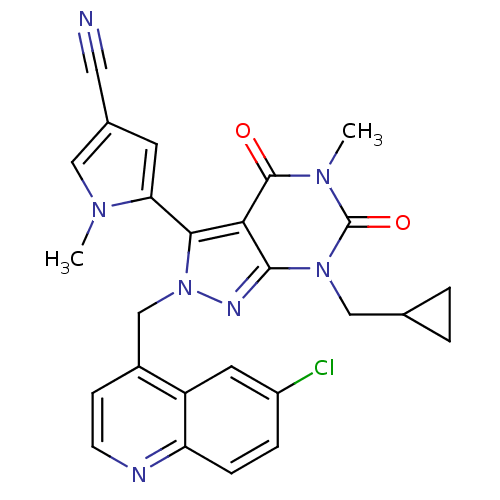
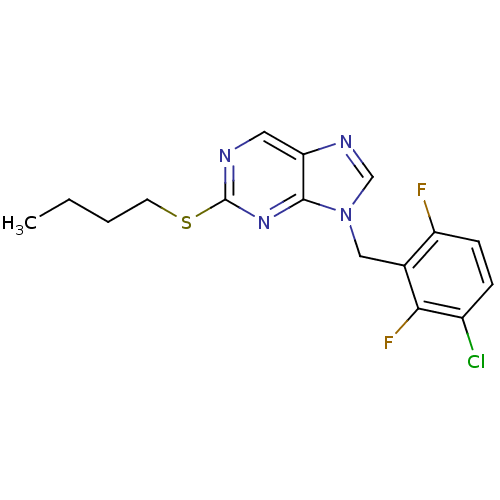
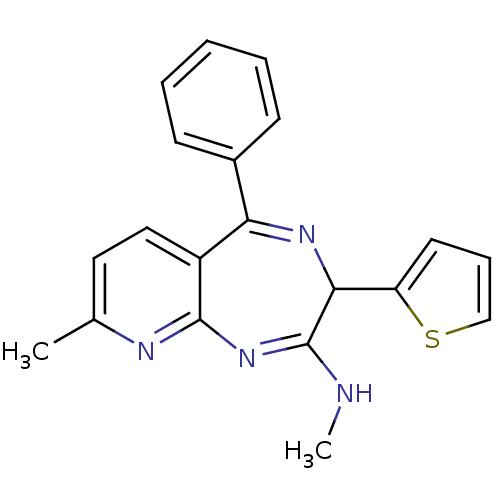
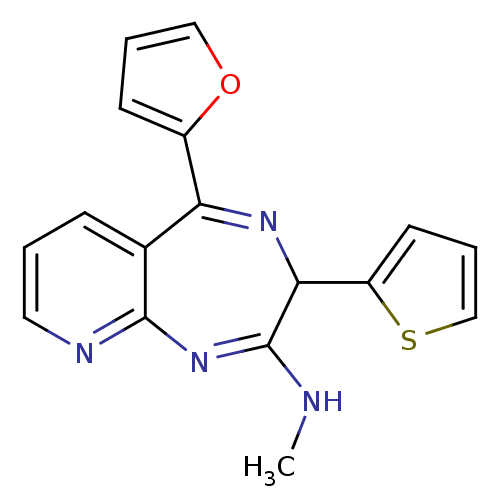
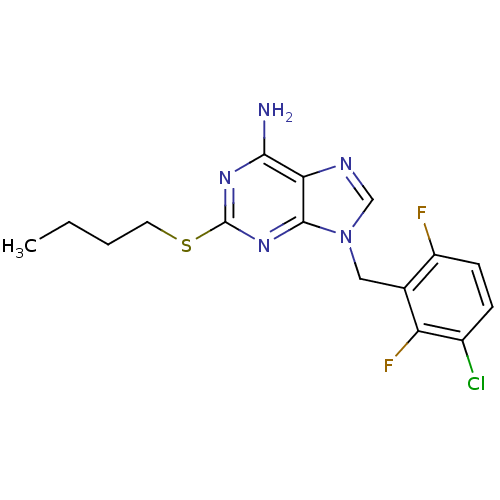
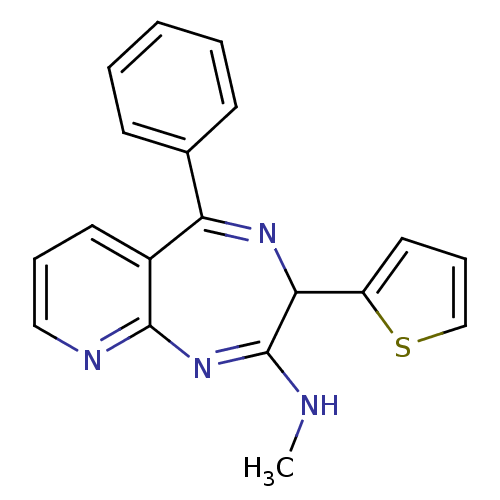
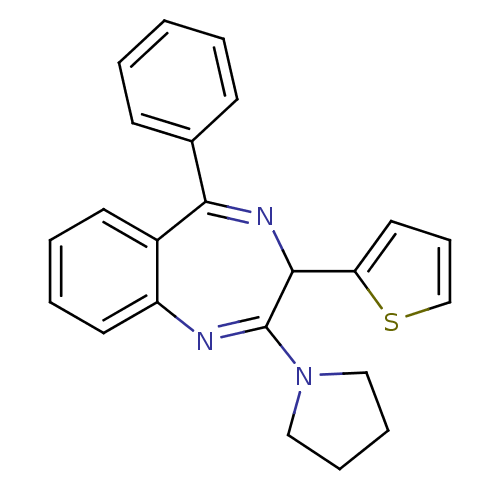
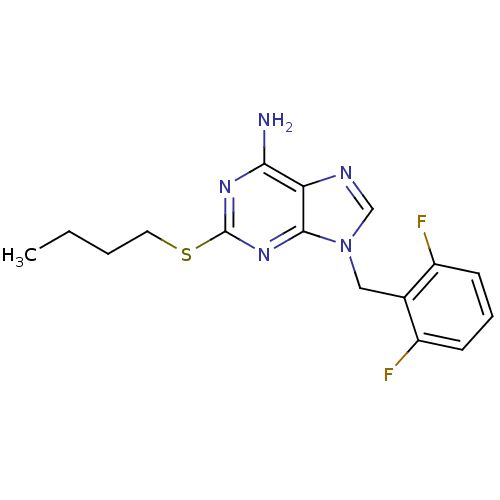
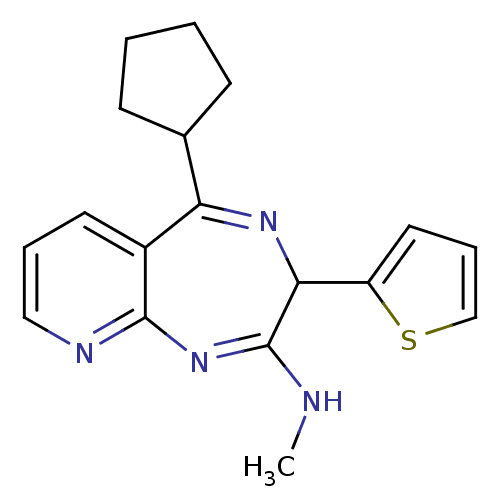
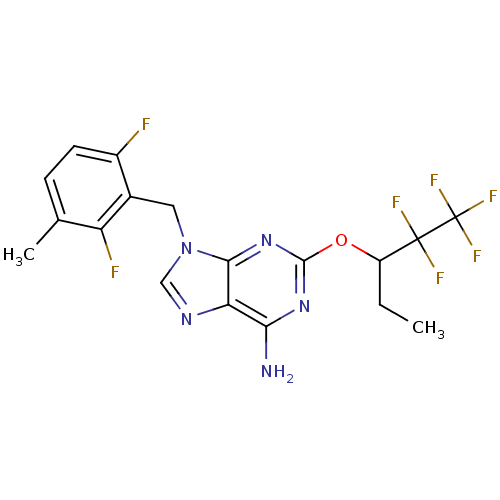
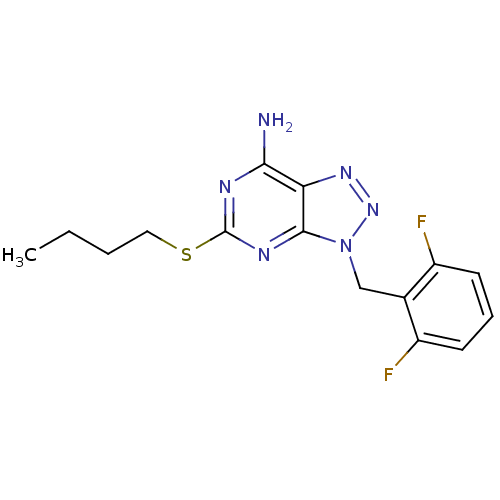
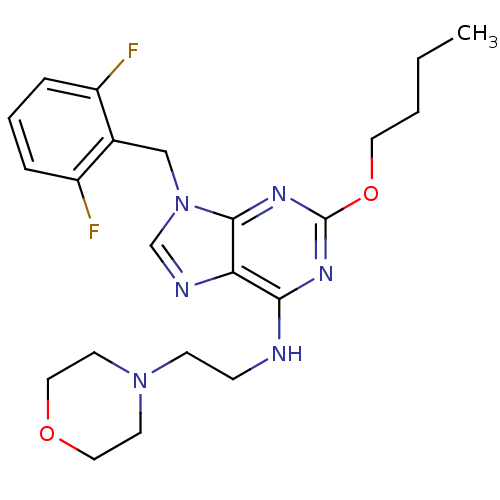
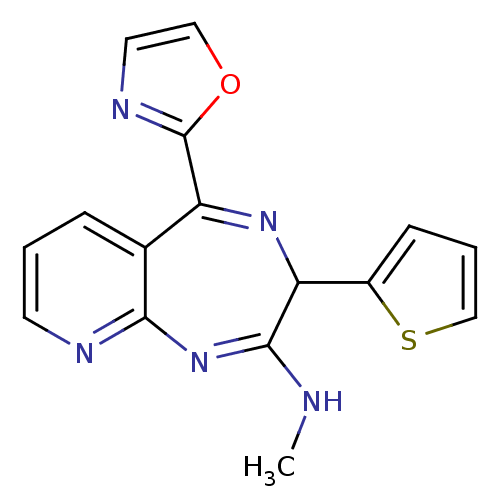
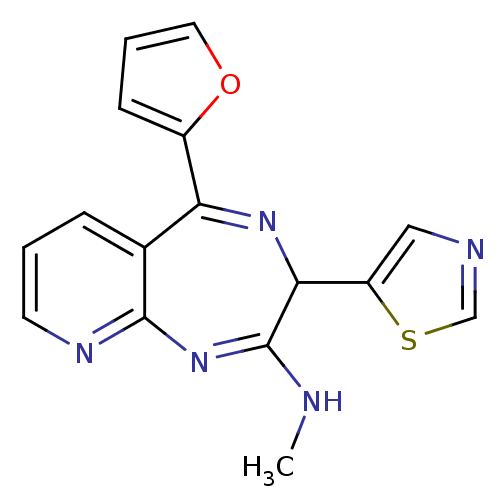
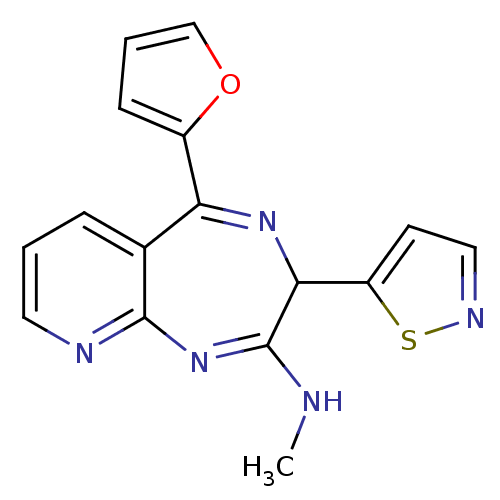
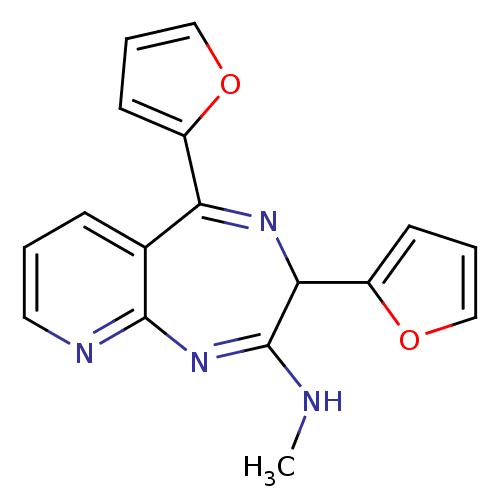
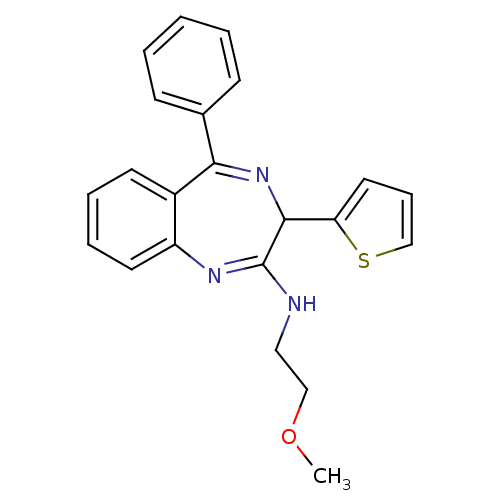
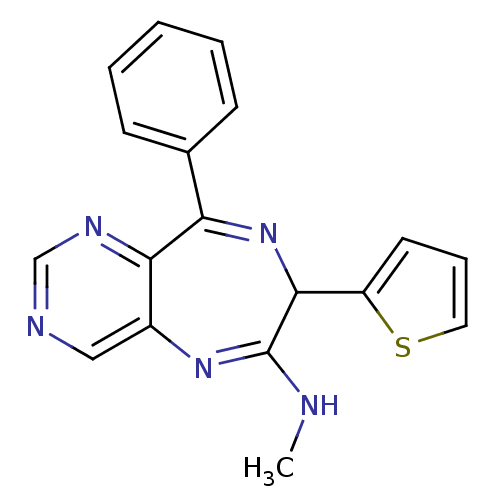
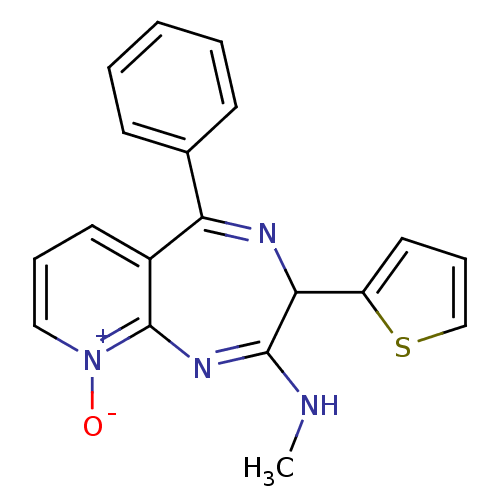
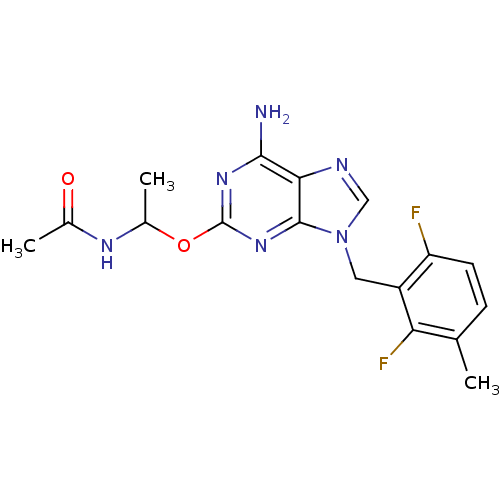
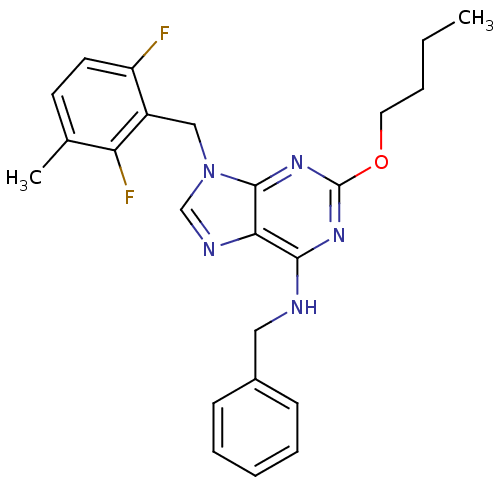
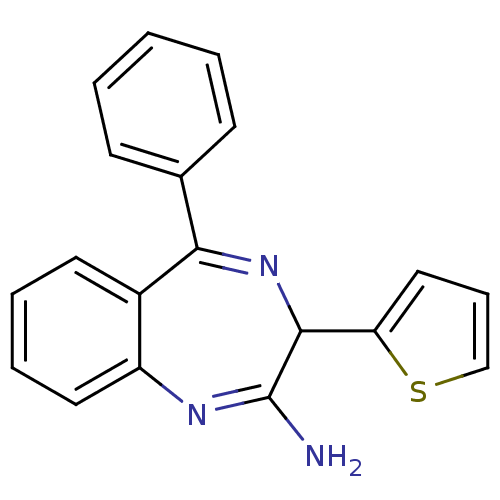
TargetGlutamate racemase(Helicobacter pylori)
Astrazeneca Global Structural Chemistry
Curated by ChEMBL
Astrazeneca Global Structural Chemistry
Curated by ChEMBL
Affinity DataKi: 5.80E+3nMAssay Description:Inhibition of Helicobacter pylori glutamate racemaseMore data for this Ligand-Target Pair
