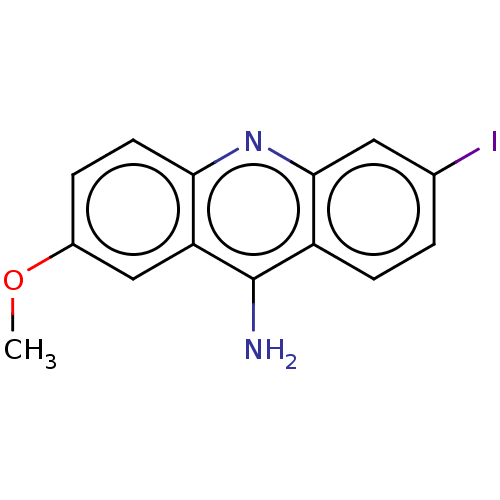
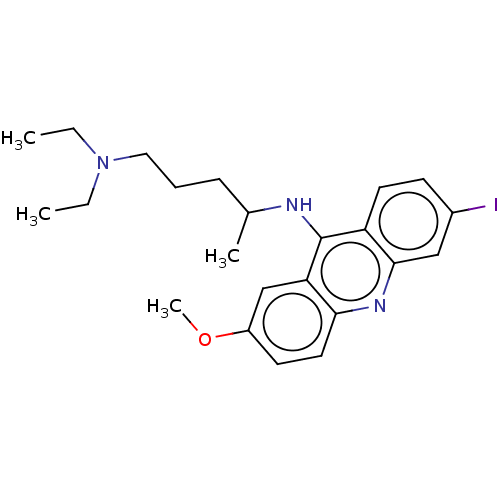
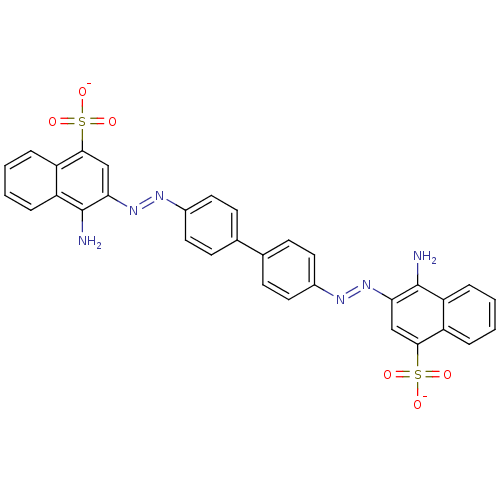
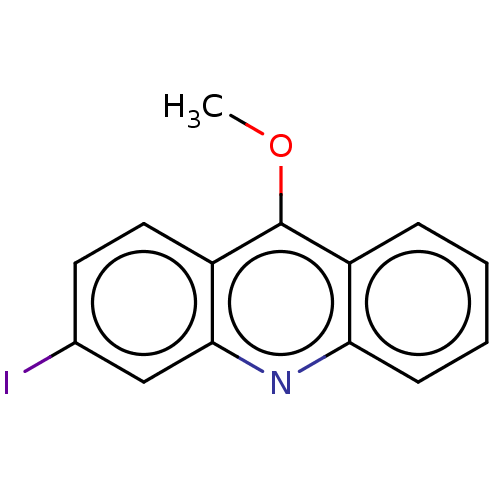
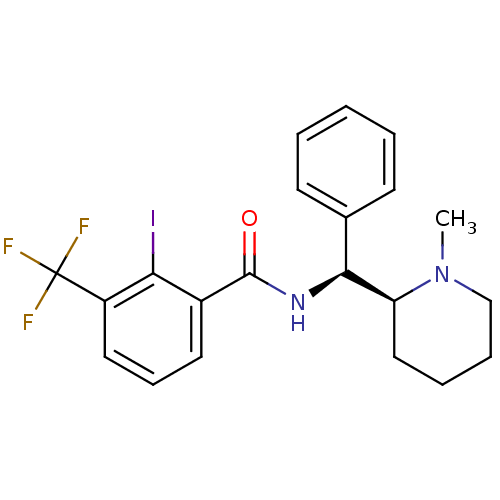
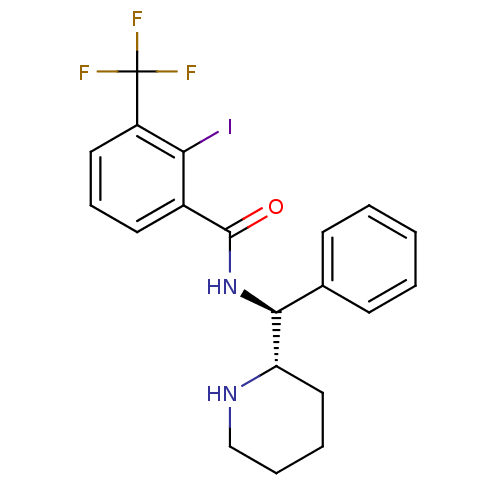
Affinity DataKi: 0.700nMAssay Description:Displacement of [125I]IMPY from beta amyloid in human corpse AD brainMore data for this Ligand-Target Pair
Affinity DataKi: 1.30nMAssay Description:Displacement of [125I]IMPY from beta amyloid in human corpse AD brainMore data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 1/2B(Homo sapiens (Human))
Hamamatsu University School Of Medicine
Curated by ChEMBL
Hamamatsu University School Of Medicine
Curated by ChEMBL
Affinity DataKi: 1.5nMAssay Description:Displacement of [3H]-(E)-N1-(2-methoxybenzyl)cinnamamidine from human NR1a/NR2b receptor expressed in mouse Ltk cellsMore data for this Ligand-Target Pair
Affinity DataKi: 2.30nMAssay Description:Displacement of [125I]IMPY from beta amyloid in human corpse AD brainMore data for this Ligand-Target Pair
Affinity DataKi: 2.80nMAssay Description:Displacement of [125I]IMPY from beta amyloid in human corpse AD brainMore data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2B(Rattus norvegicus (Rat))
Hamamatsu University School Of Medicine
Curated by ChEMBL
Hamamatsu University School Of Medicine
Curated by ChEMBL
Affinity DataKi: 3.09nMAssay Description:Displacement of [3H]ifenprodil from NR2B receptor in rat cortical synaptic membranesMore data for this Ligand-Target Pair
Affinity DataKi: 4.30nMAssay Description:Binding affinity to beta amyloid in human corpse AD brainMore data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2B(Rattus norvegicus (Rat))
Hamamatsu University School Of Medicine
Curated by ChEMBL
Hamamatsu University School Of Medicine
Curated by ChEMBL
Affinity DataKi: 5.75nMAssay Description:Displacement of [3H]ifenprodil from NR2B receptor in rat cortical synaptic membranesMore data for this Ligand-Target Pair
Affinity DataKi: 6nMAssay Description:Binding affinity to beta amyloid in human corpse AD brainMore data for this Ligand-Target Pair
Affinity DataKi: 6nMAssay Description:Displacement of [125I]DMIC from Amyloid beta (1 to 42) after 3 hrs by gamma countingMore data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2B(Rattus norvegicus (Rat))
Hamamatsu University School Of Medicine
Curated by ChEMBL
