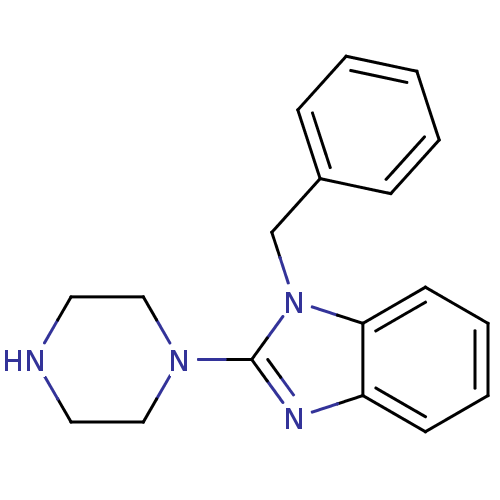
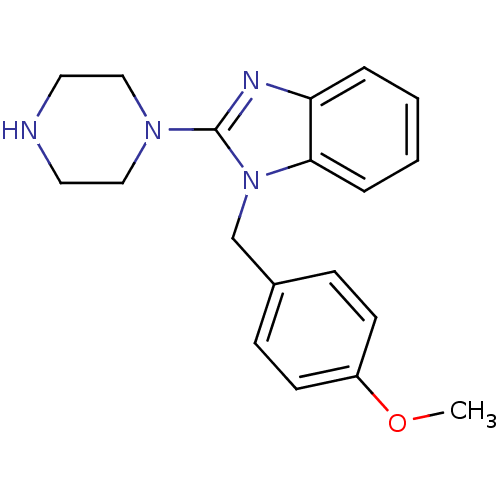
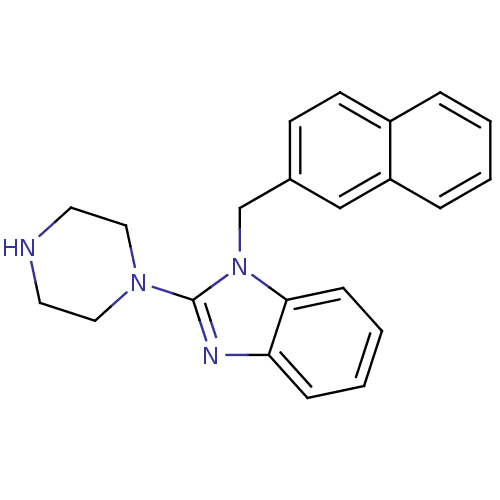
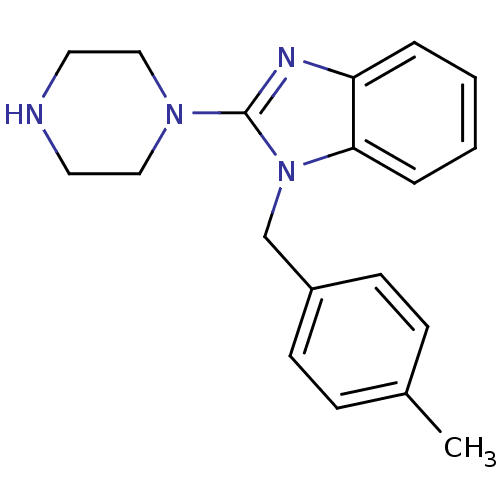
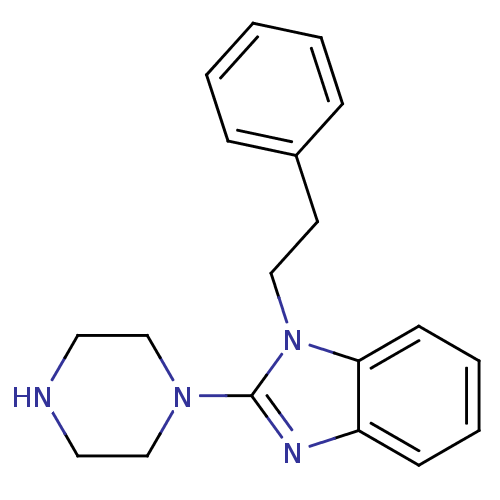
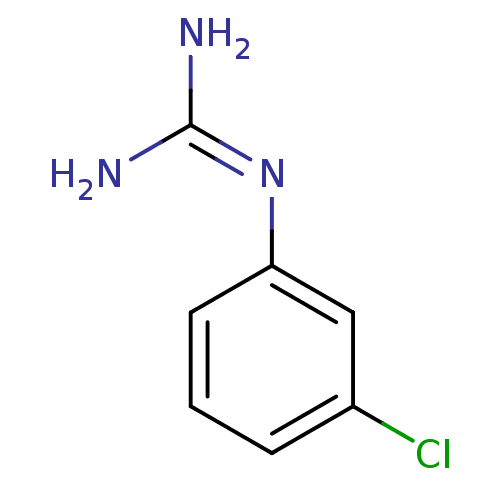
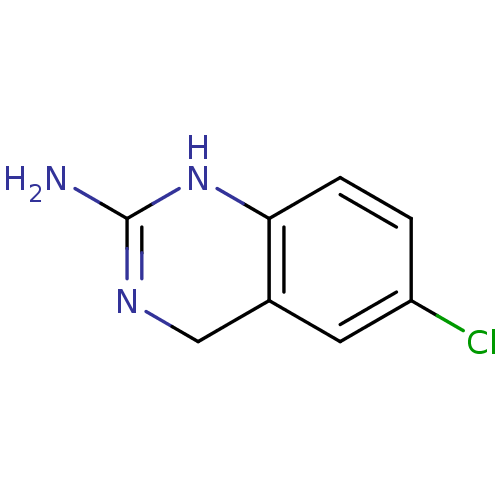
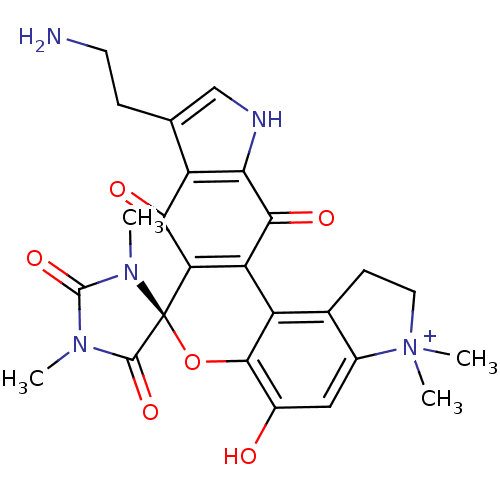
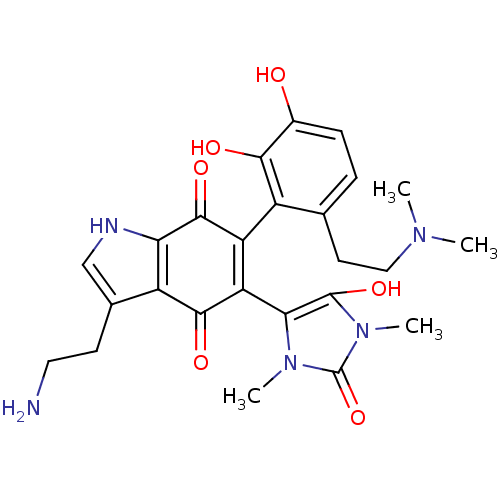
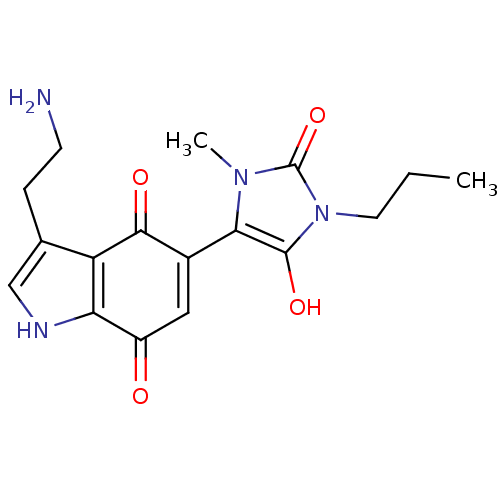
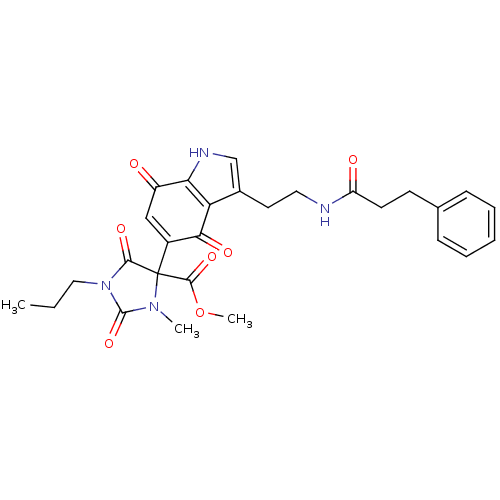
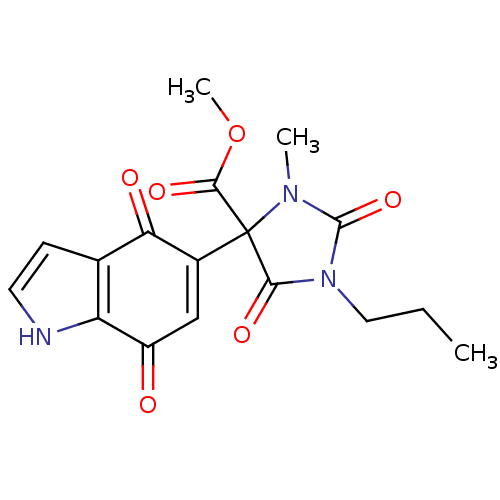
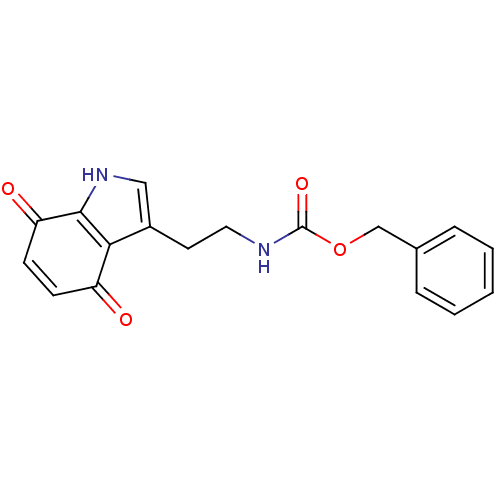
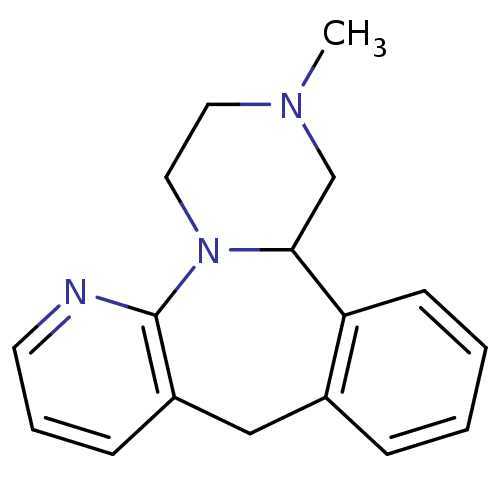
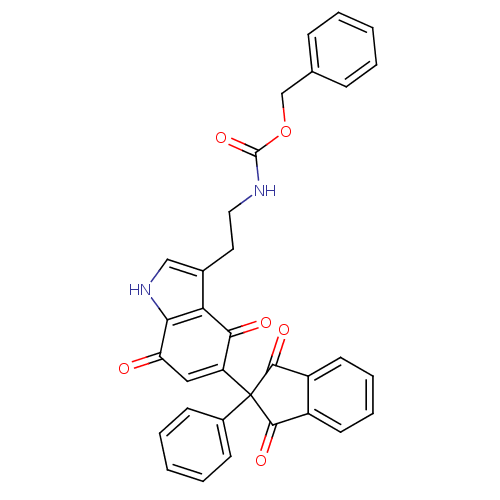
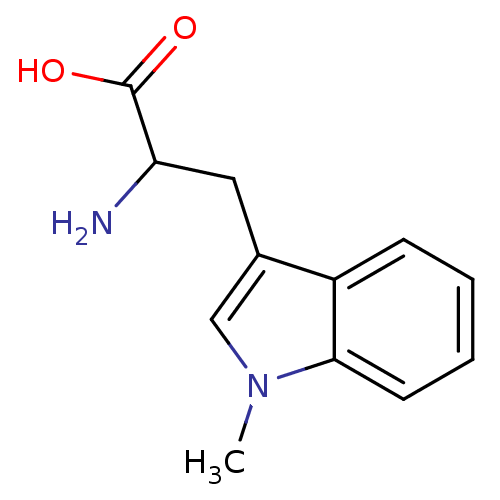
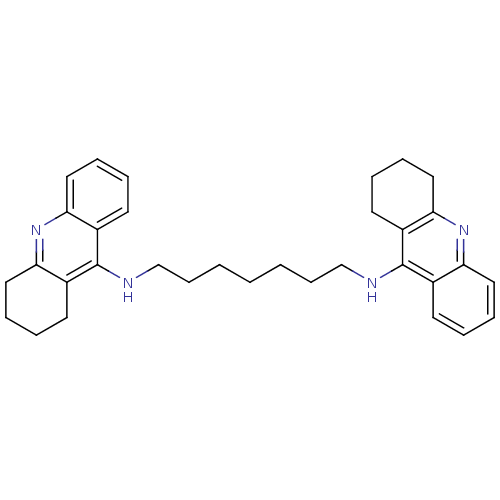
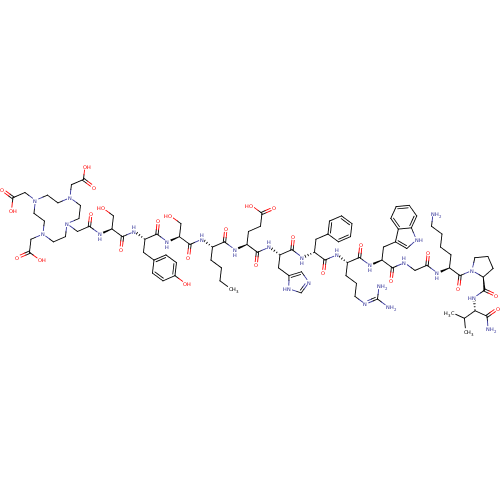
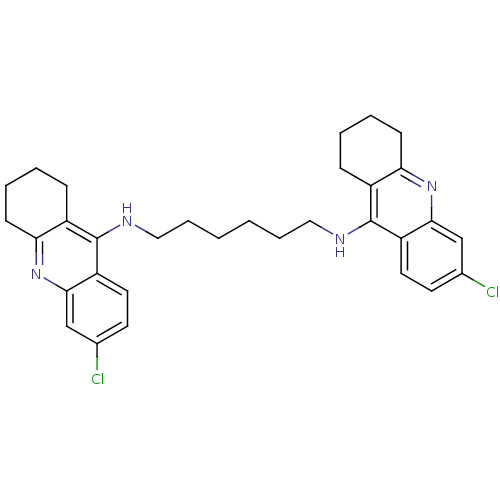
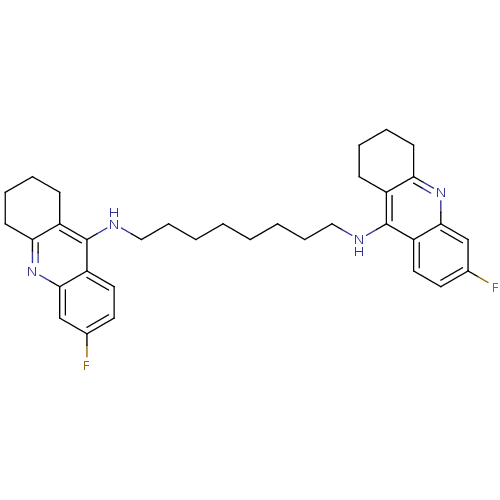
Target5-hydroxytryptamine receptor 3A(Homo sapiens (Human))
The University Of Louisiana At Monroe
Curated by ChEMBL
The University Of Louisiana At Monroe
Curated by ChEMBL
Affinity DataKi: 0.360nMAssay Description:Binding affinity of the compound against human 5-hydroxytryptamine 3A receptorMore data for this Ligand-Target Pair
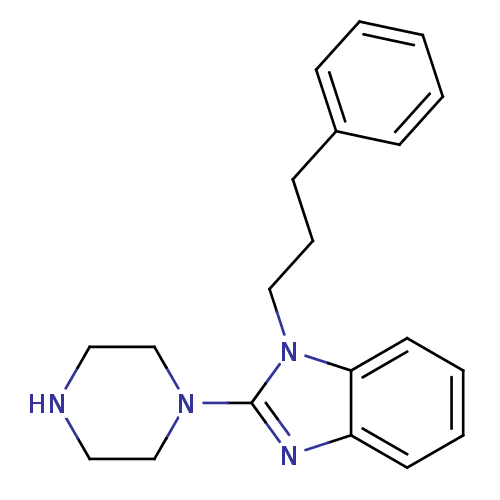
Target5-hydroxytryptamine receptor 3A(Homo sapiens (Human))
The University Of Louisiana At Monroe
Curated by ChEMBL
The University Of Louisiana At Monroe
Curated by ChEMBL
Affinity DataKi: 0.420nMAssay Description:Binding affinity of the compound against human 5-hydroxytryptamine 3A receptorMore data for this Ligand-Target Pair
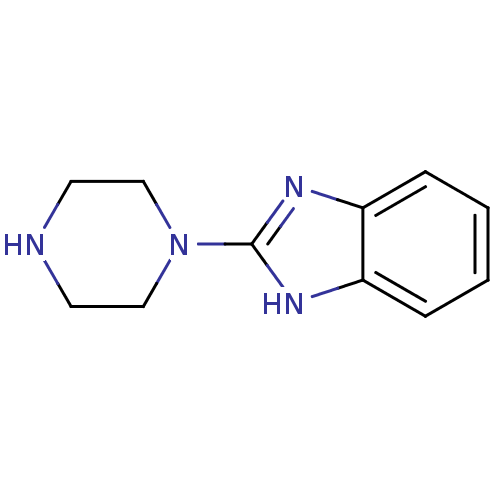
Target5-hydroxytryptamine receptor 3A(Homo sapiens (Human))
The University Of Louisiana At Monroe
Curated by ChEMBL
The University Of Louisiana At Monroe
Curated by ChEMBL
