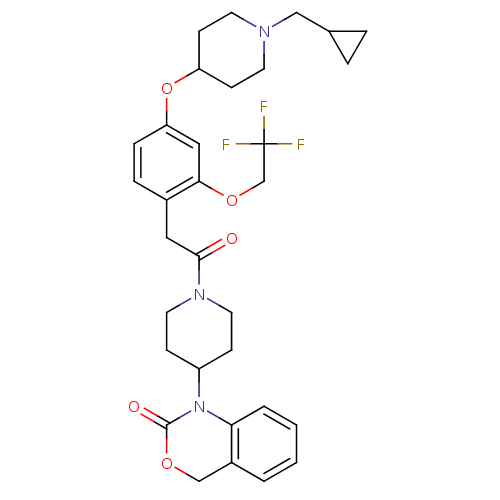
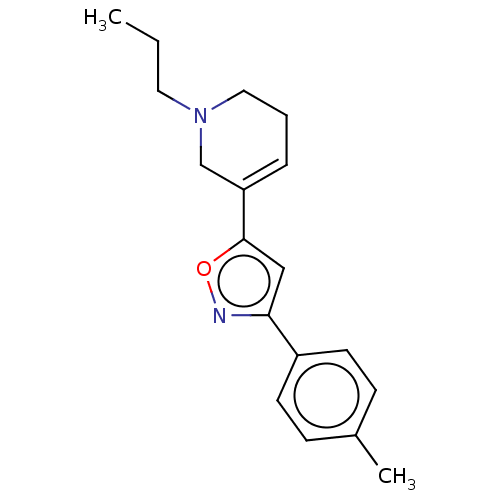
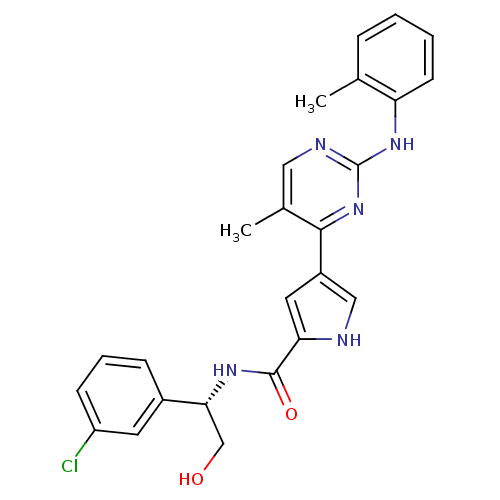
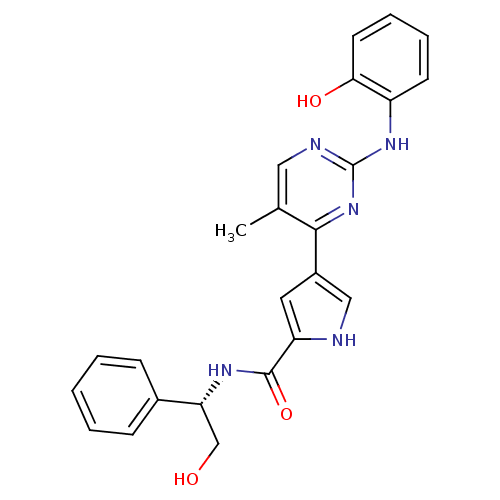
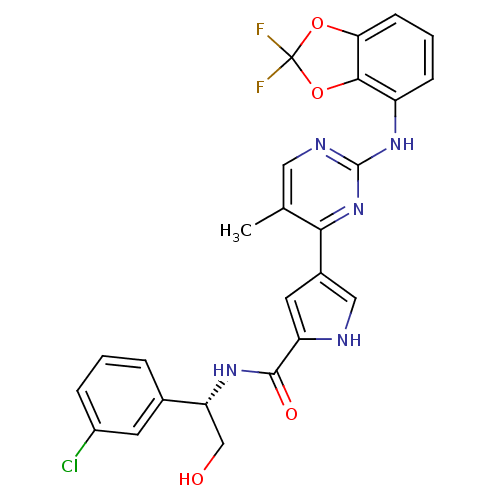
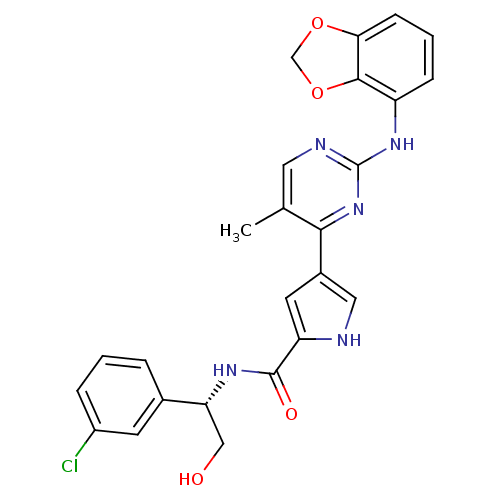
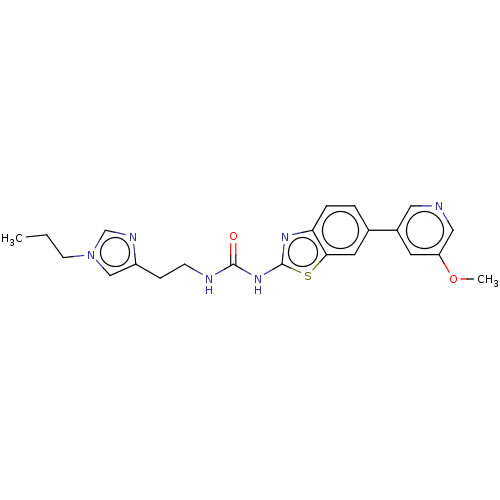
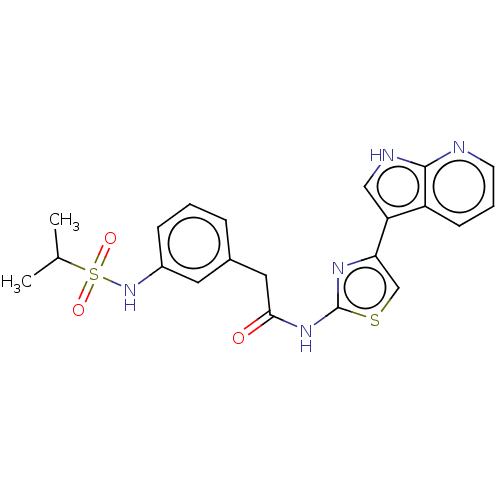
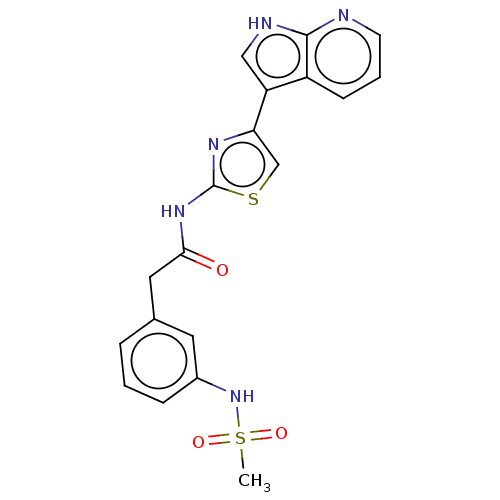
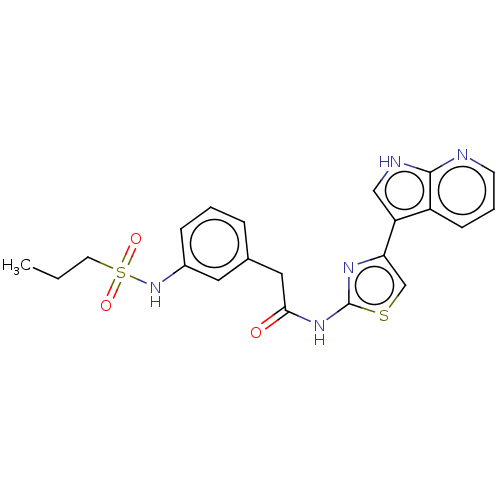
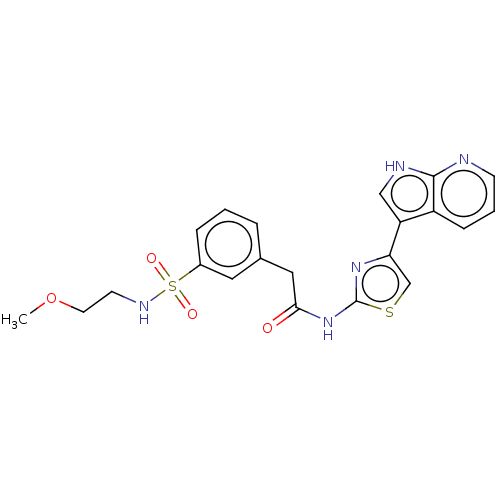
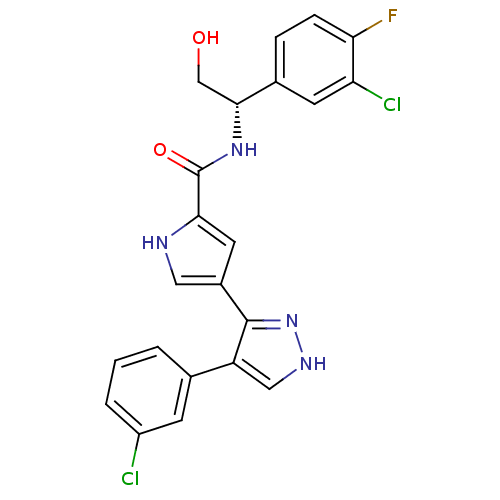
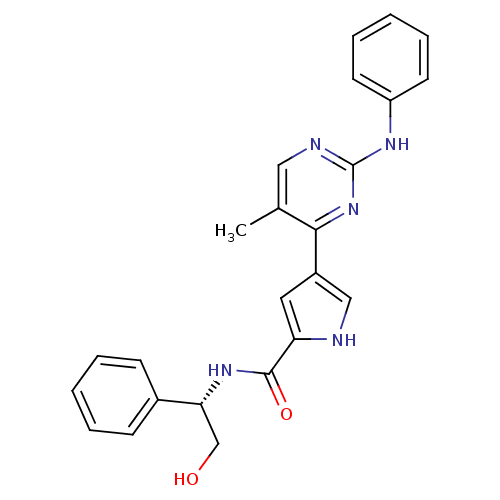
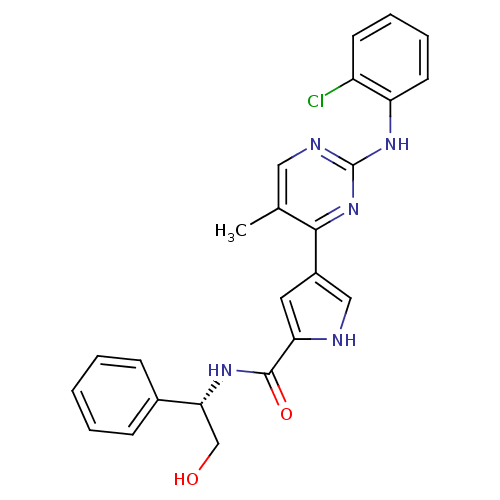
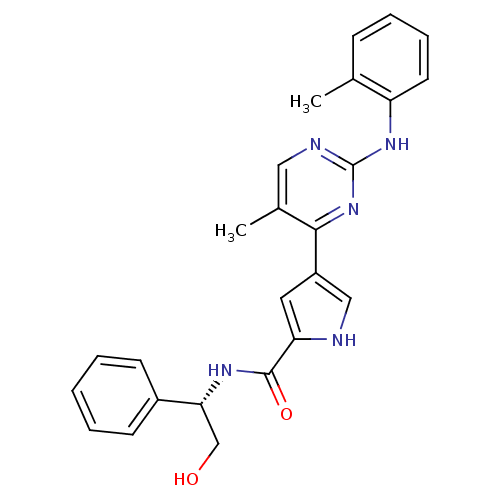
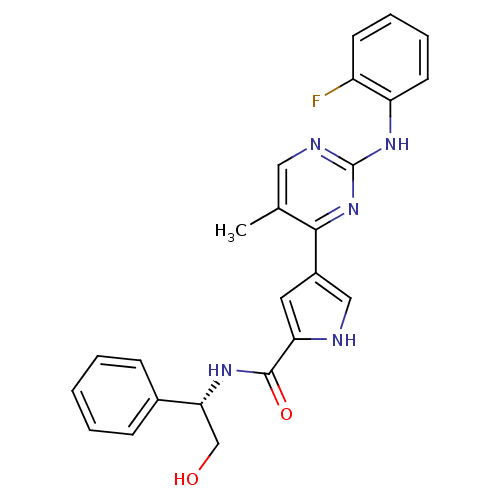
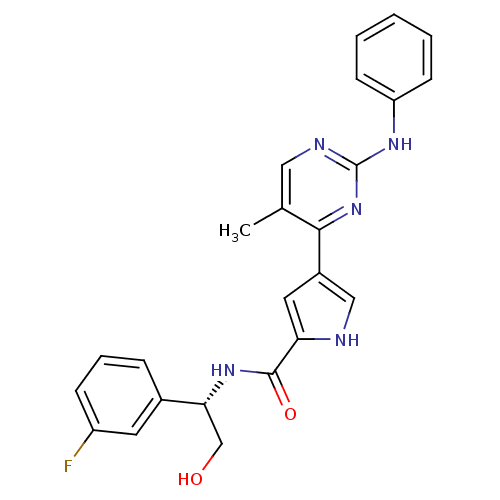
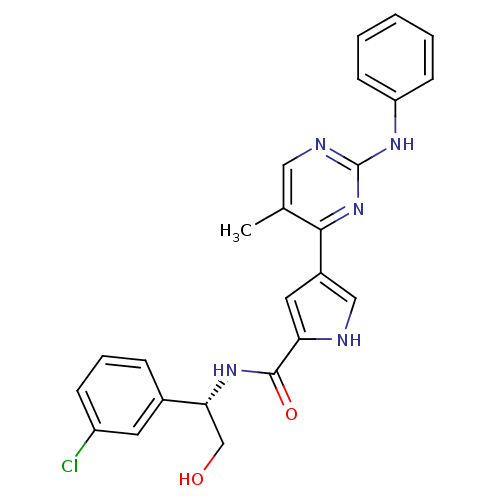
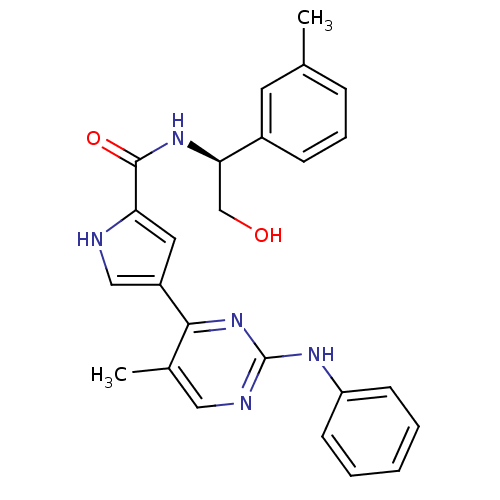
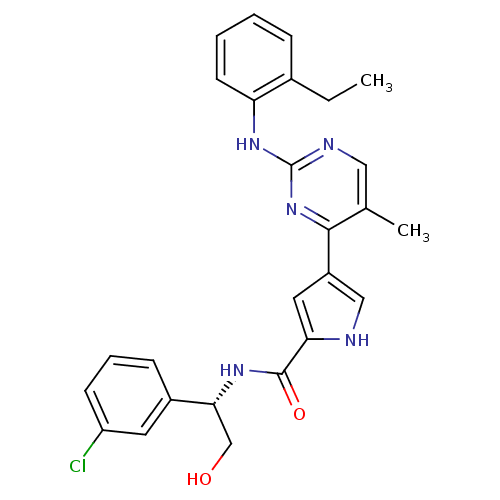
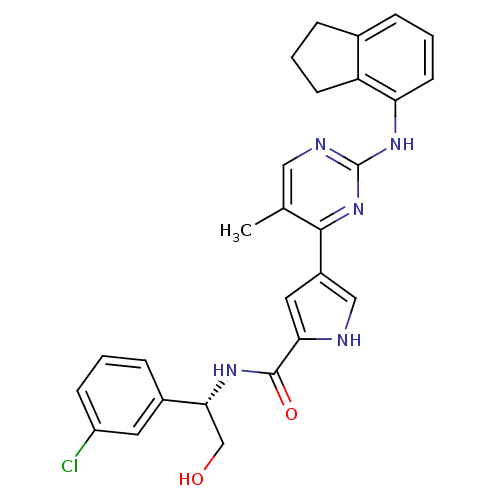
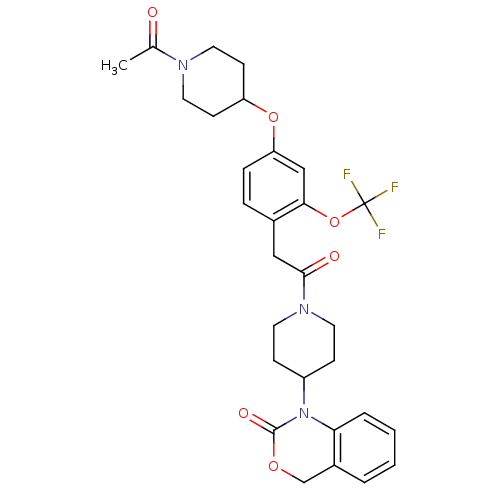
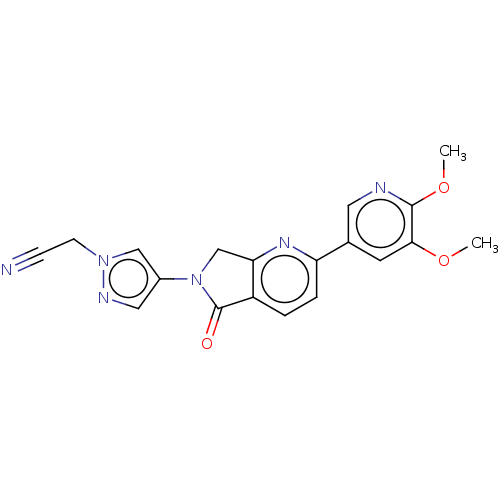
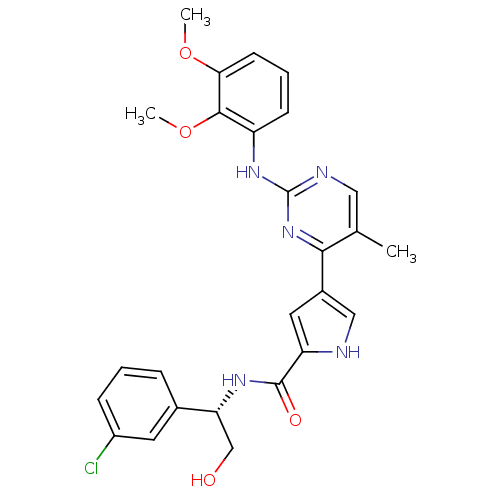
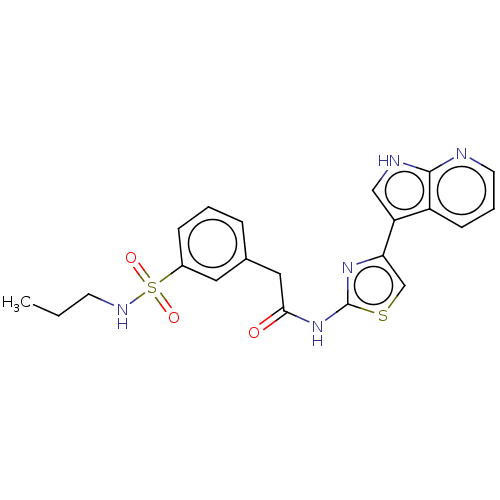
Affinity DataKi: 0.600nMAssay Description:Binding affinity for rat uterine oxytocin receptor (rOTr)More data for this Ligand-Target Pair
Affinity DataKi: 0.710nMAssay Description:Binding affinity for rat uterine oxytocin receptor (rOTr)More data for this Ligand-Target Pair
Affinity DataKi: 0.860nMAssay Description:Binding affinity for human oxytocin receptorMore data for this Ligand-Target Pair
Affinity DataKi: <1nMAssay Description:Inhibition of ROCK1 (6 to 553 residues) (unknown origin) using Lys-Lys-Arg-Asn-Arg-Thr-Leu-Ser-Val as substrate in presence of ATP by spectrophotomet...More data for this Ligand-Target Pair