Hamamatsu University School Of Medicine
Curated by ChEMBL
Affinity DataKi: 7.28nMAssay Description:Displacement of [3H]ifenprodil from NR2B receptor in rat cortical synaptic membranesMore data for this Ligand-Target Pair
Affinity DataKi: 8nMAssay Description:Displacement of [125I]IMPY from amyloid beta (1 to 42) (unknown origin) after 3 hrs by gamma countingMore data for this Ligand-Target Pair
Affinity DataKi: 11.5nMAssay Description:Displacement of [125I]IMPY from beta amyloid in human corpse AD brainMore data for this Ligand-Target Pair
Affinity DataKi: 12nMAssay Description:Displacement of [125I]IMPY from beta amyloid in human corpse AD brainMore data for this Ligand-Target Pair
Affinity DataKi: 13.2nMAssay Description:Displacement of [125I]6-iodo-4'-dimethyaminofl avone from beta amyloid (1-40) protein aggregatesMore data for this Ligand-Target Pair
Affinity DataKi: 14nMAssay Description:Displacement of [125I]DMIC from Amyloid beta (1 to 42) after 3 hrs by gamma countingMore data for this Ligand-Target Pair
Affinity DataKi: 14nMAssay Description:Displacement of [125I]IMPY from amyloid beta (1 to 42) (unknown origin) after 3 hrs by gamma countingMore data for this Ligand-Target Pair
Affinity DataKi: 15.6nMAssay Description:Displacement of [125I]6-iodo-4'-dimethyaminofl avone from beta amyloid (1-42) protein aggregatesMore data for this Ligand-Target Pair
Affinity DataKi: 20nMAssay Description:Displacement of [125I]DMIC from Amyloid beta (1 to 42) after 3 hrs by gamma countingMore data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2B(Rattus norvegicus (Rat))
Hamamatsu University School Of Medicine
Curated by ChEMBL
Hamamatsu University School Of Medicine
Curated by ChEMBL
Affinity DataKi: 20nMAssay Description:Displacement of [3H]ifenprodil from NR2B receptor in rat cortical synaptic membranesMore data for this Ligand-Target Pair
Affinity DataKi: 22.6nMAssay Description:Displacement of [125I]6-iodo-4'-dimethyaminofl avone from beta amyloid (1-40) protein aggregatesMore data for this Ligand-Target Pair
Affinity DataKi: 29nMAssay Description:Displacement of [125I]IMPY from amyloid beta (1 to 42) (unknown origin) after 3 hrs by gamma countingMore data for this Ligand-Target Pair
Affinity DataKi: 29nMAssay Description:Displacement of [125I]6-iodo-4'-dimethyaminofl avone from beta amyloid (1-40) protein aggregatesMore data for this Ligand-Target Pair
