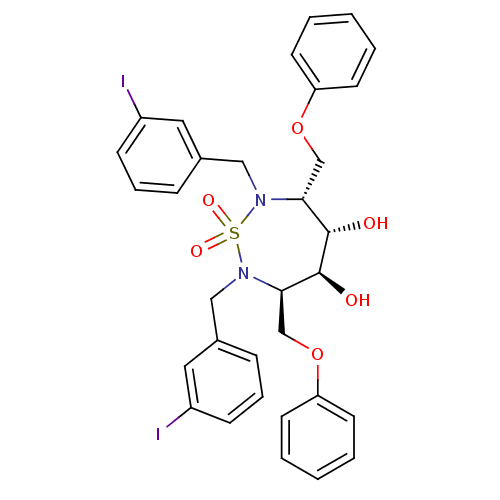
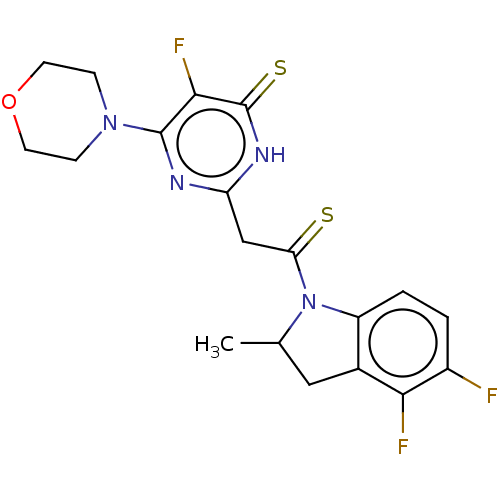
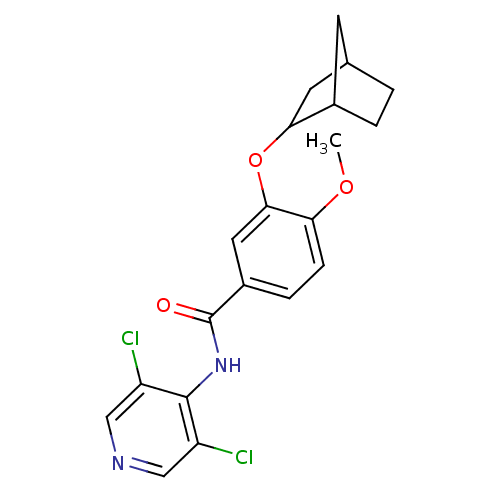
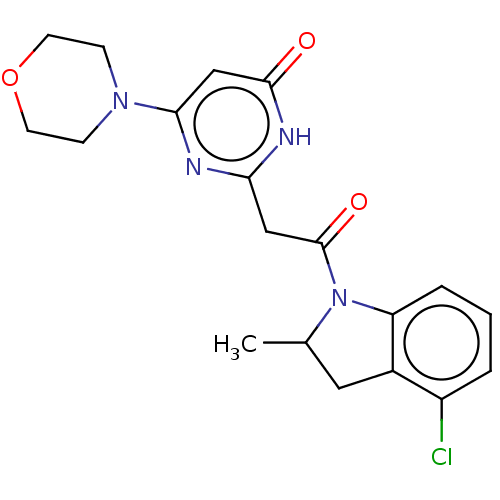
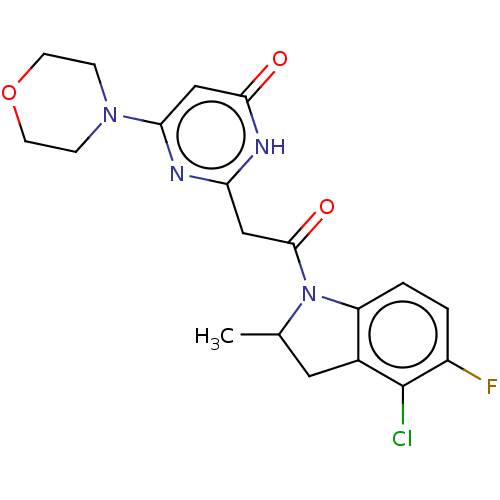
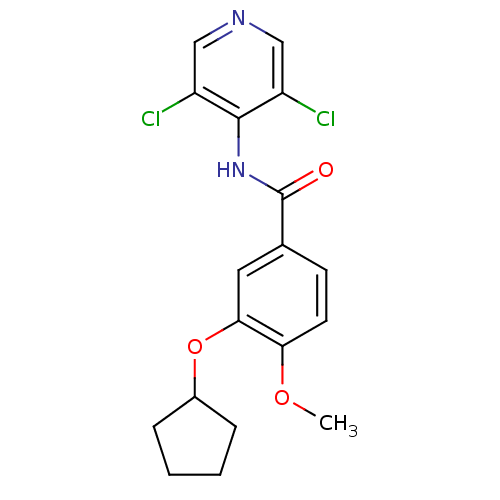
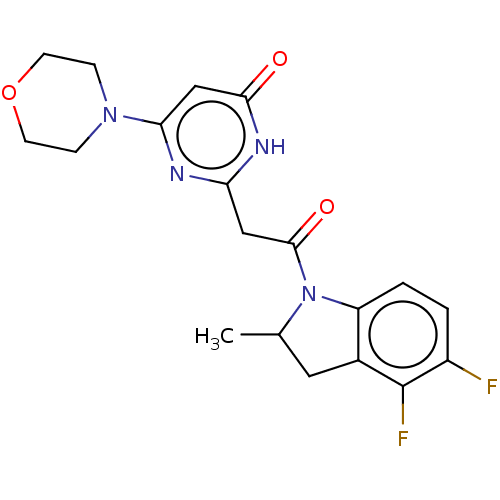
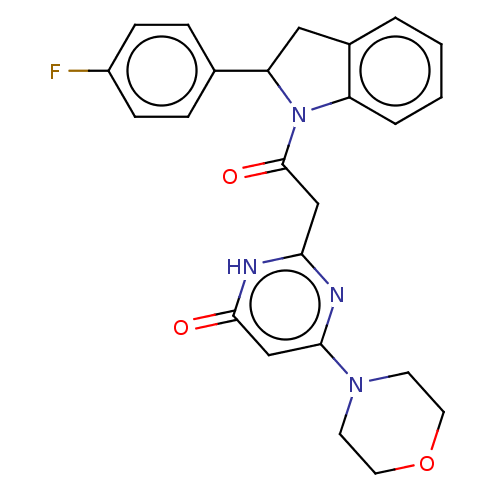
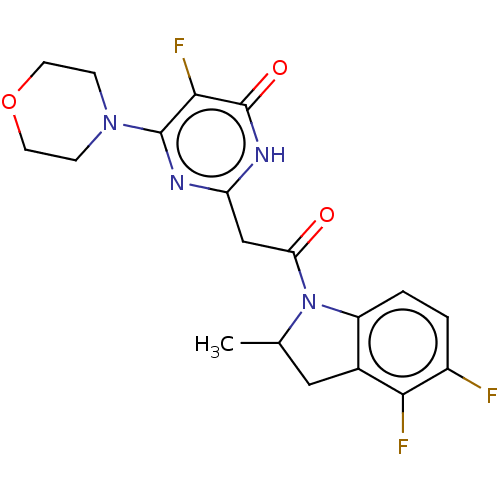
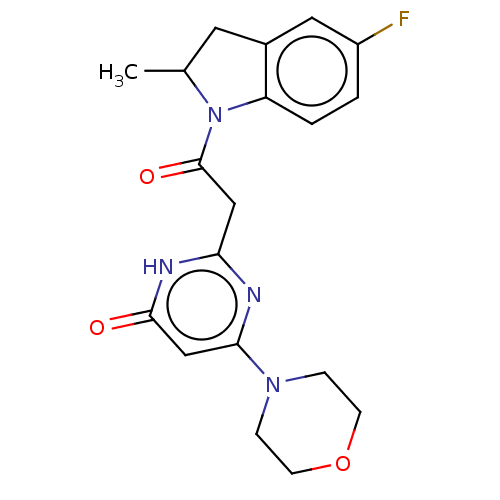
Affinity DataKi: 3.10nM ΔG°: -49.4kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
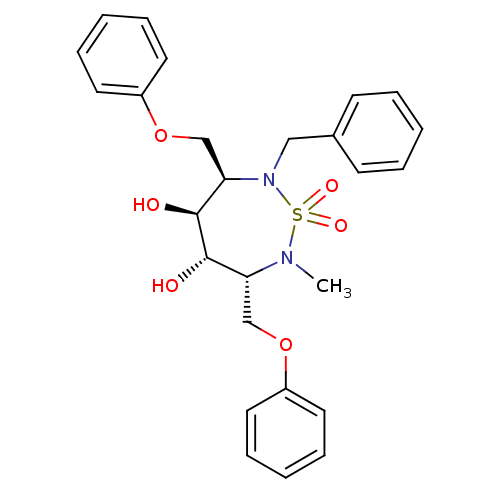
Affinity DataKi: 3.40nM ΔG°: -49.1kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
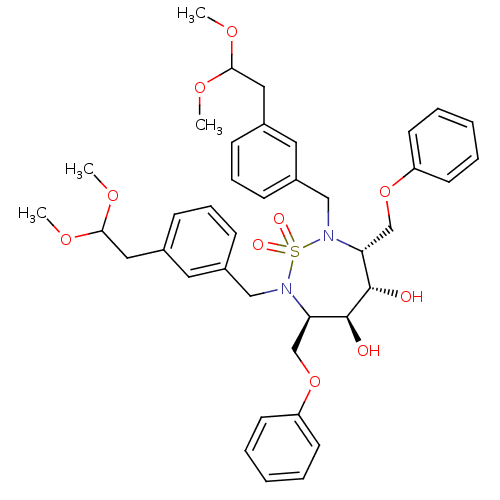
Affinity DataKi: 7.30nM ΔG°: -47.2kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
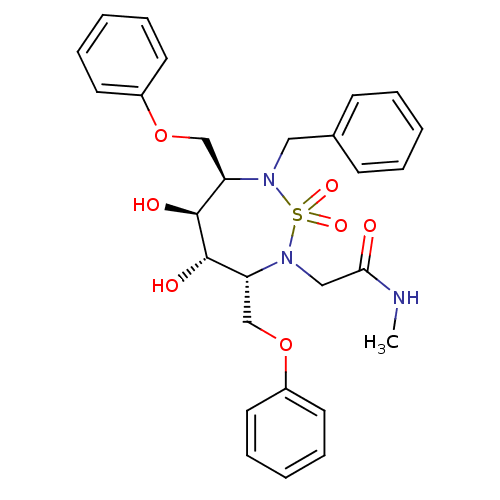
Affinity DataKi: 11nM ΔG°: -46.2kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
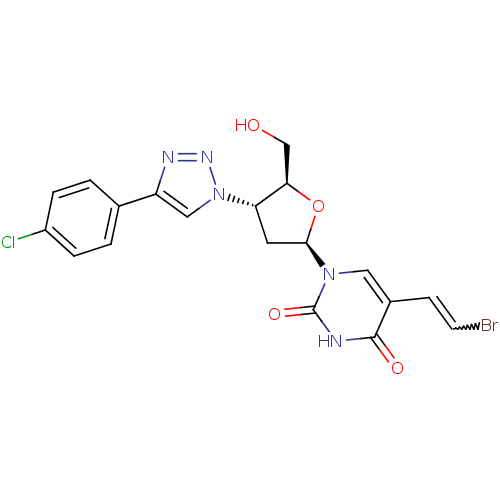
Affinity DataKi: 12nMAssay Description:Competitive inhibition of human recombinant mitochondrial thymidine kinase 2 using ATP as substrate by Lineweaver-Burk plottingMore data for this Ligand-Target Pair
Affinity DataKi: 23nM ΔG°: -44.3kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
Affinity DataKi: 39nM ΔG°: -43.0kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
Affinity DataKi: 43nM ΔG°: -42.8kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
Affinity DataKi: 59nM ΔG°: -42.0kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
Affinity DataKi: 63nM ΔG°: -41.8kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
Affinity DataKi: 84nM ΔG°: -41.1kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
Affinity DataKi: 86nM ΔG°: -41.0kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
Affinity DataKi: 93nM ΔG°: -40.8kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
Affinity DataKi: 140nM ΔG°: -39.8kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