Affinity DataKi: <1nMAssay Description:Inhibition of ROCK1 (6 to 553 residues) (unknown origin) using Lys-Lys-Arg-Asn-Arg-Thr-Leu-Ser-Val as substrate in presence of ATP by spectrophotomet...More data for this Ligand-Target Pair
Affinity DataKi: 1nMAssay Description:Binding affinity for human oxytocin receptorMore data for this Ligand-Target Pair
Affinity DataKi: 1.20nMAssay Description:Binding affinity for rat uterine oxytocin receptor (rOTr)More data for this Ligand-Target Pair
Affinity DataKi: 1.40nMAssay Description:Binding affinity for human oxytocin receptorMore data for this Ligand-Target Pair
Affinity DataKi: 1.40nMAssay Description:Binding affinity for human oxytocin receptorMore data for this Ligand-Target Pair
TargetSigma non-opioid intracellular receptor 1(Homo sapiens (Human))
Qbi Covid-19 Research Group (Qcrg)
Curated by ChEMBL
Qbi Covid-19 Research Group (Qcrg)
Curated by ChEMBL
Affinity DataKi: 2nMAssay Description:Displacement of [3H]-pentazocin from the Sigma1 receptorMore data for this Ligand-Target Pair
Affinity DataKi: <2nMAssay Description:A coupled spectrophotometric assay was used in which ADP generated by kinase was converted to ATP with the concomitant production of pyruvate from PE...More data for this Ligand-Target Pair
TargetSigma non-opioid intracellular receptor 1(Homo sapiens (Human))
Qbi Covid-19 Research Group (Qcrg)
Curated by ChEMBL
Qbi Covid-19 Research Group (Qcrg)
Curated by ChEMBL
