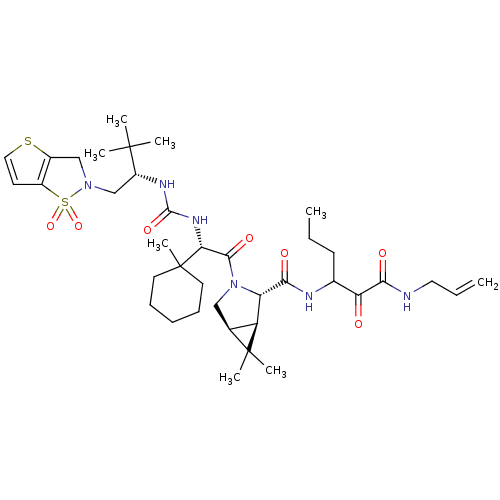
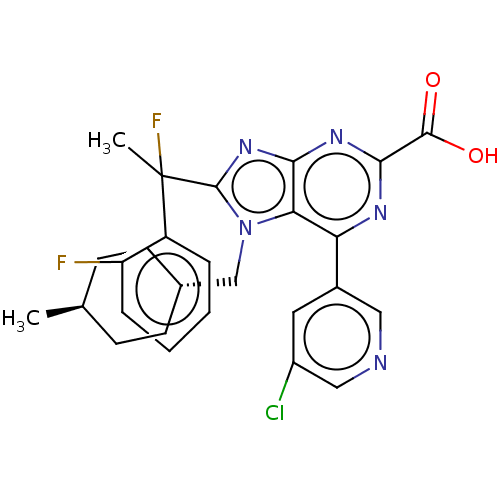
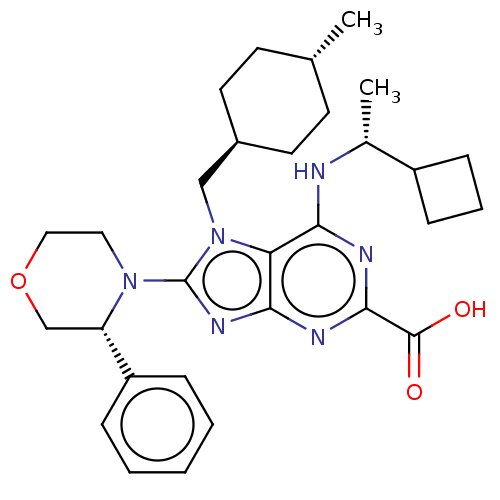
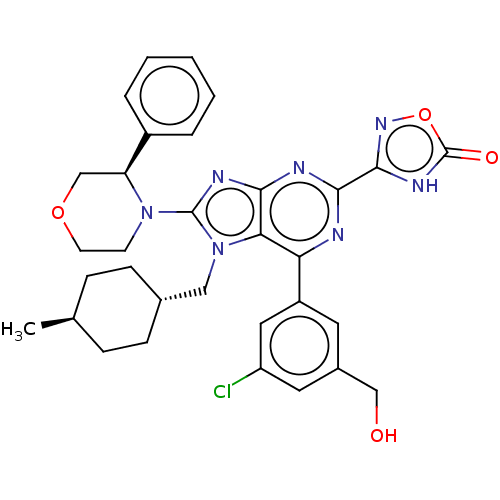
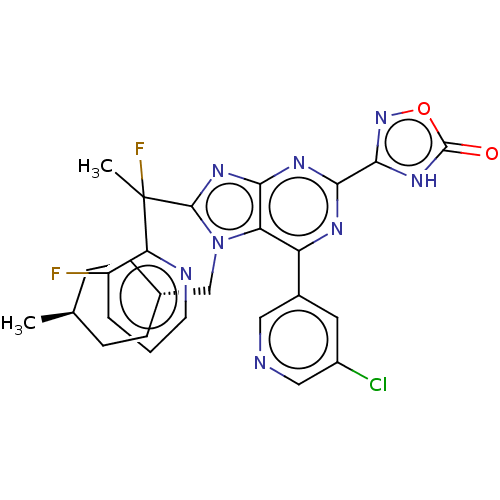
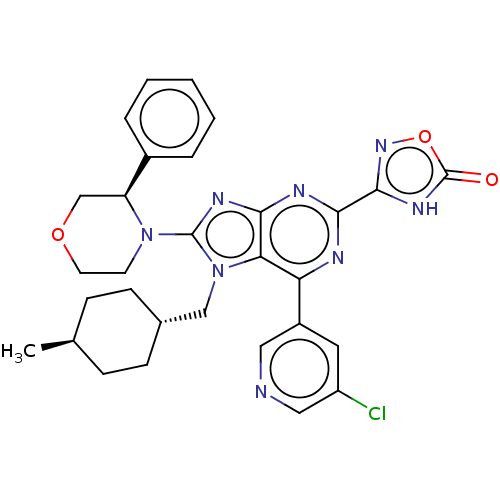
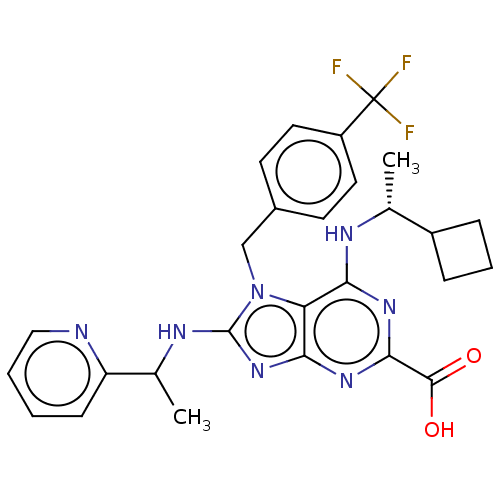
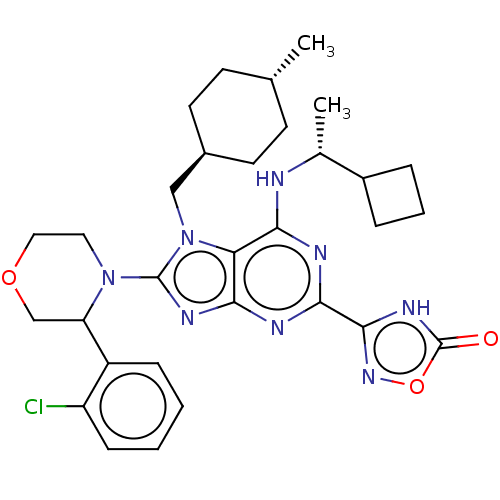
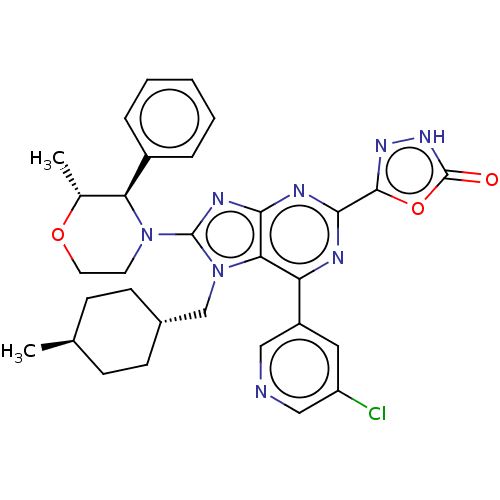
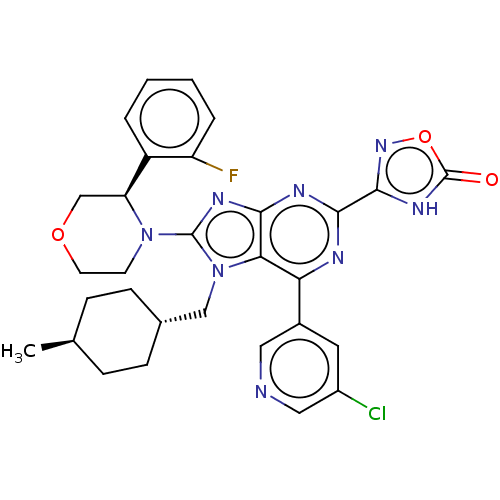
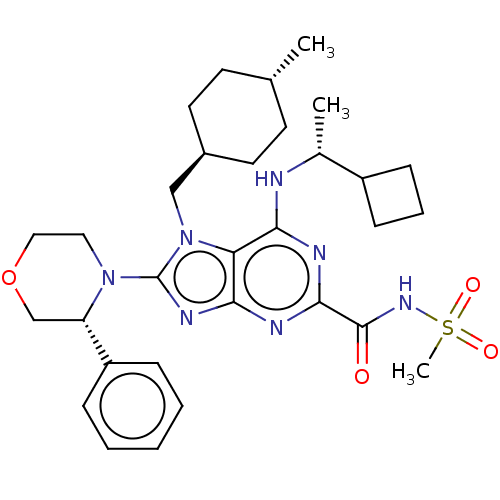
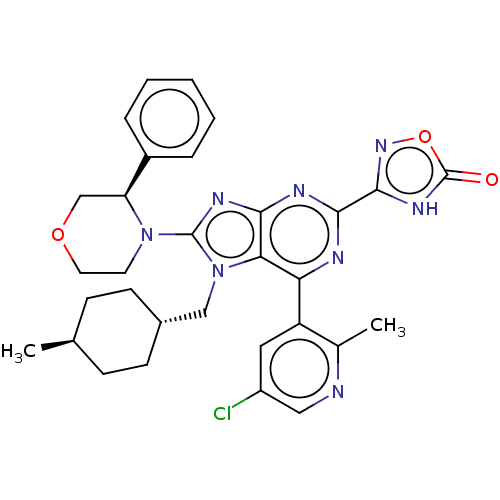
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 0.900nM ΔG°: -52.5kJ/molepH: 6.5 T: 2°CAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
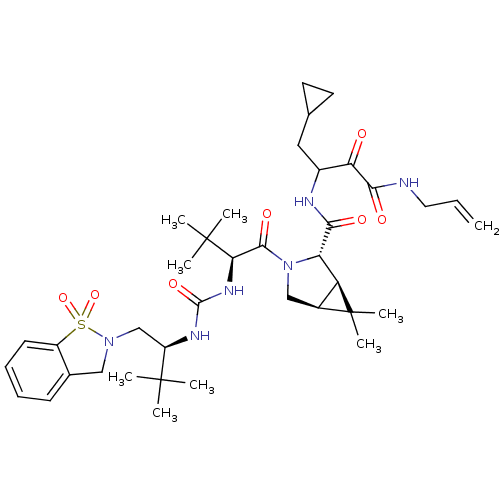
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 3nM ΔG°: -49.5kJ/molepH: 6.5 T: 2°CAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
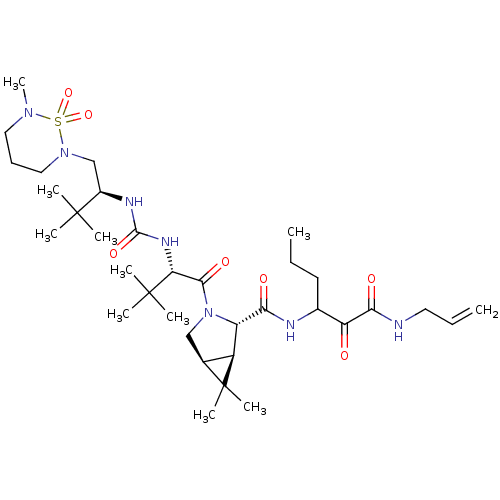
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 3nM ΔG°: -49.5kJ/molepH: 6.5 T: 2°CAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
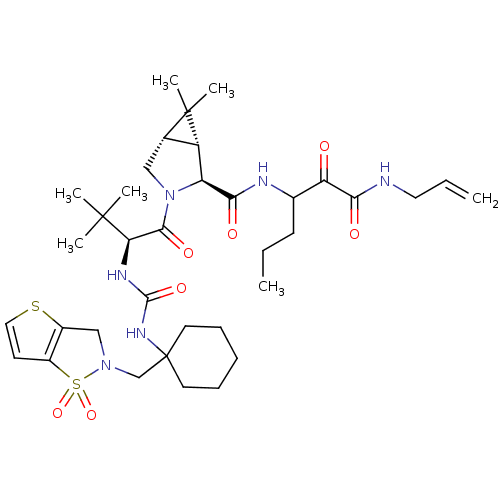
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 5nM ΔG°: -48.2kJ/molepH: 6.5 T: 2°CAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 5nM ΔG°: -48.2kJ/molepH: 6.5 T: 2°CAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 6nM ΔG°: -47.7kJ/molepH: 6.5 T: 2°CAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 7nM ΔG°: -47.3kJ/molepH: 6.5 T: 2°CAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 7nM ΔG°: -47.3kJ/molepH: 6.5 T: 2°CAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 10nM ΔG°: -46.4kJ/molepH: 6.5 T: 2°CAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 11nM ΔG°: -46.2kJ/molepH: 6.5 T: 2°CAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 14nM ΔG°: -45.6kJ/molepH: 6.5 T: 2°CAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 14nM ΔG°: -45.6kJ/molepH: 6.5 T: 2°CAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 15nM ΔG°: -45.4kJ/molepH: 6.5 T: 2°CAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 