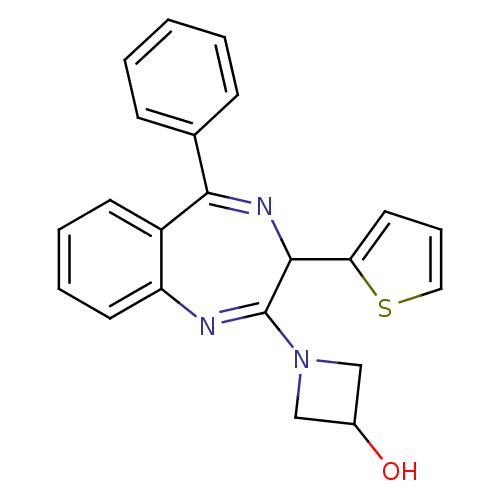
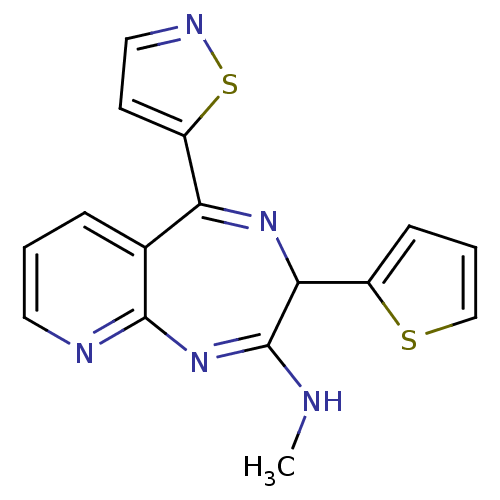
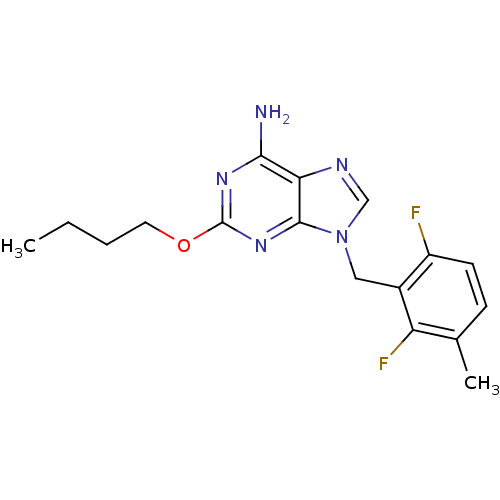
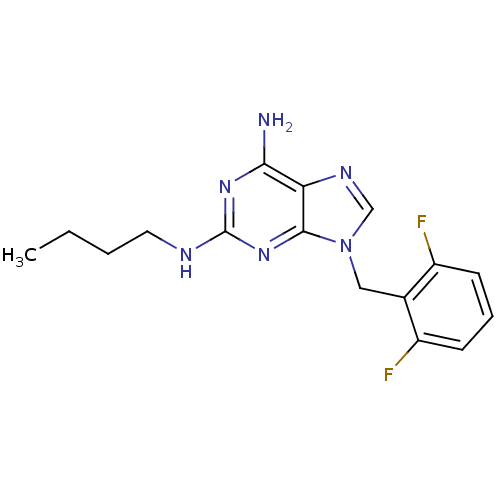
TargetGlutamate racemase(Helicobacter pylori)
Astrazeneca Global Structural Chemistry
Curated by ChEMBL
Astrazeneca Global Structural Chemistry
Curated by ChEMBL
Affinity DataIC50: 25nMAssay Description:Inhibition of Helicobacter pylori glutamate racemaseMore data for this Ligand-Target Pair
Affinity DataIC50: 500nMAssay Description:Inhibition of MurI in wild type Helicobacter pylori J99More data for this Ligand-Target Pair
Affinity DataIC50: 600nMAssay Description:Inhibition of MurI in wild type Helicobacter pylori J99More data for this Ligand-Target Pair
Affinity DataIC50: 700nMAssay Description:Inhibition of MurI in wild type Helicobacter pylori J99More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMAssay Description:Inhibition of Enterococcus faecalis MurIMore data for this Ligand-Target Pair
Affinity DataIC50: 1.10E+3nMAssay Description:Inhibition of MurI in wild type Helicobacter pylori J99More data for this Ligand-Target Pair