Affinity DataKi: 2nMAssay Description:Displacement of [3H]-pentazocin from the Sigma1 receptorMore data for this Ligand-Target Pair
Affinity DataKi: <2nMAssay Description:A coupled spectrophotometric assay was used in which ADP generated by kinase was converted to ATP with the concomitant production of pyruvate from PE...More data for this Ligand-Target Pair
Affinity DataKi: <2nM ΔG°: <-50.5kJ/molepH: 7.5 T: 2°CAssay Description:A coupled spectrophotometric assay was used in which ADP generated by kinase was converted to ATP with the concomitant production of pyruvate from PE...More data for this Ligand-Target Pair
Affinity DataKi: <2nM ΔG°: <-50.5kJ/molepH: 7.5 T: 2°CAssay Description:A coupled spectrophotometric assay was used in which ADP generated by kinase was converted to ATP with the concomitant production of pyruvate from PE...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform(Homo sapiens (Human))
Vertex Pharmaceuticals
Curated by ChEMBL
Vertex Pharmaceuticals
Curated by ChEMBL
Affinity DataKi: 2nMAssay Description:Inhibition of PI3Kgamma (unknown origin) using PIP2 as substrate after 15 mins in presence of [33P-ATP] by liquid scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 2nMAssay Description:Inhibition of ROCK1 (6 to 553 residues) (unknown origin) using Lys-Lys-Arg-Asn-Arg-Thr-Leu-Ser-Val as substrate in presence of ATP by spectrophotomet...More data for this Ligand-Target Pair
Affinity DataKi: 2nMAssay Description:Inhibition of ROCK1 (6 to 553 residues) (unknown origin) using Lys-Lys-Arg-Asn-Arg-Thr-Leu-Ser-Val as substrate in presence of ATP by spectrophotomet...More data for this Ligand-Target Pair