18nM ΔG°: -44.9kJ/molepH: 6.5 T: 2°CAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 30nM ΔG°: -43.7kJ/molepH: 6.5 T: 2°CAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 30nM ΔG°: -43.7kJ/molepH: 6.5 T: 2°CAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 41nM ΔG°: -42.9kJ/molepH: 6.5 T: 2°CAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 59nM ΔG°: -42.0kJ/molepH: 6.5 T: 2°CAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
Affinity DataIC50: 0.156nMT: 2°CAssay Description:An HDM2 FRET assay was developed to assess the compounds' inhibitory activity towards binding of p53 protein. A truncated version of HDM2 with re...More data for this Ligand-Target Pair
Affinity DataIC50: 0.176nMT: 2°CAssay Description:An HDM2 FRET assay was developed to assess the compounds' inhibitory activity towards binding of p53 protein. A truncated version of HDM2 with re...More data for this Ligand-Target Pair
Affinity DataIC50: 0.202nMT: 2°CAssay Description:An HDM2 FRET assay was developed to assess the compounds' inhibitory activity towards binding of p53 protein. A truncated version of HDM2 with re...More data for this Ligand-Target Pair
Affinity DataIC50: 0.205nMT: 2°CAssay Description:An HDM2 FRET assay was developed to assess the compounds' inhibitory activity towards binding of p53 protein. A truncated version of HDM2 with re...More data for this Ligand-Target Pair
Affinity DataIC50: 0.215nMT: 2°CAssay Description:An HDM2 FRET assay was developed to assess the compounds' inhibitory activity towards binding of p53 protein. A truncated version of HDM2 with re...More data for this Ligand-Target Pair
Affinity DataIC50: 0.289nMT: 2°CAssay Description:An HDM2 FRET assay was developed to assess the compounds' inhibitory activity towards binding of p53 protein. A truncated version of HDM2 with re...More data for this Ligand-Target Pair
Affinity DataIC50: 0.315nMT: 2°CAssay Description:An HDM2 FRET assay was developed to assess the compounds' inhibitory activity towards binding of p53 protein. A truncated version of HDM2 with re...More data for this Ligand-Target Pair
