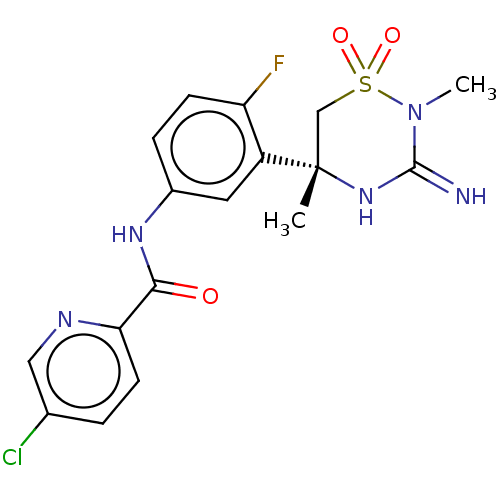
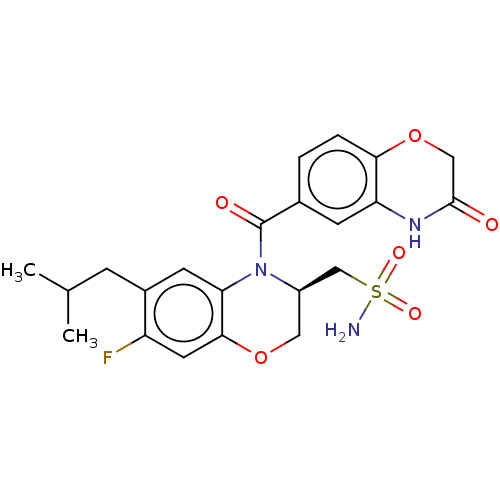
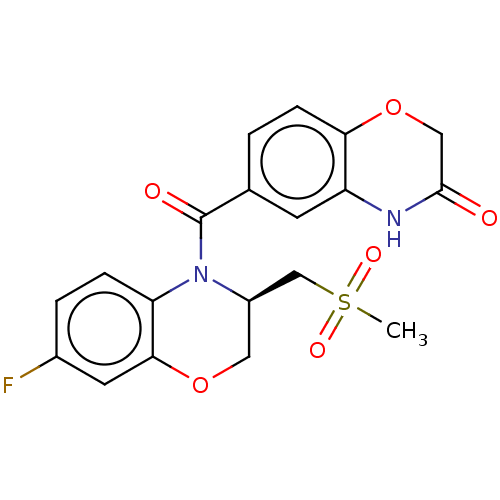
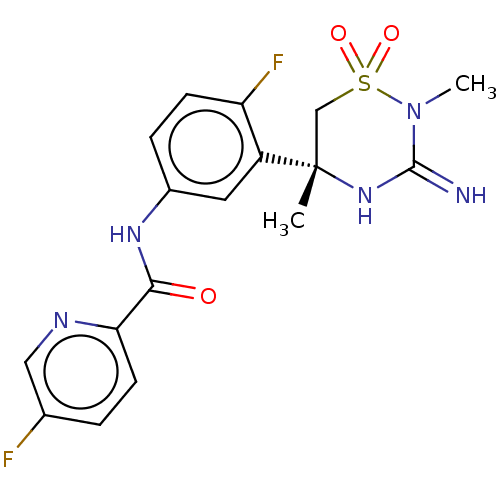
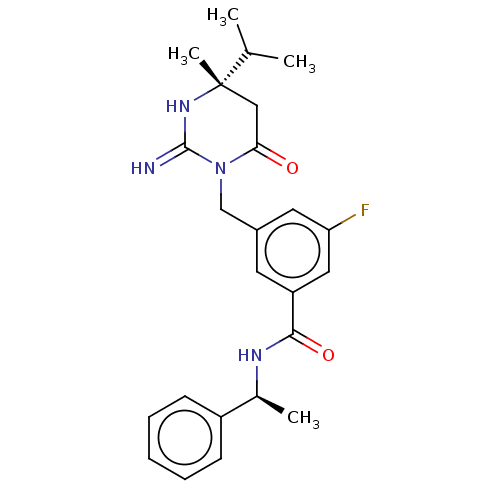
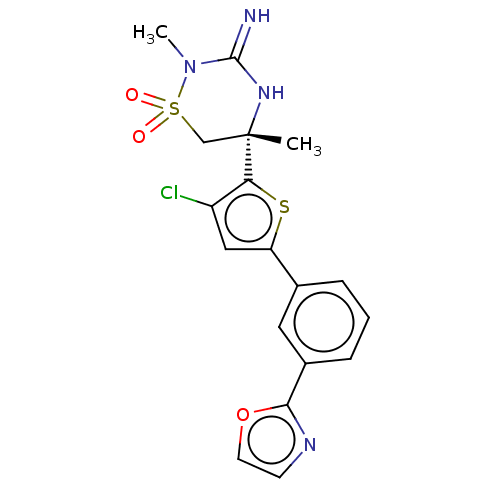
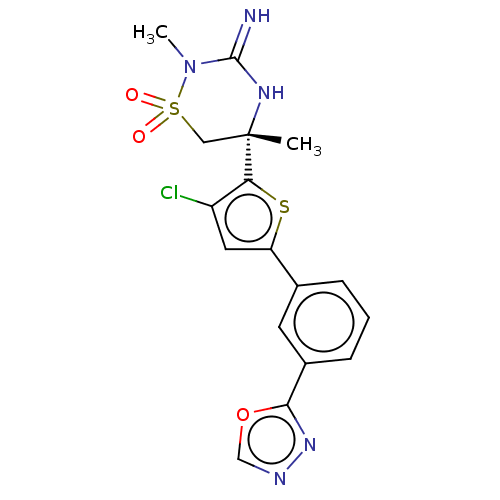
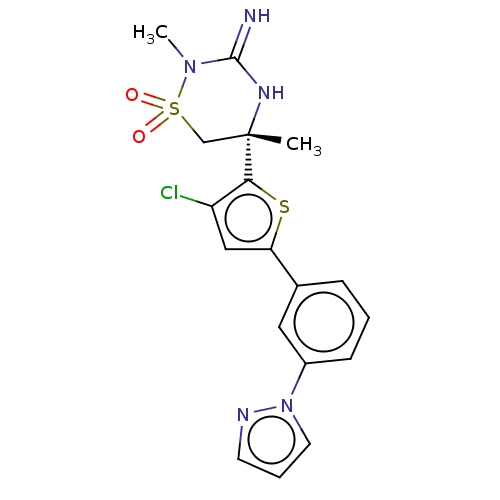
TargetTransient receptor potential cation channel subfamily V member 1(Rattus norvegicus (rat))
Seoul National University
Curated by ChEMBL
Seoul National University
Curated by ChEMBL
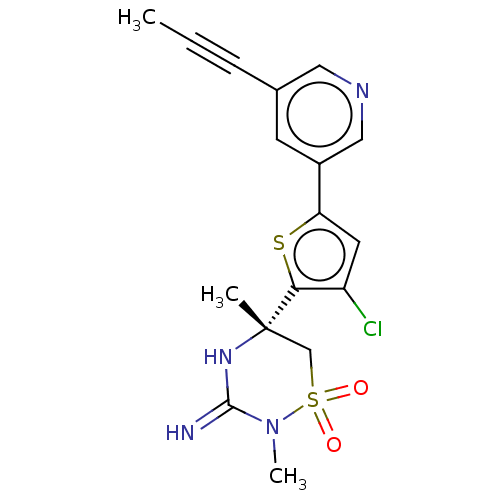
Affinity DataKi: 0.0230nMAssay Description:In vitro binding affinity towards vanilloid receptor by [3H]RTX displacement.More data for this Ligand-Target Pair
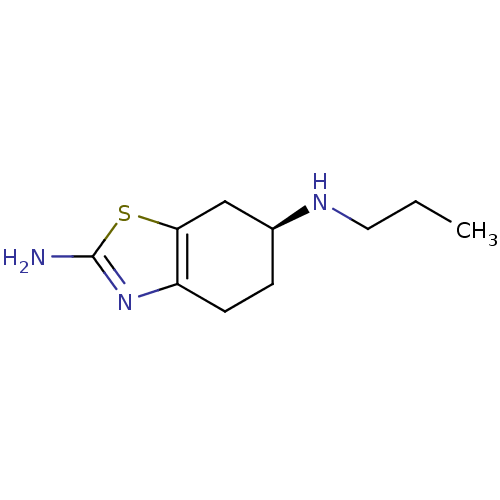