Affinity DataKi: 2nMAssay Description:Inhibition of ROCK1 (6 to 553 residues) (unknown origin) using Lys-Lys-Arg-Asn-Arg-Thr-Leu-Ser-Val as substrate in presence of ATP by spectrophotomet...More data for this Ligand-Target Pair
Affinity DataKi: 2nMAssay Description:Inhibition of ROCK1 (6 to 553 residues) (unknown origin) using Lys-Lys-Arg-Asn-Arg-Thr-Leu-Ser-Val as substrate in presence of ATP by spectrophotomet...More data for this Ligand-Target Pair
Affinity DataKi: 2nMAssay Description:Inhibition of ROCK1 (6 to 553 residues) (unknown origin) using Lys-Lys-Arg-Asn-Arg-Thr-Leu-Ser-Val as substrate in presence of ATP by spectrophotomet...More data for this Ligand-Target Pair
Affinity DataKi: 2nMAssay Description:Binding affinity towards rat liver Vasopressin V1a receptor by using functional assayMore data for this Ligand-Target Pair
Affinity DataKi: 2nMAssay Description:A coupled spectrophotometric assay was used in which ADP generated by ERK2 was converted to ATP by pyruvate kinase with the production of pyruvate fr...More data for this Ligand-Target Pair
Affinity DataKi: <2nM ΔG°: <-50.5kJ/molepH: 7.5 T: 2°CAssay Description:A coupled spectrophotometric assay was used in which ADP generated by kinase was converted to ATP with the concomitant production of pyruvate from PE...More data for this Ligand-Target Pair
Affinity DataKi: <2nM ΔG°: <-50.5kJ/molepH: 7.5 T: 2°CAssay Description:A coupled spectrophotometric assay was used in which ADP generated by kinase was converted to ATP with the concomitant production of pyruvate from PE...More data for this Ligand-Target Pair