Affinity DataIC50: 1.10E+3nMAssay Description:Inhibition of MurI in wild type Helicobacter pylori J99More data for this Ligand-Target Pair
Affinity DataIC50: 1.10E+3nMAssay Description:Inhibition of MurI in wild type Helicobacter pylori J99More data for this Ligand-Target Pair
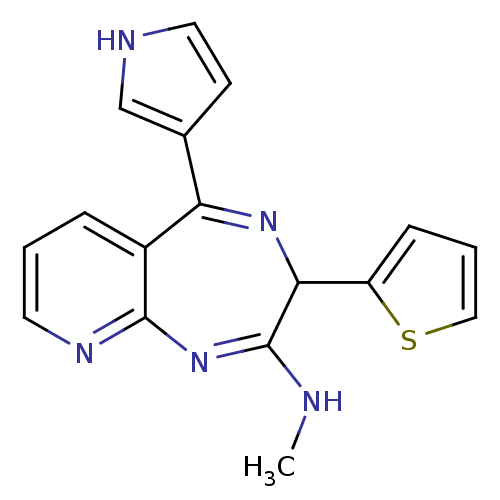
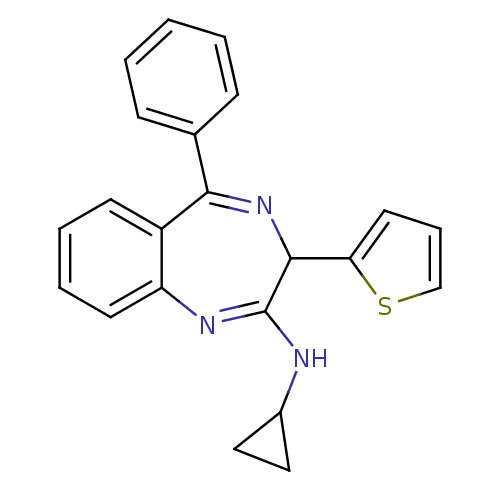
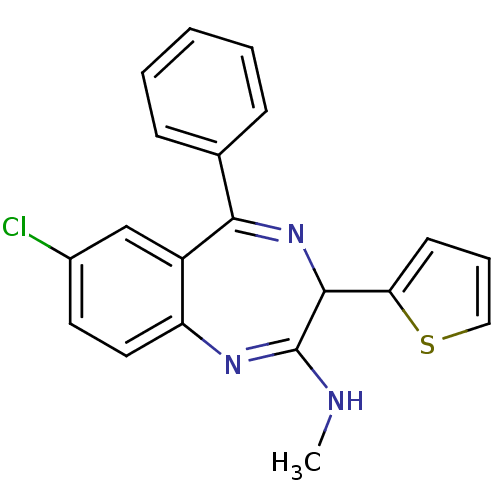
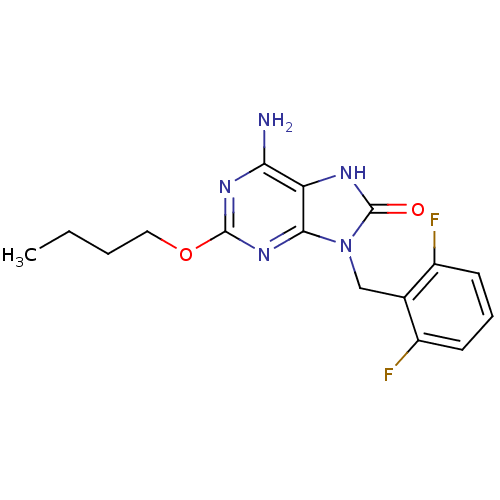
TargetGlutamate racemase(Helicobacter pylori)
Astrazeneca Global Structural Chemistry
Curated by ChEMBL
Astrazeneca Global Structural Chemistry
Curated by ChEMBL
Affinity DataIC50: 1.40E+3nMAssay Description:Inhibition of Helicobacter pylori glutamate racemaseMore data for this Ligand-Target Pair
Affinity DataIC50: 1.40E+3nMAssay Description:Inhibition of Enterococcus faecalis MurIMore data for this Ligand-Target Pair
Affinity DataIC50: 1.70E+3nMAssay Description:Inhibition of MurI in wild type Helicobacter pylori J99More data for this Ligand-Target Pair
Affinity DataIC50: 1.70E+3nMAssay Description:Inhibition of MurI in wild type Helicobacter pylori J99More data for this Ligand-Target Pair