Affinity DataKi: 1.40nMAssay Description:Binding affinity of the compound against human 5-hydroxytryptamine 3A receptorMore data for this Ligand-Target Pair
Target5-hydroxytryptamine receptor 3A(Homo sapiens (Human))
The University Of Louisiana At Monroe
Curated by ChEMBL
The University Of Louisiana At Monroe
Curated by ChEMBL
Affinity DataKi: 1.70nMAssay Description:Binding affinity of the compound against human 5-hydroxytryptamine 3A receptorMore data for this Ligand-Target Pair
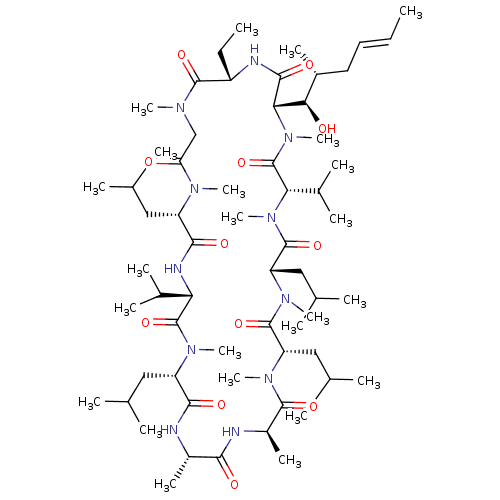
TargetPeptidyl-prolyl cis-trans isomerase FKBP1B(Homo sapiens (Human))
University Of Wisconsin-Madison
Curated by ChEMBL
University Of Wisconsin-Madison
Curated by ChEMBL
Affinity DataKi: 3nMAssay Description:The compound was tested for inhibition of Peptidyl-propyl isomerase (PPIase).More data for this Ligand-Target Pair
Target5-hydroxytryptamine receptor 3A(Homo sapiens (Human))
The University Of Louisiana At Monroe
Curated by ChEMBL
The University Of Louisiana At Monroe
Curated by ChEMBL
Affinity DataKi: 5.20nMAssay Description:Binding affinity of the compound against human 5-hydroxytryptamine 3A receptorMore data for this Ligand-Target Pair
TargetPeptidyl-prolyl cis-trans isomerase FKBP1B(Homo sapiens (Human))
University Of Wisconsin-Madison
Curated by ChEMBL
University Of Wisconsin-Madison
Curated by ChEMBL
Affinity DataKi: 6nMAssay Description:The compound was tested for inhibition of Peptidyl-propyl isomerase (PPIase).More data for this Ligand-Target Pair
TargetPeptidyl-prolyl cis-trans isomerase FKBP1B(Homo sapiens (Human))
University Of Wisconsin-Madison
Curated by ChEMBL
University Of Wisconsin-Madison
Curated by ChEMBL