Affinity DataKi: <2nM ΔG°: <-50.5kJ/molepH: 7.5 T: 2°CAssay Description:A coupled spectrophotometric assay was used in which ADP generated by kinase was converted to ATP with the concomitant production of pyruvate from PE...More data for this Ligand-Target Pair
Affinity DataKi: <2nM ΔG°: <-50.5kJ/molepH: 7.5 T: 2°CAssay Description:A coupled spectrophotometric assay was used in which ADP generated by kinase was converted to ATP with the concomitant production of pyruvate from PE...More data for this Ligand-Target Pair
Affinity DataKi: <2nM ΔG°: <-50.5kJ/molepH: 7.5 T: 2°CAssay Description:A coupled spectrophotometric assay was used in which ADP generated by kinase was converted to ATP with the concomitant production of pyruvate from PE...More data for this Ligand-Target Pair
Affinity DataKi: <2nM ΔG°: <-50.5kJ/molepH: 7.5 T: 2°CAssay Description:A coupled spectrophotometric assay was used in which ADP generated by kinase was converted to ATP with the concomitant production of pyruvate from PE...More data for this Ligand-Target Pair
Affinity DataKi: <2nM ΔG°: <-50.5kJ/molepH: 7.5 T: 2°CAssay Description:A coupled spectrophotometric assay was used in which ADP generated by kinase was converted to ATP with the concomitant production of pyruvate from PE...More data for this Ligand-Target Pair