Affinity DataKi: 150nMAssay Description:Inhibition of thymidine kinase 2More data for this Ligand-Target Pair
Affinity DataKi: 350nM ΔG°: -37.5kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
Affinity DataKi: 410nMAssay Description:Uncompetitive inhibition of human recombinant mitochondrial thymidine kinase 2 using thymidine as substrate by Lineweaver-Burk plottingMore data for this Ligand-Target Pair
Affinity DataKi: 510nM ΔG°: -36.5kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
Affinity DataKi: 510nM ΔG°: -36.5kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
Affinity DataKi: 540nM ΔG°: -36.4kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
Affinity DataKi: 710nM ΔG°: -35.7kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
Affinity DataKi: 920nM ΔG°: -35.0kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
Affinity DataKi: 1.40E+3nM ΔG°: -34.0kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
Affinity DataKi: 1.80E+3nM ΔG°: -33.3kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
Affinity DataKi: 2.20E+3nM ΔG°: -32.8kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
Affinity DataKi: 2.20E+3nM ΔG°: -32.8kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
Affinity DataKi: 6.00E+3nM ΔG°: -30.3kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
Affinity DataKi: 1.04E+4nMAssay Description:Inhibition of thymidine kinase 2More data for this Ligand-Target Pair
Affinity DataKi: 2.50E+4nM ΔG°: -26.7kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
Affinity DataIC50: 1nMAssay Description:This test is based on measuring the expression of the AKT protein phosphorylated on serine 473 (P-AKT-S473), in the PC3 human prostate carcinoma line...More data for this Ligand-Target Pair