Affinity DataKi: 30nMAssay Description:Displacement of [125I]6-iodo-4'-dimethyaminofl avone from beta amyloid (1-42) protein aggregatesMore data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2B(Rattus norvegicus (Rat))
Hamamatsu University School Of Medicine
Curated by ChEMBL
Hamamatsu University School Of Medicine
Curated by ChEMBL
Affinity DataKi: 32.5nMAssay Description:Displacement of [3H]ifenprodil from NR2B receptor in rat cortical synaptic membranesMore data for this Ligand-Target Pair
Affinity DataKi: 38.3nMAssay Description:Displacement of [125I]6-iodo-4'-dimethyaminofl avone from beta amyloid (1-42) protein aggregatesMore data for this Ligand-Target Pair
Affinity DataKi: 39nMAssay Description:Displacement of [125I]IMPY from amyloid beta (1 to 42) (unknown origin) after 3 hrs by gamma countingMore data for this Ligand-Target Pair
Affinity DataKi: 46nMAssay Description:Displacement of [125I]DMIC from Amyloid beta (1 to 42) after 3 hrs by gamma countingMore data for this Ligand-Target Pair
Affinity DataKi: 72.5nMAssay Description:Displacement of [125I]6-iodo-4'-dimethyaminofl avone from beta amyloid (1-40) protein aggregatesMore data for this Ligand-Target Pair
Affinity DataKi: 77.2nMAssay Description:Displacement of [125I]6-iodo-4'-dimethyaminofl avone from beta amyloid (1-42) protein aggregatesMore data for this Ligand-Target Pair
Affinity DataKi: 84nMAssay Description:Displacement of [125I]IMPY from amyloid beta (1 to 42) (unknown origin) after 3 hrs by gamma countingMore data for this Ligand-Target Pair
Affinity DataKi: 121nMAssay Description:Displacement of [125I]IMPY from amyloid beta (1 to 42) (unknown origin) after 3 hrs by gamma countingMore data for this Ligand-Target Pair
Affinity DataKi: 326nMAssay Description:Displacement of [125I]DMIC from Amyloid beta (1 to 42) after 3 hrs by gamma countingMore data for this Ligand-Target Pair
Affinity DataKi: 353nMAssay Description:Displacement of [125I]IMPY from amyloid beta (1 to 42) (unknown origin) after 3 hrs by gamma countingMore data for this Ligand-Target Pair
Affinity DataKi: >1.00E+3nMAssay Description:Displacement of [125I]6-iodo-4'-dimethyaminofl avone from beta amyloid (1-40) protein aggregatesMore data for this Ligand-Target Pair
Affinity DataKi: >1.00E+3nMAssay Description:Displacement of [125I]6-iodo-4'-dimethyaminofl avone from beta amyloid (1-42) protein aggregatesMore data for this Ligand-Target Pair