TargetTransient receptor potential cation channel subfamily V member 1(Rattus norvegicus (rat))
Seoul National University
Curated by ChEMBL
Seoul National University
Curated by ChEMBL
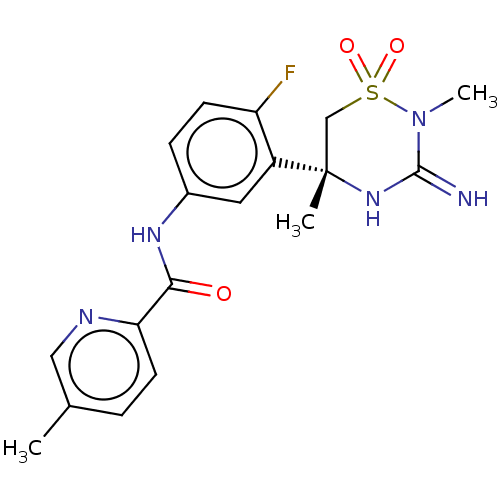
Affinity DataKi: 0.130nMAssay Description:In vitro binding to Rat Vanilloid receptor 1 (VR1) expressing CHO cells compared to capsacinMore data for this Ligand-Target Pair
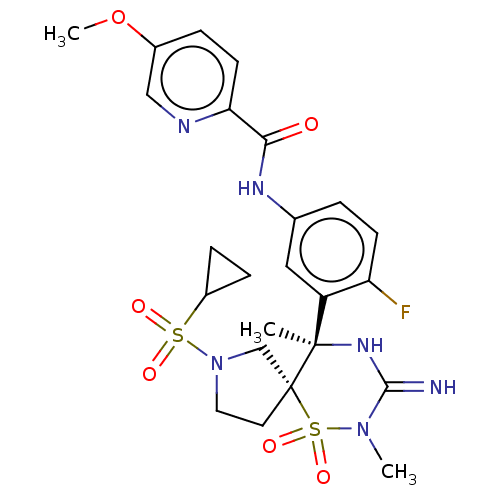
Affinity DataKi: 0.220nM ΔG°: -56.0kJ/molepH: 5.0 T: 2°CAssay Description:Inhibitor IC50s at purified human autoBACE-2 were determined in a time-resolved endpoint proteolysis assay that measures hydrolysis of the QSY7-EISEV...More data for this Ligand-Target Pair
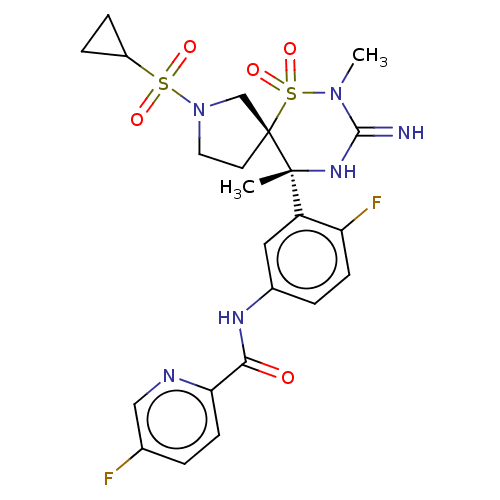