Affinity DataKi: 6nMAssay Description:The compound was tested for inhibition of Peptidyl-propyl isomerase (PPIase).More data for this Ligand-Target Pair
TargetPeptidyl-prolyl cis-trans isomerase FKBP1B(Homo sapiens (Human))
University Of Wisconsin-Madison
Curated by ChEMBL
University Of Wisconsin-Madison
Curated by ChEMBL
Affinity DataKi: 6nMAssay Description:The compound was tested for inhibition of Peptidyl-propyl isomerase (PPIase).More data for this Ligand-Target Pair
Target5-hydroxytryptamine receptor 3A(Homo sapiens (Human))
The University Of Louisiana At Monroe
Curated by ChEMBL
The University Of Louisiana At Monroe
Curated by ChEMBL
Affinity DataKi: 7.40nMAssay Description:Binding affinity of the compound against human 5-hydroxytryptamine 3A receptorMore data for this Ligand-Target Pair
Target5-hydroxytryptamine receptor 3A(Homo sapiens (Human))
The University Of Louisiana At Monroe
Curated by ChEMBL
The University Of Louisiana At Monroe
Curated by ChEMBL
Affinity DataKi: 8nMAssay Description:Binding affinity of the compound against human 5-hydroxytryptamine 3A receptorMore data for this Ligand-Target Pair
Target5-hydroxytryptamine receptor 3A(Homo sapiens (Human))
The University Of Louisiana At Monroe
Curated by ChEMBL
The University Of Louisiana At Monroe
Curated by ChEMBL
Affinity DataKi: 19nMAssay Description:Binding affinity of the compound against human 5-hydroxytryptamine 3A receptorMore data for this Ligand-Target Pair
Target5-hydroxytryptamine receptor 3A(Homo sapiens (Human))
The University Of Louisiana At Monroe
Curated by ChEMBL
The University Of Louisiana At Monroe
Curated by ChEMBL
Affinity DataKi: 32nMAssay Description:Displacement of [3H]GR65630 from 5-HT3 receptor (unknown origin) expressed in mouse/rat NG108-15 cells after 30 mins by by liquid scintillation count...More data for this Ligand-Target Pair
