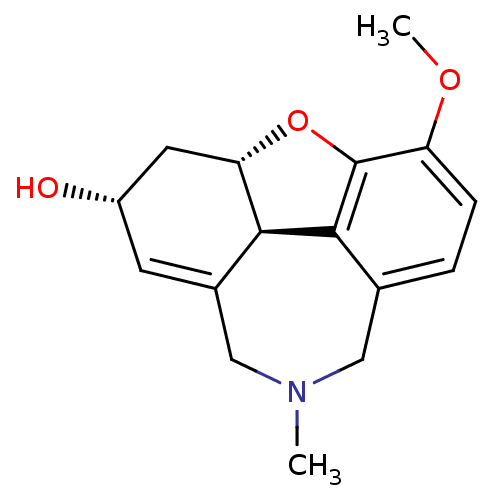
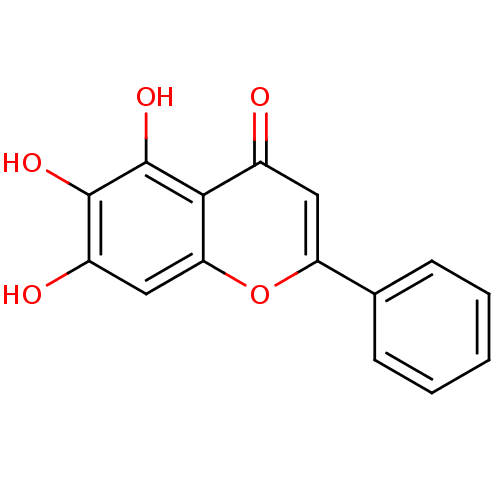
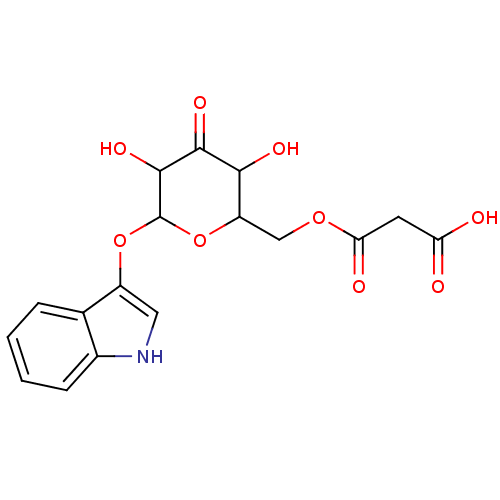
Affinity DataIC50: 8.50E+3nMpH: 8.0 T: 2°CAssay Description:Butyrylcholinesterase activity-inhibiting activities were measured by a slightly modified spectrophotometric method developed by Ellman. DTNB was us...More data for this Ligand-Target Pair
Affinity DataIC50: 1.63E+4nMpH: 8.0 T: 2°CAssay Description:Butyrylcholinesterase activity-inhibiting activities were measured by a slightly modified spectrophotometric method developed by Ellman. DTNB was us...More data for this Ligand-Target Pair
Affinity DataIC50: 1.97E+4nMpH: 8.0 T: 2°CAssay Description:Butyrylcholinesterase activity-inhibiting activities were measured by a slightly modified spectrophotometric method developed by Ellman. DTNB was us...More data for this Ligand-Target Pair
TargetLinoleate 9S-lipoxygenase-4(Glycine max (Soybean) (Glycine hispida))
Pcsir Laboratories Complex
Pcsir Laboratories Complex
Affinity DataIC50: 2.20E+4nMpH: 8.0 T: 2°CAssay Description:LOX inhibiting activity was measured by modifying the spectrophotometric method developed by Tappel. More data for this Ligand-Target Pair
Affinity DataIC50: 2.35E+4nMpH: 8.0 T: 2°CAssay Description:Butyrylcholinesterase activity-inhibiting activities were measured by a slightly modified spectrophotometric method developed by Ellman. DTNB was us...More data for this Ligand-Target Pair
Affinity DataIC50: 2.48E+4nMpH: 8.0 T: 2°CAssay Description:Butyrylcholinesterase activity-inhibiting activities were measured by a slightly modified spectrophotometric method developed by Ellman. DTNB was us...More data for this Ligand-Target Pair
TargetLinoleate 9S-lipoxygenase-4(Glycine max (Soybean) (Glycine hispida))
Pcsir Laboratories Complex
Pcsir Laboratories Complex
Affinity DataIC50: 3.06E+4nMpH: 8.0 T: 2°CAssay Description:LOX inhibiting activity was measured by modifying the spectrophotometric method developed by Tappel. More data for this Ligand-Target Pair
TargetLinoleate 9S-lipoxygenase-4(Glycine max (Soybean) (Glycine hispida))
Pcsir Laboratories Complex
Pcsir Laboratories Complex
Affinity DataIC50: 3.37E+4nMpH: 8.0 T: 2°CAssay Description:LOX inhibiting activity was measured by modifying the spectrophotometric method developed by Tappel. More data for this Ligand-Target Pair
TargetLinoleate 9S-lipoxygenase-4(Glycine max (Soybean) (Glycine hispida))
Pcsir Laboratories Complex
Pcsir Laboratories Complex
Affinity DataIC50: 3.91E+4nMpH: 8.0 T: 2°CAssay Description:LOX inhibiting activity was measured by modifying the spectrophotometric method developed by Tappel. More data for this Ligand-Target Pair
TargetLinoleate 9S-lipoxygenase-4(Glycine max (Soybean) (Glycine hispida))
Pcsir Laboratories Complex
Pcsir Laboratories Complex
Affinity DataIC50: 4.19E+4nMpH: 8.0 T: 2°CAssay Description:LOX inhibiting activity was measured by modifying the spectrophotometric method developed by Tappel. More data for this Ligand-Target Pair
Affinity DataIC50: 4.34E+4nMpH: 8.0 T: 2°CAssay Description:Butyrylcholinesterase activity-inhibiting activities were measured by a slightly modified spectrophotometric method developed by Ellman. DTNB was us...More data for this Ligand-Target Pair
Affinity DataIC50: 4.76E+4nMpH: 8.0 T: 2°CAssay Description:Butyrylcholinesterase activity-inhibiting activities were measured by a slightly modified spectrophotometric method developed by Ellman. DTNB was us...More data for this Ligand-Target Pair
TargetLinoleate 9S-lipoxygenase-4(Glycine max (Soybean) (Glycine hispida))
Pcsir Laboratories Complex
Pcsir Laboratories Complex
Affinity DataIC50: 4.93E+4nMpH: 8.0 T: 2°CAssay Description:LOX inhibiting activity was measured by modifying the spectrophotometric method developed by Tappel. More data for this Ligand-Target Pair
TargetLinoleate 9S-lipoxygenase-4(Glycine max (Soybean) (Glycine hispida))
Pcsir Laboratories Complex
Pcsir Laboratories Complex
Affinity DataIC50: 5.38E+4nMpH: 8.0 T: 2°CAssay Description:LOX inhibiting activity was measured by modifying the spectrophotometric method developed by Tappel. More data for this Ligand-Target Pair