Affinity DataIC50: 1.70E+3nMAssay Description:Inhibition of MurI in wild type Helicobacter pylori J99More data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+3nMAssay Description:Inhibition of MurI in wild type Helicobacter pylori J99More data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+3nMAssay Description:Inhibition of Enterococcus faecalis MurIMore data for this Ligand-Target Pair
Affinity DataIC50: 2.20E+3nMAssay Description:Inhibition of Enterococcus faecalis MurIMore data for this Ligand-Target Pair
Affinity DataIC50: 2.20E+3nMAssay Description:Inhibition of MurI in wild type Helicobacter pylori J99More data for this Ligand-Target Pair
Affinity DataIC50: 2.80E+3nMAssay Description:Inhibition of MurI in wild type Helicobacter pylori J99More data for this Ligand-Target Pair
Affinity DataIC50: 4.00E+3nMAssay Description:Inhibition of MurI in wild type Helicobacter pylori J99More data for this Ligand-Target Pair
Affinity DataIC50: 4.00E+3nMAssay Description:Inhibition of MurI in wild type Helicobacter pylori J99More data for this Ligand-Target Pair
Affinity DataIC50: 4.10E+3nMAssay Description:Inhibition of Enterococcus faecalis MurIMore data for this Ligand-Target Pair
Affinity DataIC50: 4.10E+3nMAssay Description:Inhibition of MurI in wild type Helicobacter pylori J99More data for this Ligand-Target Pair
Affinity DataIC50: 4.30E+3nMAssay Description:Inhibition of Enterococcus faecalis MurIMore data for this Ligand-Target Pair
Affinity DataIC50: 4.30E+3nMAssay Description:Inhibition of Enterococcus faecalis MurIMore data for this Ligand-Target Pair
Affinity DataIC50: 4.90E+3nMAssay Description:Inhibition of MurI in wild type Helicobacter pylori J99More data for this Ligand-Target Pair
Affinity DataIC50: 5.40E+3nMAssay Description:Inhibition of Enterococcus faecalis MurIMore data for this Ligand-Target Pair
Affinity DataIC50: 5.50E+3nMAssay Description:Inhibition of Enterococcus faecalis MurIMore data for this Ligand-Target Pair
Affinity DataIC50: 5.80E+3nMAssay Description:Inhibition of Enterococcus faecalis MurIMore data for this Ligand-Target Pair
Affinity DataIC50: 7.70E+3nMAssay Description:Inhibition of MurI in wild type Helicobacter pylori J99More data for this Ligand-Target Pair
