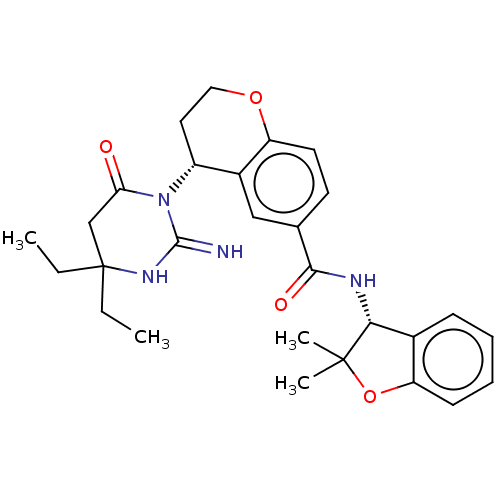
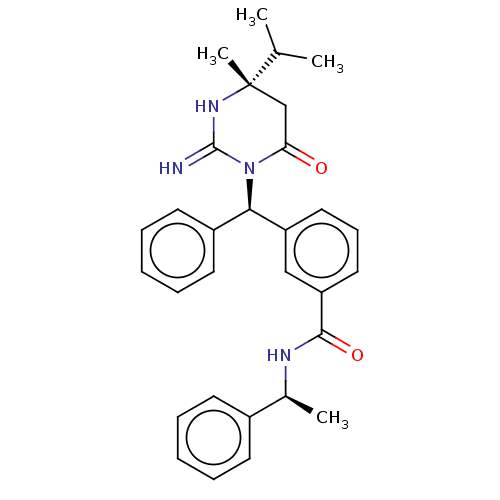
Affinity DataKi: 4.60nMAssay Description:Compound was evaluated for displacement of [3H]QNB from human Muscarinic m1 receptor in CHO cells.More data for this Ligand-Target Pair
Affinity DataKi: 5.30nMAssay Description:Compound was evaluated for displacement of [3H]-QNB from human Muscarinic m2 receptor in CHO cells.More data for this Ligand-Target Pair
TargetPyroglutamylated RF-amide peptide receptor(Homo sapiens (Human))
Schering-Plough Research Institute
Curated by PDSP Ki Database
Schering-Plough Research Institute
Curated by PDSP Ki Database
TargetPyroglutamylated RF-amide peptide receptor(Homo sapiens (Human))
Schering-Plough Research Institute
Curated by PDSP Ki Database
Schering-Plough Research Institute
Curated by PDSP Ki Database
Affinity DataKi: 12nMAssay Description:Compound was evaluated for displacement of [3H]QNB from human Muscarinic m1 receptor in CHO cells.More data for this Ligand-Target Pair
Affinity DataKi: 52nMAssay Description:Compound was evaluated for displacement of [3H]-QNB from human Muscarinic m2 receptor in CHO cells.More data for this Ligand-Target Pair
Affinity DataKi: 71nMAssay Description:Compound was evaluated for displacement of [3H]-QNB from human Muscarinic m2 receptor in CHO cells.More data for this Ligand-Target Pair
Affinity DataKi: 82nMAssay Description:Compound was evaluated for displacement of [3H]QNB from human Muscarinic m1 receptor in CHO cells.More data for this Ligand-Target Pair
Affinity DataKi: 179nMAssay Description:Compound was evaluated for displacement of [3H]QNB from human Muscarinic m1 receptor in CHO cells.More data for this Ligand-Target Pair
TargetPyroglutamylated RF-amide peptide receptor(Homo sapiens (Human))
Schering-Plough Research Institute
Curated by PDSP Ki Database
Schering-Plough Research Institute
Curated by PDSP Ki Database
TargetPyroglutamylated RF-amide peptide receptor(Homo sapiens (Human))
Schering-Plough Research Institute
Curated by PDSP Ki Database
Schering-Plough Research Institute
Curated by PDSP Ki Database
Affinity DataKi: 251nMAssay Description:Compound was evaluated for displacement of [3H]-QNB from human Muscarinic m2 receptor in CHO cells.More data for this Ligand-Target Pair
TargetPyroglutamylated RF-amide peptide receptor(Homo sapiens (Human))
Schering-Plough Research Institute
Curated by PDSP Ki Database
Schering-Plough Research Institute
Curated by PDSP Ki Database
TargetPyroglutamylated RF-amide peptide receptor(Homo sapiens (Human))
Schering-Plough Research Institute
Curated by PDSP Ki Database
Schering-Plough Research Institute
Curated by PDSP Ki Database
TargetPyroglutamylated RF-amide peptide receptor(Homo sapiens (Human))
Schering-Plough Research Institute
Curated by PDSP Ki Database
Schering-Plough Research Institute
Curated by PDSP Ki Database
Affinity DataKi: 1.70E+3nMAssay Description:Displacement of [125I]MCH from human MCHR1 expressed in CHO cell membranesMore data for this Ligand-Target Pair
Affinity DataKi: 7.50E+3nMAssay Description:Displacement of [125I]MCH from human MCHR1 expressed in CHO cell membranesMore data for this Ligand-Target Pair
Affinity DataKi: 7.50E+3nMAssay Description:Displacement of [125I]MCH from human MCHR1 expressed in CHO cell membranesMore data for this Ligand-Target Pair