Affinity DataKi: <2nM ΔG°: <-50.5kJ/molepH: 7.5 T: 2°CAssay Description:A coupled spectrophotometric assay was used in which ADP generated by kinase was converted to ATP with the concomitant production of pyruvate from PE...More data for this Ligand-Target Pair
Affinity DataKi: <2nM ΔG°: <-50.5kJ/molepH: 7.5 T: 2°CAssay Description:A coupled spectrophotometric assay was used in which ADP generated by kinase was converted to ATP with the concomitant production of pyruvate from PE...More data for this Ligand-Target Pair
Affinity DataKi: <2nM ΔG°: <-50.5kJ/molepH: 7.5 T: 2°CAssay Description:A coupled spectrophotometric assay was used in which ADP generated by kinase was converted to ATP with the concomitant production of pyruvate from PE...More data for this Ligand-Target Pair
Affinity DataKi: <2nM ΔG°: <-50.5kJ/molepH: 7.5 T: 2°CAssay Description:A coupled spectrophotometric assay was used in which ADP generated by kinase was converted to ATP with the concomitant production of pyruvate from PE...More data for this Ligand-Target Pair
Affinity DataKi: <2nM ΔG°: <-50.5kJ/molepH: 7.5 T: 2°CAssay Description:A coupled spectrophotometric assay was used in which ADP generated by kinase was converted to ATP with the concomitant production of pyruvate from PE...More data for this Ligand-Target Pair
Affinity DataKi: 2.10nMAssay Description:Binding affinity for rat uterine oxytocin receptor (rOTr)More data for this Ligand-Target Pair
Affinity DataKi: 2.40nMAssay Description:Binding affinity for human oxytocin receptorMore data for this Ligand-Target Pair
TargetSigma intracellular receptor 2(Homo sapiens (Human))
Qbi Covid-19 Research Group (Qcrg)
Curated by ChEMBL