Affinity DataIC50: 8.60E+3nMAssay Description:Inhibition of MurI in wild type Helicobacter pylori J99More data for this Ligand-Target Pair
Affinity DataIC50: 8.70E+3nMAssay Description:Inhibition of MurI in wild type Helicobacter pylori J99More data for this Ligand-Target Pair
Affinity DataIC50: 9.40E+3nMAssay Description:Inhibition of Enterococcus faecalis MurIMore data for this Ligand-Target Pair
Affinity DataIC50: 9.80E+3nMAssay Description:Inhibition of Enterococcus faecalis MurIMore data for this Ligand-Target Pair
Affinity DataIC50: 1.15E+4nMAssay Description:Inhibition of Enterococcus faecalis MurIMore data for this Ligand-Target Pair
Affinity DataIC50: 1.30E+4nMAssay Description:Inhibition of MurI in wild type Helicobacter pylori J99More data for this Ligand-Target Pair
Affinity DataIC50: 1.85E+4nMAssay Description:Inhibition of MurI in wild type Helicobacter pylori J99More data for this Ligand-Target Pair
Affinity DataIC50: 1.89E+4nMAssay Description:Inhibition of MurI in wild type Helicobacter pylori J99More data for this Ligand-Target Pair
Affinity DataIC50: 2.61E+4nMAssay Description:Inhibition of Enterococcus faecalis MurIMore data for this Ligand-Target Pair