Affinity DataIC50: 0.344nMT: 2°CAssay Description:An HDM2 FRET assay was developed to assess the compounds' inhibitory activity towards binding of p53 protein. A truncated version of HDM2 with re...More data for this Ligand-Target Pair
Affinity DataIC50: 0.359nMT: 2°CAssay Description:An HDM2 FRET assay was developed to assess the compounds' inhibitory activity towards binding of p53 protein. A truncated version of HDM2 with re...More data for this Ligand-Target Pair
Affinity DataIC50: 0.381nMT: 2°CAssay Description:An HDM2 FRET assay was developed to assess the compounds' inhibitory activity towards binding of p53 protein. A truncated version of HDM2 with re...More data for this Ligand-Target Pair
Affinity DataIC50: 0.424nMT: 2°CAssay Description:An HDM2 FRET assay was developed to assess the compounds' inhibitory activity towards binding of p53 protein. A truncated version of HDM2 with re...More data for this Ligand-Target Pair
Affinity DataIC50: 0.435nMT: 2°CAssay Description:An HDM2 FRET assay was developed to assess the compounds' inhibitory activity towards binding of p53 protein. A truncated version of HDM2 with re...More data for this Ligand-Target Pair
Affinity DataIC50: 0.452nMT: 2°CAssay Description:An HDM2 FRET assay was developed to assess the compounds' inhibitory activity towards binding of p53 protein. A truncated version of HDM2 with re...More data for this Ligand-Target Pair
Affinity DataIC50: 0.453nMT: 2°CAssay Description:An HDM2 FRET assay was developed to assess the compounds' inhibitory activity towards binding of p53 protein. A truncated version of HDM2 with re...More data for this Ligand-Target Pair
Affinity DataIC50: 0.494nMT: 2°CAssay Description:An HDM2 FRET assay was developed to assess the compounds' inhibitory activity towards binding of p53 protein. A truncated version of HDM2 with re...More data for this Ligand-Target Pair