TargetPeptidyl-prolyl cis-trans isomerase FKBP1B(Homo sapiens (Human))
University Of Wisconsin-Madison
Curated by ChEMBL
University Of Wisconsin-Madison
Curated by ChEMBL
Affinity DataKi: 33nMAssay Description:The compound was tested for inhibition of Peptidyl-propyl isomerase (PPIase).More data for this Ligand-Target Pair
Target5-hydroxytryptamine receptor 3A(Homo sapiens (Human))
The University Of Louisiana At Monroe
Curated by ChEMBL
The University Of Louisiana At Monroe
Curated by ChEMBL
Affinity DataKi: 34nMAssay Description:Displacement of [3H]GR65630 from 5-HT3 receptor (unknown origin) expressed in mouse/rat NG108-15 cells after 30 mins by by liquid scintillation count...More data for this Ligand-Target Pair
Affinity DataKi: 41nMAssay Description:Inhibition of human recombinant indoleamine-2,3-dioxygenaseMore data for this Ligand-Target Pair
Affinity DataKi: 41nM ΔG°: -43.9kJ/molepH: 6.5 T: 2°CAssay Description:The ability of the compounds prepared in this study to inhibit purified recombinant human IDO was evaluated with a steady state spectrophotometric as...More data for this Ligand-Target Pair
Affinity DataKi: 80nMAssay Description:Inhibition of human recombinant indoleamine-2,3-dioxygenaseMore data for this Ligand-Target Pair
Affinity DataKi: 80nMAssay Description:Inhibition of human recombinant indoleamine-2,3-dioxygenaseMore data for this Ligand-Target Pair
Target5-hydroxytryptamine receptor 3A(Homo sapiens (Human))
The University Of Louisiana At Monroe
Curated by ChEMBL
The University Of Louisiana At Monroe
Curated by ChEMBL