Affinity DataIC50: 3.81E+4nMAssay Description:Inhibition of Enterococcus faecalis MurIMore data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+4nMAssay Description:Inhibition of MurI in wild type Helicobacter pylori J99More data for this Ligand-Target Pair
Affinity DataIC50: 6.70E+4nMAssay Description:Inhibition of MurI in wild type Helicobacter pylori J99More data for this Ligand-Target Pair
Affinity DataIC50: 1.67E+5nMAssay Description:Inhibition of Enterococcus faecalis MurIMore data for this Ligand-Target Pair
Affinity DataIC50: >4.00E+5nMAssay Description:Inhibition of Enterococcus faecalis MurIMore data for this Ligand-Target Pair
Affinity DataIC50: >4.00E+5nMAssay Description:Inhibition of Enterococcus faecalis MurIMore data for this Ligand-Target Pair
Affinity DataIC50: 4.00E+5nMAssay Description:Inhibition of MurI in wild type Helicobacter pylori J99More data for this Ligand-Target Pair
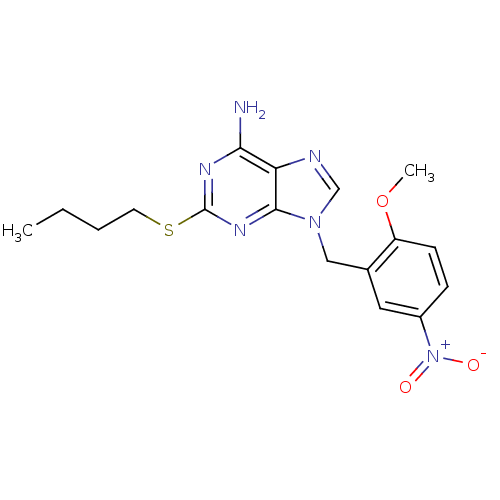
TargetGlutamate racemase(Streptococcus pneumoniae)
Astrazeneca Global Structural Chemistry
Curated by ChEMBL
Astrazeneca Global Structural Chemistry
Curated by ChEMBL
