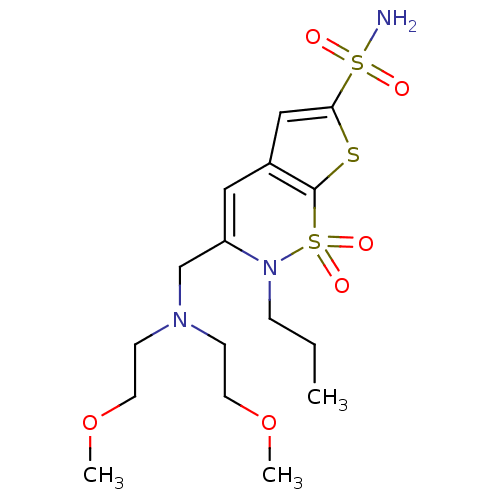
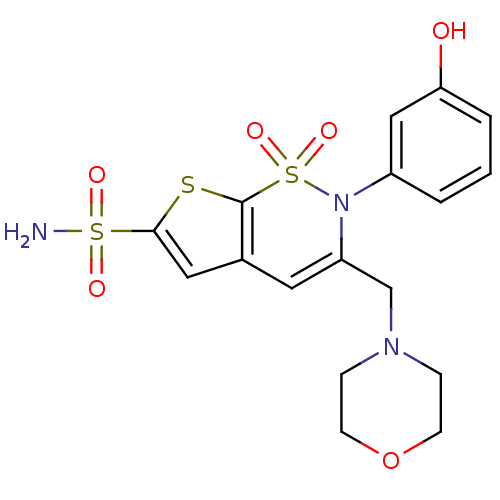
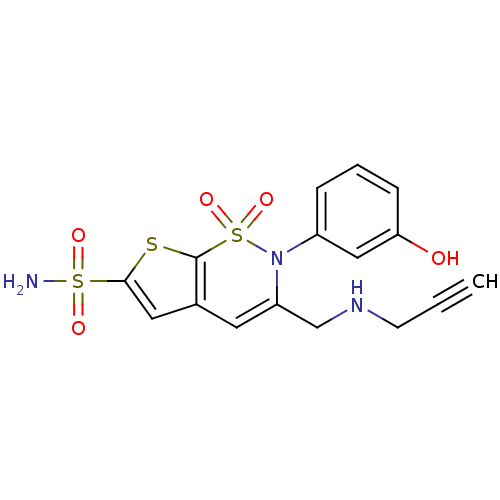
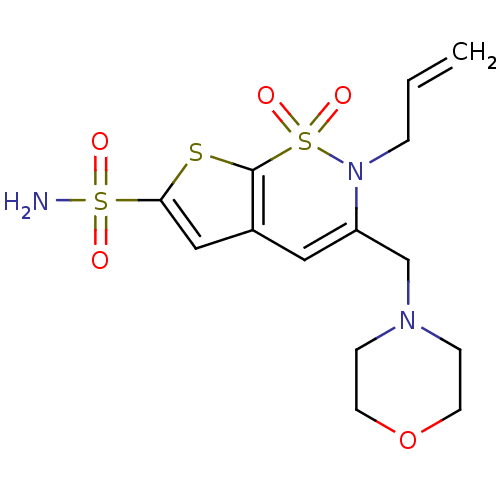
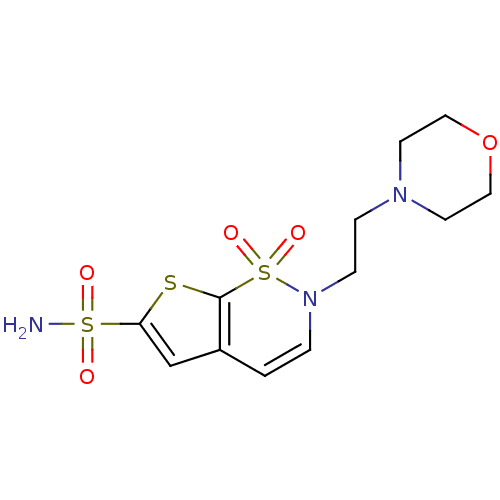
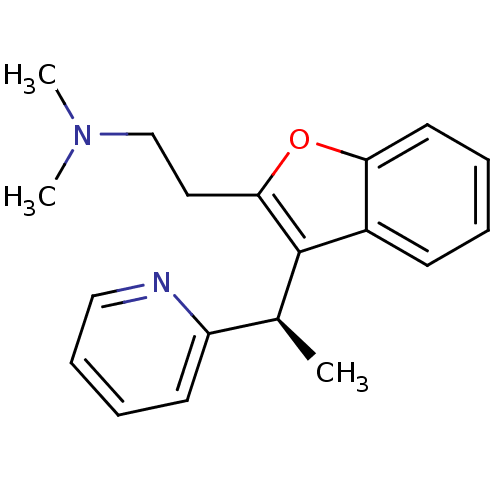
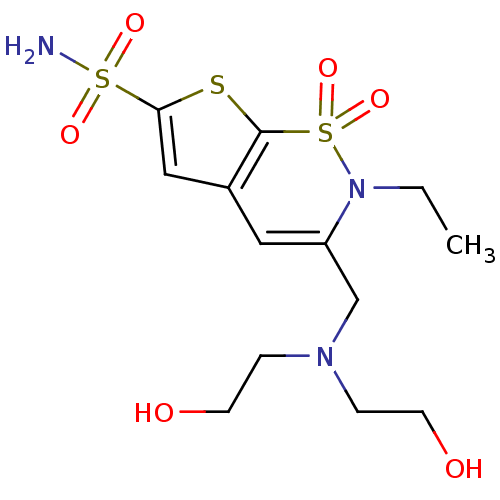
Affinity DataKi: 0.0800nMAssay Description:Inhibitor binding to CA II was determined using a fluorescence competition assay. Displacement of dansylamide and binding of the inhibitor was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.0800nM ΔG°: -59.9kJ/molepH: 7.4 T: 2°CAssay Description:Inhibitor binding to CA II was determined using a fluorescence competition assay. Displacement of dansylamide and binding of the inhibitor was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.0900nMAssay Description:Displacement of [3H]ketanserin from human 5HT2A receptor expressed in HEK293 Flp-In cells by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 0.100nMAssay Description:Inhibitor binding to CA II was determined using a fluorescence competition assay. Displacement of dansylamide and binding of the inhibitor was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.100nM ΔG°: -59.4kJ/molepH: 7.4 T: 2°CAssay Description:Inhibitor binding to CA II was determined using a fluorescence competition assay. Displacement of dansylamide and binding of the inhibitor was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.110nMAssay Description:Inhibitor binding to CA II was determined using a fluorescence competition assay. Displacement of dansylamide and binding of the inhibitor was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.110nMAssay Description:Inhibitor binding to CA II was determined using a fluorescence competition assay. Displacement of dansylamide and binding of the inhibitor was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.120nMAssay Description:Inhibitor binding to CA II was determined using a fluorescence competition assay. Displacement of dansylamide and binding of the inhibitor was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.120nMAssay Description:Inhibitor binding to CA II was determined using a fluorescence competition assay. Displacement of dansylamide and binding of the inhibitor was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.130nM ΔG°: -58.7kJ/molepH: 7.4 T: 2°CAssay Description:Inhibitor binding to CA II was determined using a fluorescence competition assay. Displacement of dansylamide and binding of the inhibitor was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.140nMAssay Description:Displacement of [3H]ketanserin from human 5HT2A receptor expressed in HEK293 Flp-In cells by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 0.150nM ΔG°: -58.3kJ/molepH: 7.4 T: 2°CAssay Description:Inhibitor binding to CA II was determined