Affinity DataKi: 80nMAssay Description:Displacement of [3H]GR65630 from human 5-HT3 receptor expressed in African green monkey COS cells after 90 mins by scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataKi: 190nM ΔG°: -39.9kJ/molepH: 6.5 T: 2°CAssay Description:The ability of the compounds prepared in this study to inhibit purified recombinant human IDO was evaluated with a steady state spectrophotometric as...More data for this Ligand-Target Pair
Affinity DataKi: 200nM ΔG°: -39.8kJ/molepH: 6.5 T: 2°CAssay Description:The ability of the compounds prepared in this study to inhibit purified recombinant human IDO was evaluated with a steady state spectrophotometric as...More data for this Ligand-Target Pair
Affinity DataKi: 260nM ΔG°: -39.1kJ/molepH: 6.5 T: 2°CAssay Description:The ability of the compounds prepared in this study to inhibit purified recombinant human IDO was evaluated with a steady state spectrophotometric as...More data for this Ligand-Target Pair
Affinity DataKi: 260nM ΔG°: -39.1kJ/molepH: 6.5 T: 2°CAssay Description:The ability of the compounds prepared in this study to inhibit purified recombinant human IDO was evaluated with a steady state spectrophotometric as...More data for this Ligand-Target Pair
Affinity DataKi: 420nM ΔG°: -37.9kJ/molepH: 6.5 T: 2°CAssay Description:The ability of the compounds prepared in this study to inhibit purified recombinant human IDO was evaluated with a steady state spectrophotometric as...More data for this Ligand-Target Pair
Affinity DataKi: 1.49E+3nM ΔG°: -34.6kJ/molepH: 6.5 T: 2°CAssay Description:The ability of the compounds prepared in this study to inhibit purified recombinant human IDO was evaluated with a steady state spectrophotometric as...More data for this Ligand-Target Pair
Target5-hydroxytryptamine receptor 3A(Homo sapiens (Human))
