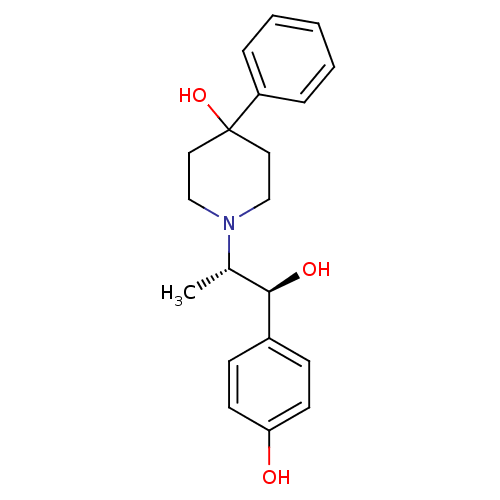
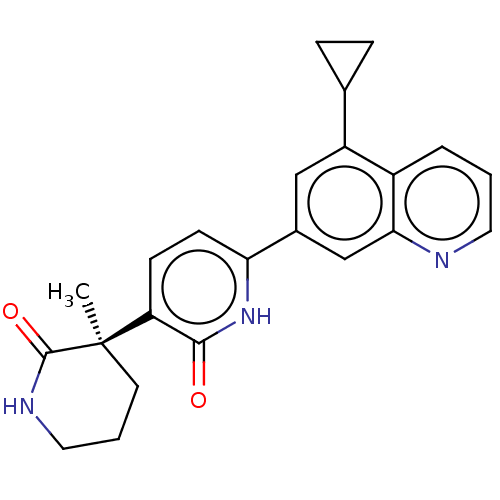
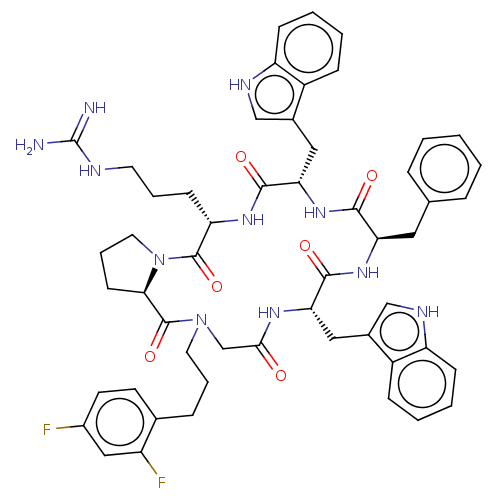
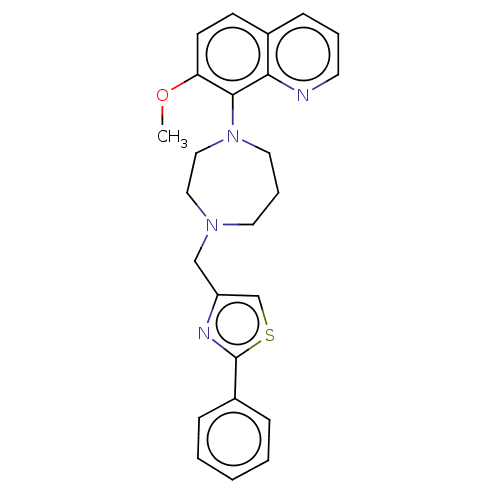
Affinity DataKi: 2nM ΔG°: -49.7kJ/molepH: 7.4 T: 2°CAssay Description:Test compounds were half log serially diluted in 100% DMSO (J. T. Baker #922401). 1 uL of each compound was added to appropriate wells of a 384-well ...More data for this Ligand-Target Pair
Affinity DataKi: 3.30nMAssay Description:To measure the ability of test compounds in the present invention to bind to the human EP3 receptor, and therefore have the potential to antagonize P...More data for this Ligand-Target Pair
Affinity DataKi: 3.60nM ΔG°: -48.2kJ/molepH: 7.4 T: 2°CAssay Description:Test compounds were half log serially diluted in 100% DMSO (J. T. Baker #922401). 1 uL of each compound was added to appropriate wells of a 384-well ...More data for this Ligand-Target Pair
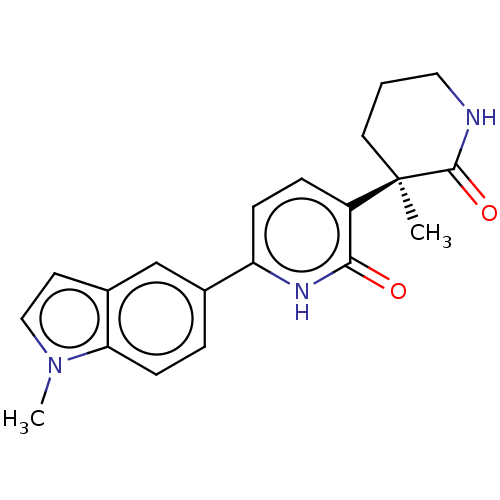
TargetGlutamate receptor ionotropic, NMDA 2B(Rattus norvegicus (Rat))
Astrazeneca Pharmaceuticals
Curated by ChEMBL
Astrazeneca Pharmaceuticals
Curated by ChEMBL
Affinity DataKi: 4.60nMAssay Description:Displacement of [3H]CP101606 from NR2B in rat brain minus cerebellum membraneMore data for this Ligand-Target Pair
Affinity DataKi: 4.60nMAssay Description:To measure the ability of test compounds in the present invention to bind to the human EP3 receptor, and therefore have the potential to antagonize P...More data for this Ligand-Target Pair
Affinity DataKi: 5nMAssay Description:Displacement of [125I]-CXCL12 from human CXCR7 expressed in CHOK1 cell membranes after 2 hrs by scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 5.01nMAssay Description:Displacement of [125I]-CXCL12 from human CXCR7 expressed in CHOK1 cell membranes after 2 hrs by scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 7.20nMAssay Description:To measure the ability of test compounds in the present invention to bind to the human EP3 receptor, and therefore have the potential to antagonize P...More data for this Ligand-Target Pair
Affinity DataKi: 7.30nMAssay Description:To measure the ability of test compounds in the present invention to bind to the human EP3 receptor, and therefore have the potential to antagonize P...More data for this Ligand-Target Pair
Affinity DataKi: 7.80nM ΔG°: -46.3kJ/molepH: 7.4 T: 2°CAssay Description:Test compounds were half log serially diluted in 100% DMSO (J. T. Baker #922401). 1 uL of each compound was added to appropriate wells of a 384-well ...More data for this Ligand-Target Pair