Affinity DataIC50: 1nMAssay Description:This test is based on measurement of the expression of AKT protein phosphorylated on serine 473 (P-AKT-S473) in the PC3 human prostate carcinoma line...More data for this Ligand-Target Pair
Affinity DataIC50: 1nMAssay Description:Inhibitory potency against pig aortic PDE IVMore data for this Ligand-Target Pair
Affinity DataIC50: 1nMAssay Description:This test is based on measurement of the expression of AKT protein phosphorylated on serine 473 (P-AKT-S473) in the PC3 human prostate carcinoma line...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta isoform(Homo sapiens (Human))
Sanofi
Curated by ChEMBL
Sanofi
Curated by ChEMBL
Affinity DataIC50: 1nMAssay Description:Inhibition of N-terminal ploy-His-tagged human PI3Kbeta expressed in baculovirus-infected sf21 cells using PI(4,5)P2 as substrate after 15 mins by HT...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta isoform(Homo sapiens (Human))
Sanofi
Curated by ChEMBL
Sanofi
Curated by ChEMBL
Affinity DataIC50: 1nMAssay Description:Inhibition of N-terminal ploy-His-tagged human PI3Kbeta expressed in baculovirus-infected sf21 cells using PI(4,5)P2 as substrate after 15 mins by HT...More data for this Ligand-Target Pair
Affinity DataIC50: 1nMAssay Description:This test is based on measurement of the expression of AKT protein phosphorylated on serine 473 (P-AKT-S473) in the PC3 human prostate carcinoma line...More data for this Ligand-Target Pair
Affinity DataIC50: 1nMAssay Description:Inhibitory potency against pig aortic PDE IVMore data for this Ligand-Target Pair
Affinity DataIC50: 1nMAssay Description:Inhibitory potency against pig aortic PDE IVMore data for this Ligand-Target Pair
Affinity DataIC50: 1nMAssay Description:This test is based on measurement of the expression of AKT protein phosphorylated on serine 473 (P-AKT-S473) in the PC3 human prostate carcinoma line...More data for this Ligand-Target Pair