Affinity DataIC50: 0.504nMT: 2°CAssay Description:An HDM2 FRET assay was developed to assess the compounds' inhibitory activity towards binding of p53 protein. A truncated version of HDM2 with re...More data for this Ligand-Target Pair
Affinity DataIC50: 0.507nMT: 2°CAssay Description:An HDM2 FRET assay was developed to assess the compounds' inhibitory activity towards binding of p53 protein. A truncated version of HDM2 with re...More data for this Ligand-Target Pair
Affinity DataIC50: 0.515nMT: 2°CAssay Description:An HDM2 FRET assay was developed to assess the compounds' inhibitory activity towards binding of p53 protein. A truncated version of HDM2 with re...More data for this Ligand-Target Pair
Affinity DataIC50: 0.517nMT: 2°CAssay Description:An HDM2 FRET assay was developed to assess the compounds' inhibitory activity towards binding of p53 protein. A truncated version of HDM2 with re...More data for this Ligand-Target Pair
Affinity DataIC50: 0.526nMT: 2°CAssay Description:An HDM2 FRET assay was developed to assess the compounds' inhibitory activity towards binding of p53 protein. A truncated version of HDM2 with re...More data for this Ligand-Target Pair
Affinity DataIC50: 0.548nMT: 2°CAssay Description:An HDM2 FRET assay was developed to assess the compounds' inhibitory activity towards binding of p53 protein. A truncated version of HDM2 with re...More data for this Ligand-Target Pair
Affinity DataIC50: 0.563nMT: 2°CAssay Description:An HDM2 FRET assay was developed to assess the compounds' inhibitory activity towards binding of p53 protein. A truncated version of HDM2 with re...More data for this Ligand-Target Pair
Affinity DataIC50: 0.568nMT: 2°CAssay Description:An HDM2 FRET assay was developed to assess the compounds' inhibitory activity towards binding of p53 protein. A truncated version of HDM2 with re...More data for this Ligand-Target Pair
Affinity DataIC50: 0.569nMT: 2°CAssay Description:An HDM2 FRET assay was developed to assess the compounds' inhibitory activity towards binding of p53 protein. A truncated version of HDM2 with re...More data for this Ligand-Target Pair
