target Pair
Affinity DataKi: 0.0830nMAssay Description:The Ki (binding affinity) for μ opioid receptors was determined using a competitive displacement assay as previously described in Neumeyer (Jour...More data for this Ligand-Target Pair


 3D Structure (crystal)
3D Structure (crystal)