Affinity DataIC50: 1.60nMAssay Description:Steroid binding against the rat thymus glucocorticoid receptorMore data for this Ligand-Target Pair
Affinity DataIC50: 1.70nMAssay Description:Steroid binding against the rat thymus glucocorticoid receptorMore data for this Ligand-Target Pair
Affinity DataIC50: 1.80nMAssay Description:Steroid binding against the rat thymus glucocorticoid receptorMore data for this Ligand-Target Pair
Affinity DataIC50: 2nMAssay Description:This test is based on measurement of the expression of AKT protein phosphorylated on serine 473 (P-AKT-S473) in the PC3 human prostate carcinoma line...More data for this Ligand-Target Pair
Affinity DataIC50: 2nMAssay Description:This test is based on measurement of the expression of AKT protein phosphorylated on serine 473 (P-AKT-S473) in the PC3 human prostate carcinoma line...More data for this Ligand-Target Pair
Affinity DataIC50: 2nMAssay Description:This test is based on measurement of the expression of AKT protein phosphorylated on serine 473 (P-AKT-S473) in the PC3 human prostate carcinoma line...More data for this Ligand-Target Pair
Affinity DataIC50: 2nMAssay Description:This test is based on measurement of the expression of AKT protein phosphorylated on serine 473 (P-AKT-S473) in the PC3 human prostate carcinoma line...More data for this Ligand-Target Pair
Affinity DataIC50: 2nMAssay Description:This test is based on measurement of the expression of AKT protein phosphorylated on serine 473 (P-AKT-S473) in the PC3 human prostate carcinoma line...More data for this Ligand-Target Pair
Affinity DataIC50: 2nMAssay Description:This test is based on measurement of the expression of AKT protein phosphorylated on serine 473 (P-AKT-S473) in the PC3 human prostate carcinoma line...More data for this Ligand-Target Pair



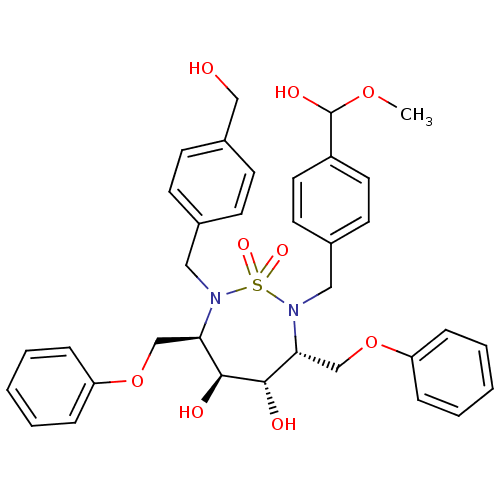
 3D Structure (crystal)
3D Structure (crystal)