using a fluorescence competition assay. Displacement of dansylamide and binding of the inhibitor was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.150nMAssay Description:Inhibitor binding to CA II was determined using a fluorescence competition assay. Displacement of dansylamide and binding of the inhibitor was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.150nM ΔG°: -58.3kJ/molepH: 7.4 T: 2°CAssay Description:Inhibitor binding to CA II was determined using a fluorescence competition assay. Displacement of dansylamide and binding of the inhibitor was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.150nM ΔG°: -58.3kJ/molepH: 7.4 T: 2°CAssay Description:Inhibitor binding to CA II was determined using a fluorescence competition assay. Displacement of dansylamide and binding of the inhibitor was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.160nMAssay Description:Inhibitor binding to CA II was determined using a fluorescence competition assay. Displacement of dansylamide and binding of the inhibitor was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.160nMAssay Description:Inhibitor binding to CA II was determined using a fluorescence competition assay. Displacement of dansylamide and binding of the inhibitor was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.190nM ΔG°: -54.9kJ/molepH: 7.4 T: 2°CAssay Description:The membranes prepared from HEK cells transfected with adenosine receptors were used in binding assays. Nonspecific binding was determined in the pre...More data for this Ligand-Target Pair
Affinity DataKi: 0.200nMAssay Description:Inhibitor binding to CA II was determined using a fluorescence competition assay. Displacement of dansylamide and binding of the inhibitor was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.200nMAssay Description:Displacement of [3H]ZM241385 from human adenosine A2A receptor expressed in HEK293 cellsMore data for this Ligand-Target Pair
TargetCorticotropin-releasing factor receptor 1/2(Rattus norvegicus (rat))
Bristol-Myers Squibb
Curated by ChEMBL
Bristol-Myers Squibb
Curated by ChEMBL
Affinity DataKi: 0.200nMAssay Description:In vitro binding affinity to the CRF receptor in rat cortical homogenatesMore data for this Ligand-Target Pair
Affinity DataKi: 0.200nMAssay Description:Displacement of [3H]-ZM241385 from human adenosine A2A receptor expressed in HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 0.200nMAssay Description:Displacement of [3H]ZM241385 from human adenosine A2A receptor expressed in HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 0.210nM ΔG°: -57.5kJ/molepH: 7.4 T: 2°CAssay Description:Inhibitor binding to CA II was determined using a fluorescence competition assay. Displacement of dansylamide and binding of the inhibitor