Qbi Covid-19 Research Group (Qcrg)
Curated by ChEMBL
Affinity DataKi: 2.5nMAssay Description:Displacement of [3H]-DTG from the Sigma2 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 2.80nMAssay Description:Binding affinity for rat uterine oxytocin receptor (rOTr)More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform(Homo sapiens (Human))
Vertex Pharmaceuticals
Curated by ChEMBL
Vertex Pharmaceuticals
Curated by ChEMBL
Affinity DataKi: 3nMAssay Description:Inhibition of PI3Kgamma (unknown origin) using PIP2 as substrate measured after 15 mins in presence of [33P]-ATP by liquid scintillation counting met...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform(Homo sapiens (Human))
Vertex Pharmaceuticals
Curated by ChEMBL
Vertex Pharmaceuticals
Curated by ChEMBL
Affinity DataKi: 3nMAssay Description:Inhibition of PI3Kgamma (unknown origin) using PIP2 as substrate measured after 15 mins in presence of [33P]-ATP by liquid scintillation counting met...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform(Homo sapiens (Human))
Vertex Pharmaceuticals
Curated by ChEMBL
Vertex Pharmaceuticals
Curated by ChEMBL
Affinity DataKi: 3nMAssay Description:Inhibition of PI3Kgamma (unknown origin) using PIP2 as substrate measured after 15 mins in presence of [33P]-ATP by liquid scintillation counting met...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform(Homo sapiens (Human))
Vertex Pharmaceuticals
Curated by ChEMBL
Vertex Pharmaceuticals
Curated by ChEMBL