The University Of Louisiana At Monroe
Curated by ChEMBL
The University Of Louisiana At Monroe
Curated by ChEMBL
Affinity DataKi: 2.90E+3nMAssay Description:Displacement of [3H]GR65630 from human 5-HT3A receptor expressed in HEK293 cells by liquid scintillation counting analysisMore data for this Ligand-Target Pair
TargetSigma non-opioid intracellular receptor 1(Cavia porcellus (Guinea pig))
Virginia Commonwealth University
Curated by ChEMBL
Virginia Commonwealth University
Curated by ChEMBL
Affinity DataKi: 6.15E+3nMAssay Description:Displacement of [3H]Pentazocine from guinea pig sigma1 receptor after 90 mins by scintillation counting analysisMore data for this Ligand-Target Pair
TargetAcetylcholine receptor subunit alpha/Neuronal acetylcholine receptor subunit beta-4(Homo sapiens (Human))
Virginia Commonwealth University
Curated by ChEMBL
Virginia Commonwealth University
Curated by ChEMBL
Affinity DataKi: >1.00E+4nMAssay Description:Displacement of [3H]-epibatidine from human alpha2beta4 nAChR transfected in HEK293 cells by scintillation counting analysisMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-4/beta-2(Homo sapiens (Human))
Virginia Commonwealth University
Curated by ChEMBL
Virginia Commonwealth University
Curated by ChEMBL
Affinity DataKi: >1.00E+4nMAssay Description:Displacement of [3H]-epibatidine from human alpha4beta2 nAChR transfected in HEK293 cells by scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataKi: >1.00E+4nMAssay Description:Displacement of [[3H]N-methylspiperone from human recombinant dopamine D2 receptor expressed in human fibroblasts after 90 mins by scintillation coun...More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+4nMAssay Description:Displacement of [3H]Pyrilamine from human recombinant histamine H1 receptor expressed in HEK cells after 90 mins by scintillation counting analysisMore data for this Ligand-Target Pair