Affinity DataIC50: >4.00E+5nMAssay Description:Inhibition of MurI in Streptococcus pneumoniaeMore data for this Ligand-Target Pair
TargetGlutamate racemase(Streptococcus pneumoniae)
Astrazeneca Global Structural Chemistry
Curated by ChEMBL
Astrazeneca Global Structural Chemistry
Curated by ChEMBL
Affinity DataIC50: >4.00E+5nMAssay Description:Inhibition of MurI in Streptococcus pneumoniaeMore data for this Ligand-Target Pair
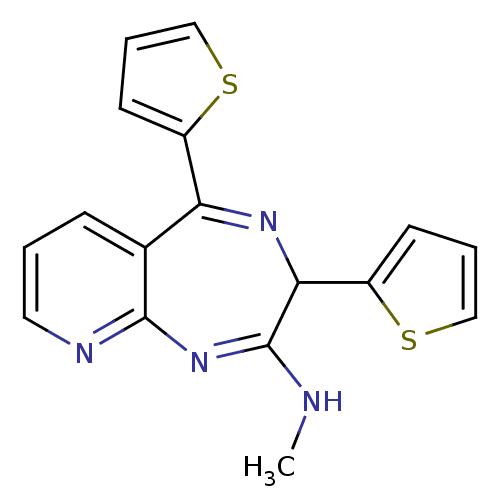
TargetGlutamate racemase(Helicobacter pylori)
Astrazeneca Global Structural Chemistry
Curated by ChEMBL
Astrazeneca Global Structural Chemistry
Curated by ChEMBL
Affinity DataKd: 3.90E+3nMAssay Description:Binding affinity to Helicobacter pylori MurI by protein fluorescence binding assayMore data for this Ligand-Target Pair
TargetGlutamate racemase(Helicobacter pylori)
Astrazeneca Global Structural Chemistry
Curated by ChEMBL
Astrazeneca Global Structural Chemistry
Curated by ChEMBL
Affinity DataKd: 2.90E+3nMAssay Description:Binding affinity to Helicobacter pylori MurI by isothermal titration calorimetryMore data for this Ligand-Target Pair

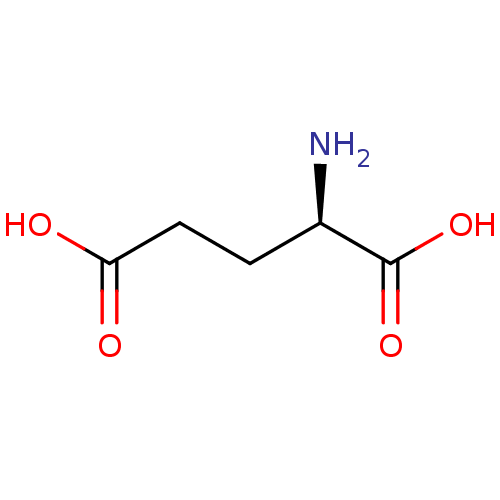
 3D Structure (crystal)
3D Structure (crystal)