Affinity DataKi: 7.80E+3nMAssay Description:Displacement of [125I]MCH from human MCHR1 expressed in CHO cell membranesMore data for this Ligand-Target Pair
Affinity DataKi: 8.10E+3nMAssay Description:Displacement of [125I]MCH from human MCHR1 expressed in CHO cell membranesMore data for this Ligand-Target Pair
TargetPyroglutamylated RF-amide peptide receptor(Homo sapiens (Human))
Schering-Plough Research Institute
Curated by PDSP Ki Database
Schering-Plough Research Institute
Curated by PDSP Ki Database
Affinity DataKi: 1.13E+4nMAssay Description:Displacement of [125I]MCH from human MCHR1 expressed in CHO cell membranesMore data for this Ligand-Target Pair
Affinity DataKi: 1.19E+4nMAssay Description:Displacement of [125I]MCH from human MCHR1 expressed in CHO cell membranesMore data for this Ligand-Target Pair
Affinity DataKi: 2.09E+4nMAssay Description:Displacement of [125I]MCH from human MCHR1 expressed in CHO cell membranesMore data for this Ligand-Target Pair
Affinity DataIC50: 0.0300nMAssay Description:Inhibition of Plasmodium falciparum 3D7 HA-glms-tagged PMX using DABCYL-HSFIQEGKEE-EDANS as substrate measured after 2 hrs by FRET assayMore data for this Ligand-Target Pair
Affinity DataIC50: 0.0600nMAssay Description:Inhibition of Plasmodium falciparum 3D7 HA-glms-tagged PMX using DABCYL-HSFIQEGKEE-EDANS as substrate measured after 2 hrs by FRET assayMore data for this Ligand-Target Pair
Affinity DataIC50: 0.0700nMAssay Description:Inhibition of Plasmodium falciparum 3D7 HA-glms-tagged PMX using DABCYL-HSFIQEGKEE-EDANS as substrate measured after 2 hrs by FRET assayMore data for this Ligand-Target Pair
Affinity DataIC50: 0.300nMAssay Description:Inhibition of Plasmodium falciparum 3D7 HA-glms-tagged PMX using DABCYL-HSFIQEGKEE-EDANS as substrate measured after 2 hrs by FRET assayMore data for this Ligand-Target Pair
Affinity DataIC50: 0.400nMAssay Description:Inhibition of Plasmodium falciparum 3D7 HA-glms-tagged PMX using DABCYL-HSFIQEGKEE-EDANS as substrate measured after 2 hrs by FRET assayMore data for this Ligand-Target Pair
Affinity DataIC50: 0.600nMAssay Description:Inhibition of Plasmodium falciparum 3D7 HA-glms-tagged PMX using DABCYL-HSFIQEGKEE-EDANS as substrate measured after 2 hrs by FRET assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.40nMAssay Description:Inhibition of Plasmodium falciparum 3D7 HA-glms-tagged PMIX using DABCYL-HSFIQEGKEE-EDANS as substrate measured after 2 hrs by FRET assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.90nMAssay Description:Inhibition of Plasmodium falciparum 3D7 HA-glms-tagged PMX using DABCYL-HSFIQEGKEE-EDANS as substrate measured after 2 hrs by FRET assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2.20nMAssay Description:Inhibition of Plasmodium falciparum 3D7 HA-glms-tagged PMIX using DABCYL-HSFIQEGKEE-EDANS as substrate measured after 2 hrs by FRET assayMore data for this Ligand-Target Pair
Affinity DataIC50: 4.5nMAssay Description:Inhibition of Plasmodium falciparum 3D7 HA-glms-tagged PMIX using DABCYL-HSFIQEGKEE-EDANS as substrate measured after 2 hrs by FRET assayMore data for this Ligand-Target Pair
Affinity DataIC50: 5.70nMAssay Description:Inhibition of Plasmodium falciparum 3D7 HA-glms-tagged PMX using DABCYL-HSFIQEGKEE-EDANS as substrate measured after 2 hrs by FRET assayMore data for this Ligand-Target Pair