TargetAcetylcholine receptor subunit alpha/Neuronal acetylcholine receptor subunit beta-2(Homo sapiens (Human))
Virginia Commonwealth University
Curated by ChEMBL
Virginia Commonwealth University
Curated by ChEMBL
Affinity DataKi: >1.00E+4nMAssay Description:Displacement of [3H]-epibatidine from human alpha2beta2 nAChR transfected in HEK293 cells by scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataKi: 1.09E+4nM ΔG°: -29.5kJ/molepH: 6.5 T: 2°CAssay Description:The ability of the compounds prepared in this study to inhibit purified recombinant human IDO was evaluated with a steady state spectrophotometric as...More data for this Ligand-Target Pair
Affinity DataKi: 6.20E+4nM ΔG°: -25.0kJ/molepH: 6.5 T: 2°CAssay Description:The ability of the compounds prepared in this study to inhibit purified recombinant human IDO was evaluated with a steady state spectrophotometric as...More data for this Ligand-Target Pair
Affinity DataIC50: 0.0700nMpH: 7.4 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at...More data for this Ligand-Target Pair
Affinity DataIC50: 0.200nMpH: 7.4 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at...More data for this Ligand-Target Pair
Affinity DataIC50: 0.210nMAssay Description:Displacement of [125I]-[Nle4-D-Phe7]-alpha-MSH from MC1R in mouse B16-F10 cells after 1.5 hrs by gamma countingMore data for this Ligand-Target Pair
Affinity DataIC50: 0.25nMAssay Description:Displacement of [125I]-[Nle4-D-Phe7]-alpha-MSH from MC1R in mouse B16-F10 cells after 1.5 hrs by gamma countingMore data for this Ligand-Target Pair