Affinity DataKi: >1.00E+3nMAssay Description:Displacement of [125I]6-iodo-4'-dimethyaminofl avone from beta amyloid (1-42) protein aggregatesMore data for this Ligand-Target Pair
Affinity DataKi: >1.00E+3nMAssay Description:Displacement of [125I]6-iodo-4'-dimethyaminofl avone from beta amyloid (1-40) protein aggregatesMore data for this Ligand-Target Pair
Affinity DataKi: >1.00E+4nMAssay Description:Displacement of [125I]IMPY from amyloid beta (1 to 42) (unknown origin) after 3 hrs by gamma countingMore data for this Ligand-Target Pair
TargetSodium- and chloride-dependent glycine transporter 1(Homo sapiens (Human))
Nagasaki University
Curated by ChEMBL
Nagasaki University
Curated by ChEMBL
Affinity DataIC50: 0.390nMAssay Description:Inhibition of GlyT1 in human JAR cells assessed as inhibition of [3H]glycine uptake preincubated for 1 mins measured after 2 hrs by liquid scintillat...More data for this Ligand-Target Pair
TargetSodium- and chloride-dependent glycine transporter 1(Homo sapiens (Human))
Nagasaki University
Curated by ChEMBL
Nagasaki University
Curated by ChEMBL
Affinity DataIC50: 2.40nMAssay Description:Inhibition of GlyT1 in human JAR cells assessed as inhibition of [3H]glycine uptake preincubated for 1 mins measured after 2 hrs by liquid scintillat...More data for this Ligand-Target Pair
TargetSodium- and chloride-dependent glycine transporter 1(Homo sapiens (Human))
Nagasaki University
Curated by ChEMBL
Nagasaki University
Curated by ChEMBL
Affinity DataIC50: 2.5nMAssay Description:Inhibition of GlyT1 in human JAR cells assessed as inhibition of [3H]glycine uptake preincubated for 1 mins measured after 2 hrs by liquid scintillat...More data for this Ligand-Target Pair
TargetSodium- and chloride-dependent glycine transporter 1(Homo sapiens (Human))
Nagasaki University
Curated by ChEMBL
Nagasaki University
Curated by ChEMBL
Affinity DataIC50: 7.5nMAssay Description:Inhibition of GlyT1 in human JAR cells assessed as inhibition of [3H]glycine uptake preincubated for 1 mins measured after 2 hrs by liquid scintillat...More data for this Ligand-Target Pair
TargetSodium- and chloride-dependent glycine transporter 1(Homo sapiens (Human))
Nagasaki University
Curated by ChEMBL
Nagasaki University
Curated by ChEMBL
Affinity DataIC50: 43nMAssay Description:Inhibition of GlyT1 in human JAR cells assessed as inhibition of [3H]glycine uptake preincubated for 1 mins measured after 2 hrs by liquid scintillat...More data for this Ligand-Target Pair