Affinity DataIC50: 12nMAssay Description:Inhibition of Plasmodium falciparum 3D7 HA-glms-tagged PMIX using DABCYL-HSFIQEGKEE-EDANS as substrate measured after 2 hrs by FRET assayMore data for this Ligand-Target Pair
Affinity DataIC50: 20nMT: 2°CAssay Description:Fluorescence-detected RNAP inhibition assays in Tables 1 and 4 were performed using a DNA fragment containing the lacUV5 promoter as a template and t...More data for this Ligand-Target Pair
Affinity DataIC50: 24nMAssay Description:Inhibition of Plasmodium falciparum 3D7 HA-glms-tagged PMIX using DABCYL-HSFIQEGKEE-EDANS as substrate measured after 2 hrs by FRET assayMore data for this Ligand-Target Pair
Affinity DataIC50: 30nMT: 2°CAssay Description:Fluorescence-detected RNAP inhibition assays in Tables 1 and 4 were performed using a DNA fragment containing the lacUV5 promoter as a template and t...More data for this Ligand-Target Pair
Affinity DataIC50: 68nMAssay Description:Inhibition of Plasmodium falciparum 3D7 HA-glms-tagged PMIX using DABCYL-HSFIQEGKEE-EDANS as substrate measured after 2 hrs by FRET assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+3nMT: 2°CAssay Description:Fluorescence-detected RNAP inhibition assays in Tables 1 and 4 were performed using a DNA fragment containing the lacUV5 promoter as a template and t...More data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+3nMT: 2°CAssay Description:Fluorescence-detected RNAP inhibition assays in Tables 1 and 4 were performed using a DNA fragment containing the lacUV5 promoter as a template and t...More data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+3nMT: 2°CAssay Description:Fluorescence-detected RNAP inhibition assays in Tables 1 and 4 were performed using a DNA fragment containing the lacUV5 promoter as a template and t...More data for this Ligand-Target Pair
Affinity DataIC50: 4.00E+3nMT: 2°CAssay Description:Fluorescence-detected RNAP inhibition assays in Tables 1 and 4 were performed using a DNA fragment containing the lacUV5 promoter as a template and t...More data for this Ligand-Target Pair
Affinity DataIC50: 4.00E+3nMT: 2°CAssay Description:Fluorescence-detected RNAP inhibition assays in Tables 1 and 4 were performed using a DNA fragment containing the lacUV5 promoter as a template and t...More data for this Ligand-Target Pair
Affinity DataIC50: >3.00E+4nMAssay Description:Inhibition of human PXRMore data for this Ligand-Target Pair
Ligand InfoPDB
Affinity DataIC50: >3.00E+4nMAssay Description:Reversible inhibition of CYP2C9 (unknown origin)More data for this Ligand-Target Pair
Ligand InfoPDB
Affinity DataIC50: >3.00E+4nMAssay Description:Reversible inhibition of CYP2D6 (unknown origin)More data for this Ligand-Target Pair
Ligand InfoPDB
Affinity DataIC50: >3.00E+4nMAssay Description:Reversible inhibition of CYP3A4 (unknown origin)More data for this Ligand-Target Pair
Ligand InfoPDB
Affinity DataIC50: >3.00E+4nMAssay Description:Inhibition of Nav1.5 (unknown origin)More data for this Ligand-Target Pair
Ligand InfoPDB
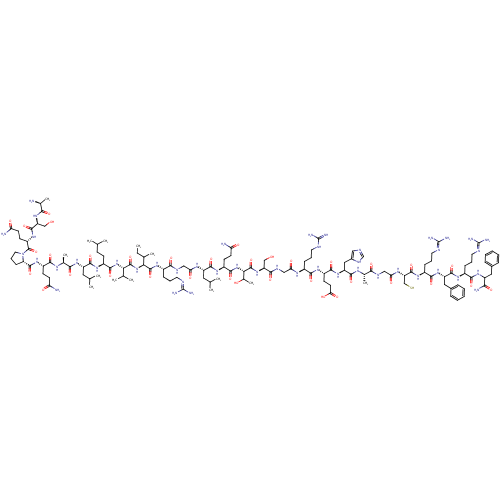

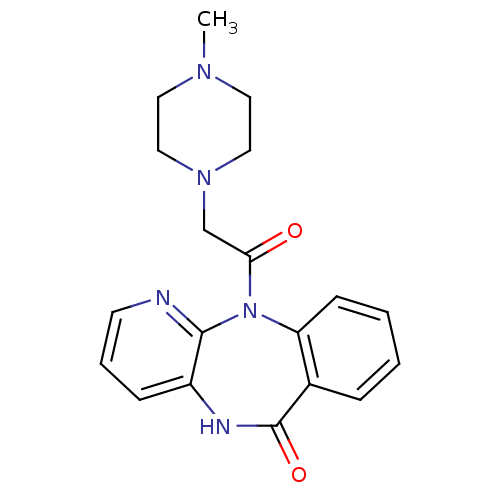
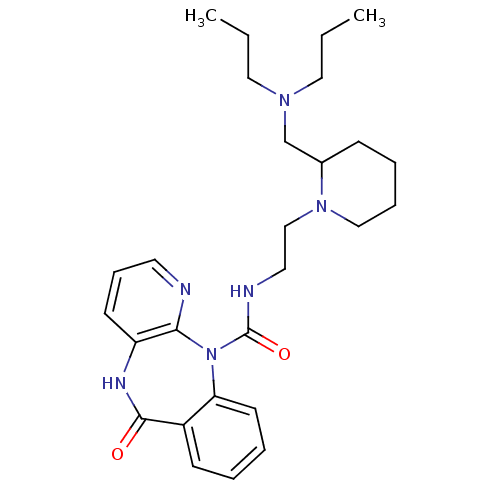










 3D Structure (crystal)
3D Structure (crystal)