Affinity DataIC50: 0.300nMpH: 7.4 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at...More data for this Ligand-Target Pair
Affinity DataIC50: 0.590nMAssay Description:Displacement of [125I]-[Nle4-D-Phe7]-alpha-MSH from MC1R in mouse B16-F10 cells after 1.5 hrs by gamma countingMore data for this Ligand-Target Pair
Affinity DataIC50: 0.600nMpH: 7.4 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at...More data for this Ligand-Target Pair
Affinity DataIC50: 0.600nMpH: 7.4 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at...More data for this Ligand-Target Pair
Affinity DataIC50: 0.700nMpH: 7.4 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at...More data for this Ligand-Target Pair
Affinity DataIC50: 0.760nMAssay Description:Displacement of [125I]-[Nle4-D-Phe7]-alpha-MSH from MC1R in mouse B16-F10 cells after 1.5 hrs by gamma countingMore data for this Ligand-Target Pair
Affinity DataIC50: 0.820nMAssay Description:Displacement of [125I]-[Nle4-D-Phe7]-alpha-MSH from MC1R in mouse B16-F10 cells after 1.5 hrs by gamma countingMore data for this Ligand-Target Pair
Affinity DataIC50: 0.900nMpH: 7.4 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at...More data for this Ligand-Target Pair
Affinity DataIC50: 1.30nMpH: 7.4 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at...More data for this Ligand-Target Pair
Affinity DataIC50: 1.40nMpH: 7.4 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at...More data for this Ligand-Target Pair
Affinity DataIC50: 1.60nMpH: 7.4 T: 2°CAssay Description:The cholinesterase assays were performed using colorimetric method reported by Ellman. Enzyme activity was determined by measuring the absorbance at...More data for this Ligand-Target Pair