was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.210nMAssay Description:Inhibitor binding to CA II was determined using a fluorescence competition assay. Displacement of dansylamide and binding of the inhibitor was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.210nMAssay Description:Inhibitor binding to CA II was determined using a fluorescence competition assay. Displacement of dansylamide and binding of the inhibitor was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.210nM ΔG°: -57.5kJ/molepH: 7.4 T: 2°CAssay Description:Inhibitor binding to CA II was determined using a fluorescence competition assay. Displacement of dansylamide and binding of the inhibitor was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.220nMAssay Description:Binding affinity against WT human adenosine A2A receptor expressed in CHO cells using [3H]- ZM-241385More data for this Ligand-Target Pair
Affinity DataKi: 0.220nMAssay Description:Inhibitor binding to CA II was determined using a fluorescence competition assay. Displacement of dansylamide and binding of the inhibitor was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.240nM ΔG°: -57.1kJ/molepH: 7.4 T: 2°CAssay Description:Inhibitor binding to CA II was determined using a fluorescence competition assay. Displacement of dansylamide and binding of the inhibitor was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.25nM ΔG°: -57.0kJ/molepH: 7.4 T: 2°CAssay Description:Inhibitor binding to CA II was determined using a fluorescence competition assay. Displacement of dansylamide and binding of the inhibitor was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.25nMAssay Description:Inhibitor binding to CA II was determined using a fluorescence competition assay. Displacement of dansylamide and binding of the inhibitor was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.260nM ΔG°: -56.9kJ/molepH: 7.4 T: 2°CAssay Description:Inhibitor binding to CA II was determined using a fluorescence competition assay. Displacement of dansylamide and binding of the inhibitor was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.260nM ΔG°: -54.2kJ/molepH: 7.4 T: 2°CAssay Description:The membranes prepared from HEK cells transfected with adenosine receptors were used in binding assays. Nonspecific binding was determined in the pre...More data for this Ligand-Target Pair
Affinity DataKi: 0.260nMAssay Description:Inhibitor binding to CA II was determined using a fluorescence competition assay. Displacement of dansylamide and binding of the inhibitor was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.270nMAssay Description:Inhibitor binding to CA II was determined using a fluorescence competition assay. Displacement of dansylamide and binding of the inhibitor was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.300nMAssay Description:Inhibitor binding to CA II was determined using a fluorescence competition assay. Displacement of dansylamide and binding of the inhibitor was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.300nMAssay Description:Binding affinity at adenosine A2A receptorMore data for this Ligand-Target Pair