Affinity DataKi: 3nMAssay Description:Inhibition of PI3Kgamma (unknown origin) using PIP2 as substrate after 15 mins in presence of [33P-ATP] by liquid scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 3nMAssay Description:Inhibition of ROCK1 (6 to 553 residues) (unknown origin) using Lys-Lys-Arg-Asn-Arg-Thr-Leu-Ser-Val as substrate in presence of ATP by spectrophotomet...More data for this Ligand-Target Pair
Affinity DataKi: 3nM ΔG°: -49.5kJ/molepH: 7.5 T: 2°CAssay Description:A coupled spectrophotometric assay was used in which ADP generated by kinase was converted to ATP with the concomitant production of pyruvate from PE...More data for this Ligand-Target Pair
Affinity DataKi: 3nMAssay Description:Inhibition of ROCK1 (6 to 553 residues) (unknown origin) using Lys-Lys-Arg-Asn-Arg-Thr-Leu-Ser-Val as substrate in presence of ATP by spectrophotomet...More data for this Ligand-Target Pair
Affinity DataKi: 3nMAssay Description:Inhibition of ROCK1 (6 to 553 residues) (unknown origin) using Lys-Lys-Arg-Asn-Arg-Thr-Leu-Ser-Val as substrate in presence of ATP by spectrophotomet...More data for this Ligand-Target Pair
Affinity DataKi: 3nMAssay Description:Inhibition of ROCK1 (6 to 553 residues) (unknown origin) using Lys-Lys-Arg-Asn-Arg-Thr-Leu-Ser-Val as substrate in presence of ATP by spectrophotomet...More data for this Ligand-Target Pair

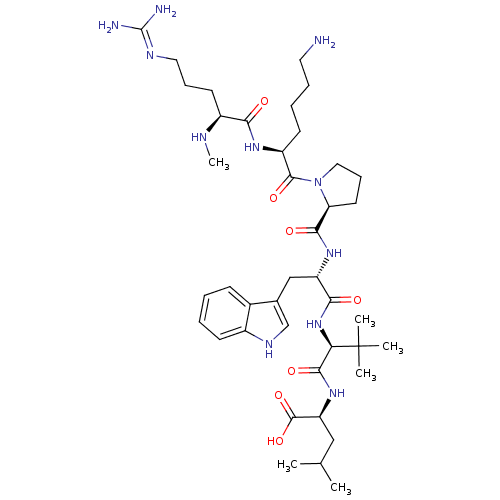

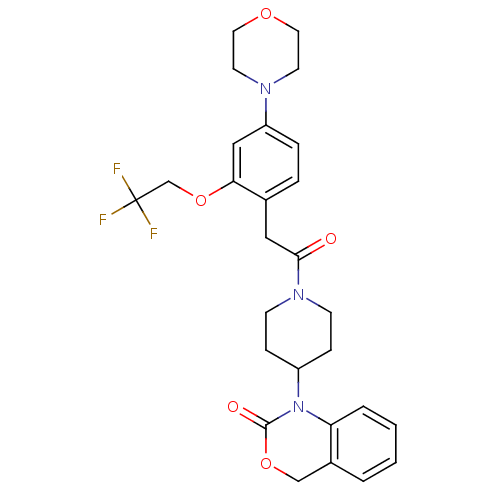
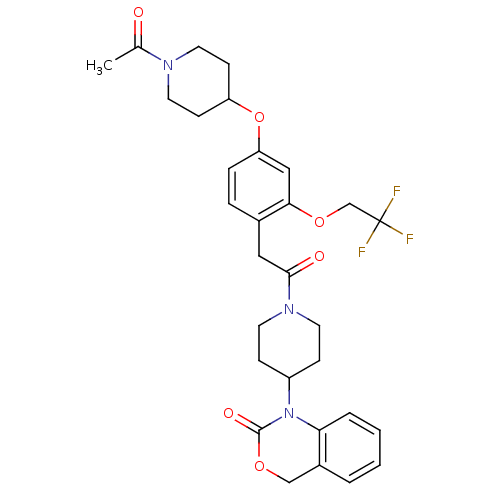
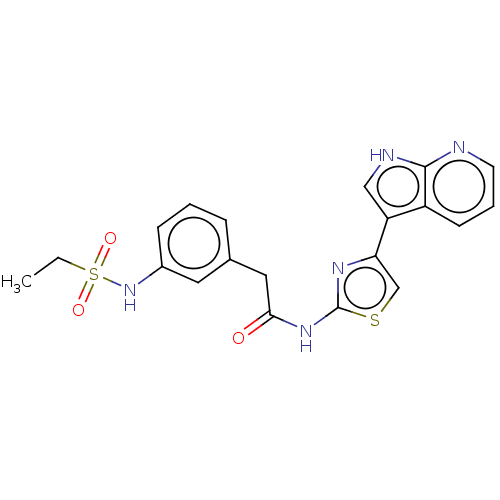
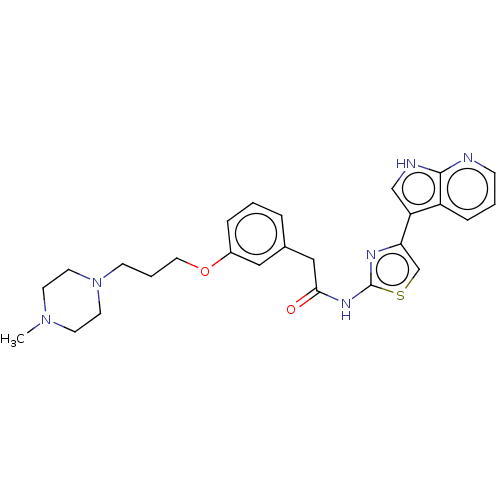
 3D Structure (crystal)
3D Structure (crystal)