Affinity DataKi: 7.90nMAssay Description:Displacement of [125I]-CXCL12 from human CXCR7 expressed in CHOK1 cell membranes after 2 hrs by scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 7.90nMAssay Description:Displacement of [125I]CXCL12 from human CXCR7 expressed in CHO-K1 cell membranes after 2 hrs by scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 7.94nMAssay Description:Displacement of [125I]CXCL12 from human CXCR7 expressed in CHO-K1 cell membranes after 2 hrs by scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 8nMAssay Description:Displacement of [125I]-CXCL12 from human CXCR7 expressed in CHOK1 cell membranes after 2 hrs by scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 8.90nMAssay Description:To measure the ability of test compounds in the present invention to bind to the human EP3 receptor, and therefore have the potential to antagonize P...More data for this Ligand-Target Pair
Affinity DataKi: 9nMAssay Description:Displacement of [125I]-CXCL12 from human CXCR7 expressed in CHOK1 cell membranes after 2 hrs by scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 9nMAssay Description:Displacement of [125I]-CXCL12 from human CXCR7 expressed in CHOK1 cell membranes after 2 hrs by scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 9nMAssay Description:Displacement of [125I]-CXCL12 from human CXCR7 expressed in CHOK1 cell membranes after 2 hrs by scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 9nMAssay Description:Displacement of [125I]-CXCL12 from human CXCR7 expressed in CHOK1 cell membranes after 2 hrs by scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 9nMAssay Description:Binding affinity to CXCR7 (unknown origin) assessed as inhibition constantMore data for this Ligand-Target Pair
Affinity DataKi: 9.5nM ΔG°: -45.8kJ/molepH: 7.4 T: 2°CAssay Description:Test compounds were half log serially diluted in 100% DMSO (J. T. Baker #922401). 1 uL of each compound was added to appropriate wells of a 384-well ...More data for this Ligand-Target Pair
Affinity DataKi: 11.2nM ΔG°: -45.4kJ/molepH: 7.4 T: 2°CAssay Description:Test compounds were half log serially diluted in 100% DMSO (J. T. Baker #922401). 1 uL of each compound was added to appropriate wells of a 384-well ...More data for this Ligand-Target Pair
Affinity DataKi: 11.9nM ΔG°: -45.2kJ/molepH: 7.4 T: 2°CAssay Description:Test compounds were half log serially diluted in 100% DMSO (J. T. Baker #922401). 1 uL of each compound was added to appropriate wells of a 384-well ...More data for this Ligand-Target Pair
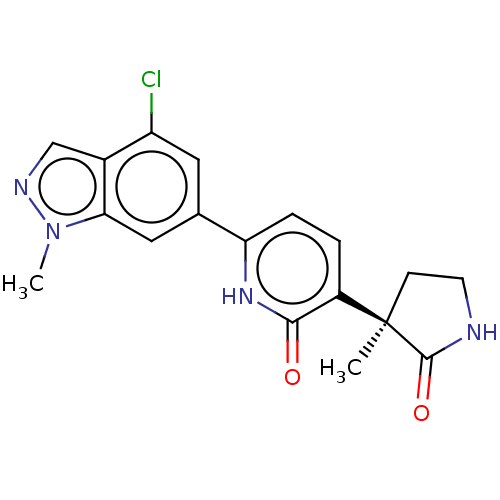
TargetGlutamate receptor ionotropic, NMDA 2B(Rattus norvegicus (Rat))
Astrazeneca Pharmaceuticals
Curated by ChEMBL
Astrazeneca Pharmaceuticals
Curated by ChEMBL
Affinity DataKi: 12nMAssay Description:Displacement of [3H]CP101606 from NR2B in rat brain minus cerebellum membraneMore data for this Ligand-Target Pair