TargetSodium- and chloride-dependent glycine transporter 1(Homo sapiens (Human))
Nagasaki University
Curated by ChEMBL
Nagasaki University
Curated by ChEMBL
Affinity DataIC50: 80nMAssay Description:Inhibition of GlyT1 in human JAR cells assessed as inhibition of [3H]glycine uptake preincubated for 1 mins measured after 2 hrs by liquid scintillat...More data for this Ligand-Target Pair
TargetSodium- and chloride-dependent glycine transporter 1(Homo sapiens (Human))
Nagasaki University
Curated by ChEMBL
Nagasaki University
Curated by ChEMBL
Affinity DataIC50: 314nMAssay Description:Inhibition of human GlyT1 expressed in CHO cells assessed as inhibition of [3H]glycine uptake preincubated for 1 mins measured after 2 hrs by liquid ...More data for this Ligand-Target Pair
TargetBaculoviral IAP repeat-containing protein 5(Homo sapiens (Human))
Nagasaki University
Curated by ChEMBL
Nagasaki University
Curated by ChEMBL
Affinity DataKd: 34nMAssay Description:Binding affinity to recombinant full length N-terminal GST-tagged human survivin expressed in Escherichia coli BL21 incubated for 1 hr under shaking ...More data for this Ligand-Target Pair
Affinity DataKd: 48nMAssay Description:Binding affinity to amyloid beta aggregate (unknown origin) by centrifugation methodMore data for this Ligand-Target Pair
TargetBaculoviral IAP repeat-containing protein 5(Homo sapiens (Human))
Nagasaki University
Curated by ChEMBL
Nagasaki University
Curated by ChEMBL
Affinity DataKd: 44nMAssay Description:Binding affinity to recombinant full length N-terminal GST-tagged human survivin expressed in Escherichia coli BL21 incubated for 1 hr under shaking ...More data for this Ligand-Target Pair
TargetSodium- and chloride-dependent glycine transporter 1(attus norvegicus (Rat))
Nagasaki University
Curated by ChEMBL
Nagasaki University
Curated by ChEMBL
Affinity DataKd: 1.60nMAssay Description:Binding affinity to GlyT1 in rat cortical brain homogenate after 60 mins by gamma countingMore data for this Ligand-Target Pair

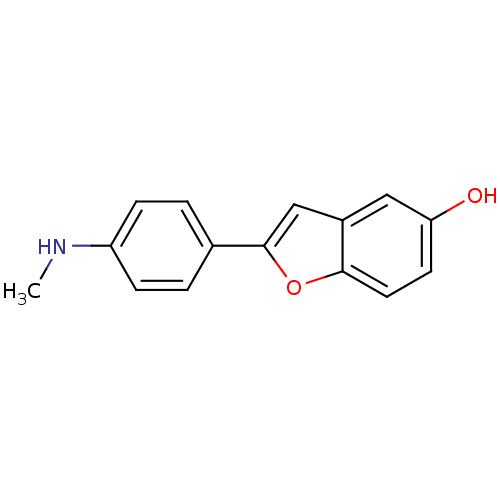


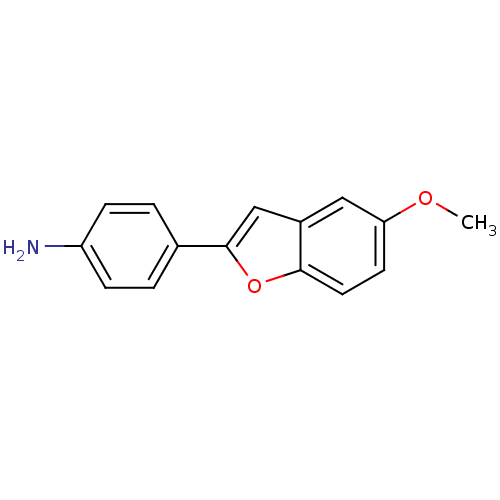
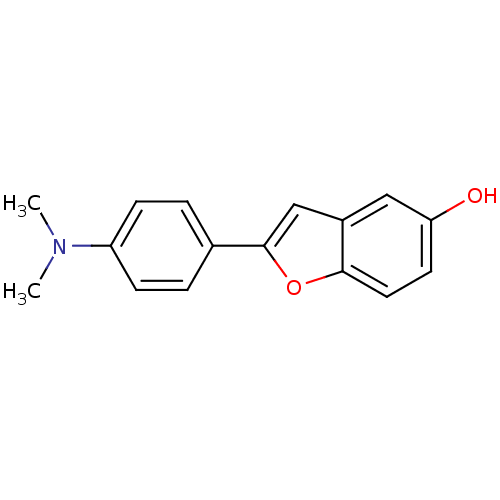
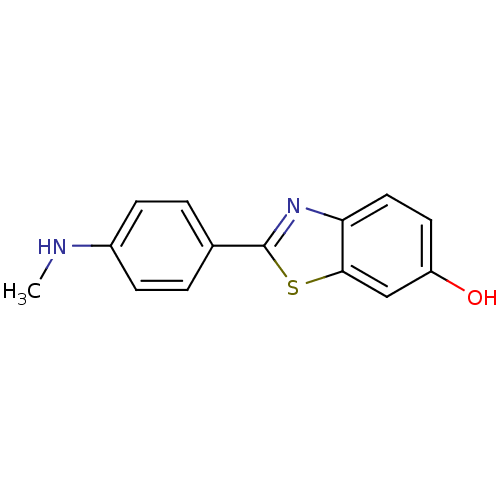

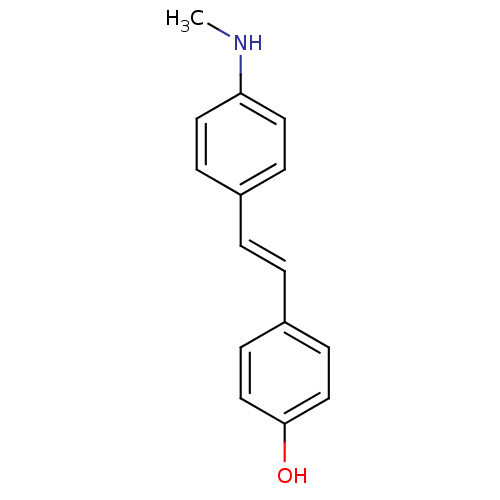


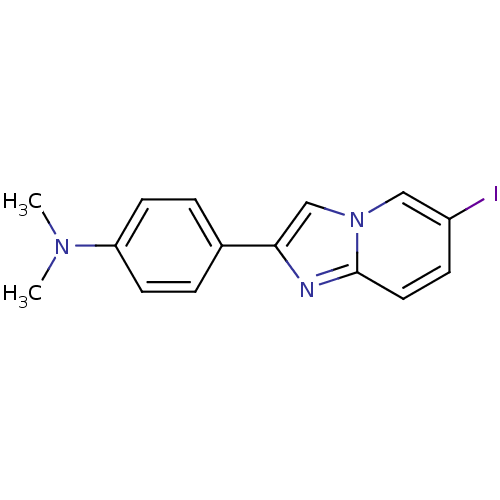




 3D Structure (crystal)
3D Structure (crystal)