Affinity DataKi: 0.300nMAssay Description:Binding affinity at adenosine A2A receptorMore data for this Ligand-Target Pair
Affinity DataKi: 0.300nMAssay Description:Displacement of [3H]-ZM241385 from human adenosine A2A receptor expressed in HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 0.300nMAssay Description:Displacement of [3H]-ZM241385 from human adenosine A2A receptor expressed in HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 0.300nMAssay Description:Displacement of [3H]-ZM241385 from human adenosine A2A receptor expressed in HEK293 cellsMore data for this Ligand-Target Pair
Affinity DataKi: 0.300nMAssay Description:Binding affinity at adenosine A2A receptorMore data for this Ligand-Target Pair
Affinity DataKi: 0.300nMAssay Description:Displacement of [3H]-ZM241385 from human adenosine A2A receptor expressed in HEK293 cellsMore data for this Ligand-Target Pair
TargetGamma-aminobutyric acid receptor subunit alpha-1(Rattus norvegicus (Rat))
Neurocrine Biosciences
Curated by PDSP Ki Database
Neurocrine Biosciences
Curated by PDSP Ki Database
Affinity DataKi: 0.310nMAssay Description:Inhibitor binding to CA II was determined using a fluorescence competition assay. Displacement of dansylamide and binding of the inhibitor was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.320nM ΔG°: -56.4kJ/molepH: 7.4 T: 2°CAssay Description:Inhibitor binding to CA II was determined using a fluorescence competition assay. Displacement of dansylamide and binding of the inhibitor was determ...More data for this Ligand-Target Pair
TargetMuscarinic acetylcholine receptor M1(Homo sapiens (Human))
Neurocrine Biosciences
Curated by ChEMBL
Neurocrine Biosciences
Curated by ChEMBL
Affinity DataKi: 0.320nMAssay Description:Displacement of [3H]N-methylscopolamine from human muscarinic M1 receptor expressed in CHO Flp-In cells by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 0.330nMAssay Description:Inhibitor binding to CA II was determined using a fluorescence competition assay. Displacement of dansylamide and binding of the inhibitor was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.330nMAssay Description:Inhibitor binding to CA II was determined using a fluorescence competition assay. Displacement of dansylamide and binding of the inhibitor was determ...More data for this Ligand-Target Pair

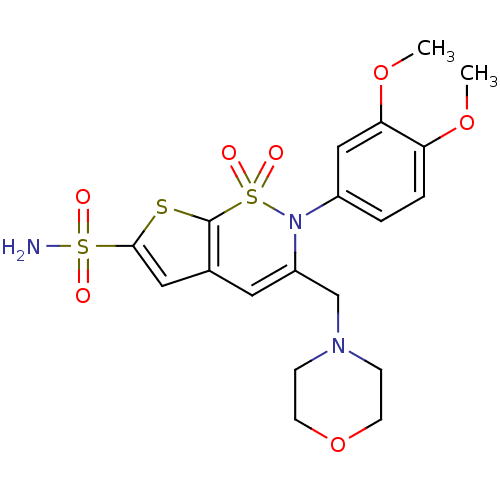



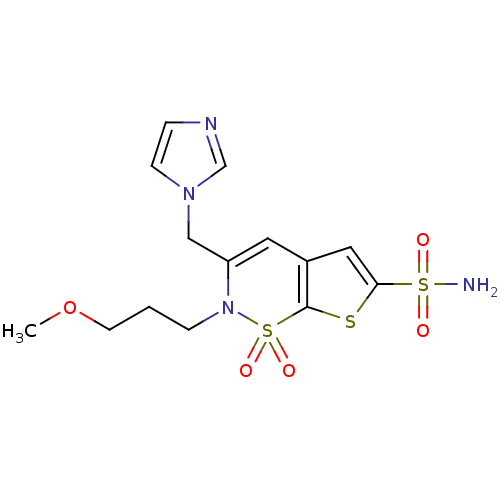
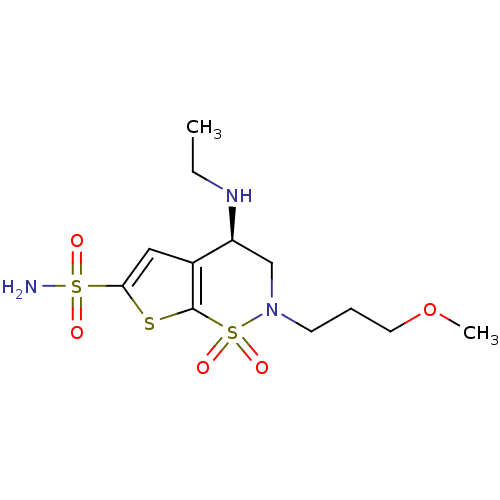
 3D Structure (crystal)
3D Structure (crystal)