Affinity DataKi: 12.2nM ΔG°: -45.2kJ/molepH: 7.4 T: 2°CAssay Description:Test compounds were half log serially diluted in 100% DMSO (J. T. Baker #922401). 1 uL of each compound was added to appropriate wells of a 384-well ...More data for this Ligand-Target Pair
Affinity DataKi: 12.6nMAssay Description:Displacement of [125I]CXCL12 from human CXCR7 expressed in CHO-K1 cell membranes after 2 hrs by scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 12.6nMAssay Description:Displacement of [125I]CXCL12 from human CXCR7 expressed in CHO-K1 cell membranes after 2 hrs by scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 13nMAssay Description:Displacement of [125I]CXCL12 from human CXCR7 expressed in CHO-K1 cell membranes after 2 hrs by scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 13nMAssay Description:Displacement of [125I]CXCL12 from human CXCR7 expressed in CHO-K1 cell membranes after 2 hrs by scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 13nMAssay Description:Displacement of [125I]CXCL12 from human CXCR7 expressed in CHO-K1 cell membranes after 2 hrs by scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 13nMAssay Description:Displacement of [125I]CXCL12 from human CXCR7 expressed in CHO-K1 cell membranes after 2 hrs by scintillation counting methodMore data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Homo sapiens (Human))
Pfizer
Curated by ChEMBL
Pfizer
Curated by ChEMBL
Affinity DataKi: <13nMAssay Description:Inhibition of human ERG by membrane-competitive-binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 13.9nM ΔG°: -44.8kJ/molepH: 7.4 T: 2°CAssay Description:Test compounds were half log serially diluted in 100% DMSO (J. T. Baker #922401). 1 uL of each compound was added to appropriate wells of a 384-well ...More data for this Ligand-Target Pair
Affinity DataKi: 14.2nM ΔG°: -44.8kJ/molepH: 7.4 T: 2°CAssay Description:Test compounds were half log serially diluted in 100% DMSO (J. T. Baker #922401). 1 uL of each compound was added to appropriate wells of a 384-well ...More data for this Ligand-Target Pair
Affinity DataKi: 17nMAssay Description:Binding affinity to CXCR7 (unknown origin) assessed as inhibition constantMore data for this Ligand-Target Pair
Affinity DataKi: 18nMAssay Description:Displacement of [125I]CXCL12 from mouse CXCR7 expressed in CHO-K1 cell membranes after 2 hrs by scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 18.6nMAssay Description:To measure the ability of test compounds in the present invention to bind to the human EP3 receptor, and therefore have the potential to antagonize P...More data for this Ligand-Target Pair