Affinity DataIC50: 0.585nMT: 2°CAssay Description:An HDM2 FRET assay was developed to assess the compounds' inhibitory activity towards binding of p53 protein. A truncated version of HDM2 with re...More data for this Ligand-Target Pair
Affinity DataIC50: 0.585nMT: 2°CAssay Description:An HDM2 FRET assay was developed to assess the compounds' inhibitory activity towards binding of p53 protein. A truncated version of HDM2 with re...More data for this Ligand-Target Pair
Affinity DataIC50: 0.585nMT: 2°CAssay Description:An HDM2 FRET assay was developed to assess the compounds' inhibitory activity towards binding of p53 protein. A truncated version of HDM2 with re...More data for this Ligand-Target Pair
Affinity DataIC50: 0.598nMT: 2°CAssay Description:An HDM2 FRET assay was developed to assess the compounds' inhibitory activity towards binding of p53 protein. A truncated version of HDM2 with re...More data for this Ligand-Target Pair
Affinity DataIC50: 0.601nMT: 2°CAssay Description:An HDM2 FRET assay was developed to assess the compounds' inhibitory activity towards binding of p53 protein. A truncated version of HDM2 with re...More data for this Ligand-Target Pair
Affinity DataIC50: 0.620nMT: 2°CAssay Description:An HDM2 FRET assay was developed to assess the compounds' inhibitory activity towards binding of p53 protein. A truncated version of HDM2 with re...More data for this Ligand-Target Pair
Affinity DataIC50: 0.623nMT: 2°CAssay Description:An HDM2 FRET assay was developed to assess the compounds' inhibitory activity towards binding of p53 protein. A truncated version of HDM2 with re...More data for this Ligand-Target Pair
Affinity DataIC50: 0.626nMT: 2°CAssay Description:An HDM2 FRET assay was developed to assess the compounds' inhibitory activity towards binding of p53 protein. A truncated version of HDM2 with re...More data for this Ligand-Target Pair