Affinity DataKi: 19.9nM ΔG°: -44.0kJ/molepH: 7.4 T: 2°CAssay Description:Test compounds were half log serially diluted in 100% DMSO (J. T. Baker #922401). 1 uL of each compound was added to appropriate wells of a 384-well ...More data for this Ligand-Target Pair
Affinity DataKi: 20nMAssay Description:Displacement of [125I]-CXCL12 from human CXCR7 expressed in CHOK1 cell membranes after 2 hrs by scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 20nMAssay Description:Displacement of [125I]-CXCL12 from human CXCR7 expressed in CHOK1 cell membranes after 2 hrs by scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 23nMAssay Description:Displacement of [125I]-CXCL12 from human CXCR7 expressed in CHOK1 cell membranes after 2 hrs by scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 24nMAssay Description:Displacement of [125I]-CXCL12 from human CXCR7 expressed in CHOK1 cell membranes after 2 hrs by scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 25nMAssay Description:Modulation of ProLink-fused human CXCR7 expressed in CHOK1 cell membranes assessed as induction of EA-tagged beta-arrestin recruitment after 30 mins ...More data for this Ligand-Target Pair
Affinity DataKi: 25nMAssay Description:Displacement of [125I]-CXCL12 from human CXCR7 expressed in CHOK1 cell membranes after 2 hrs by scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 25nMAssay Description:Displacement of [125I]-CXCL12 from human CXCR7 expressed in CHOK1 cell membranes after 2 hrs by scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 25nM ΔG°: -43.4kJ/molepH: 7.4 T: 2°CAssay Description:Test compounds were half log serially diluted in 100% DMSO (J. T. Baker #922401). 1 uL of each compound was added to appropriate wells of a 384-well ...More data for this Ligand-Target Pair
Affinity DataKi: 25nMAssay Description:Displacement of [125I]-CXCL12 from human CXCR7 expressed in CHOK1 cell membranes after 2 hrs by scintillation counting methodMore data for this Ligand-Target Pair
TargetGlutamate receptor ionotropic, NMDA 2B(Rattus norvegicus (Rat))
Astrazeneca Pharmaceuticals
Curated by ChEMBL
Astrazeneca Pharmaceuticals
Curated by ChEMBL
Affinity DataKi: 27nMAssay Description:Displacement of [3H]CP101606 from NR2B in rat brain minus cerebellum membraneMore data for this Ligand-Target Pair
Affinity DataKi: 29nMAssay Description:Displacement of [125I]CXCL12 from human CXCR7 expressed in CHO-K1 cell membranes after 2 hrs by scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 29.2nM ΔG°: -43.0kJ/molepH: 7.4 T: 2°CAssay Description:Test compounds were half log serially diluted in 100% DMSO (J. T. Baker #922401). 1 uL of each compound was added to appropriate wells of a 384-well ...More data for this Ligand-Target Pair