Affinity DataKi: 0.240nM ΔG°: -55.8kJ/molepH: 5.0 T: 2°CAssay Description:Inhibitor compounds, prepared at 3× the desired final concentration in 1×BACE assay buffer (20 mM sodium acetate pH 5.0, 10% glycerol, 0....More data for this Ligand-Target Pair
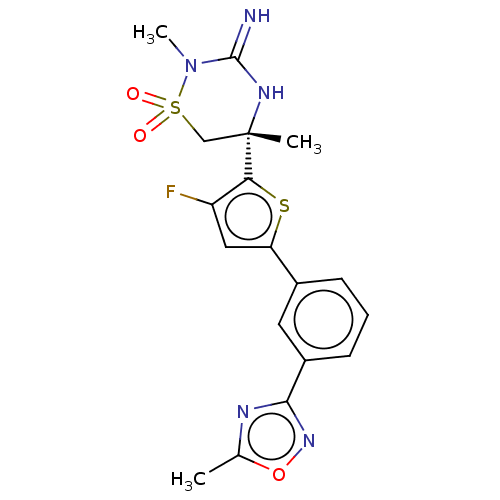
Affinity DataKi: 0.280nM ΔG°: -55.4kJ/molepH: 5.0 T: 2°CAssay Description:Inhibitor compounds, prepared at 3× the desired final concentration in 1×BACE assay buffer (20 mM sodium acetate pH 5.0, 10% glycerol, 0....More data for this Ligand-Target Pair
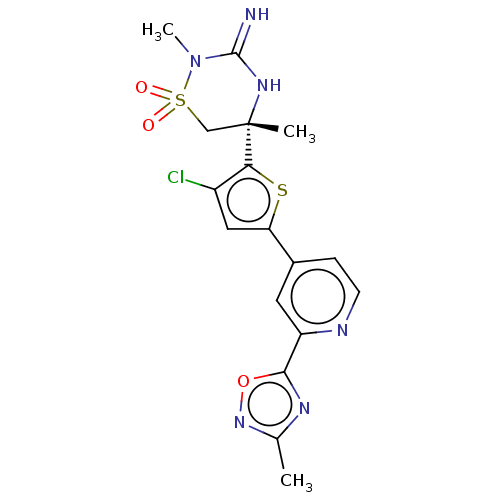
Affinity DataKi: 0.310nM ΔG°: -55.2kJ/molepH: 5.0 T: 2°CAssay Description:Inhibitor compounds, prepared at 3× the desired final concentration in 1×BACE assay buffer (20 mM sodium acetate pH 5.0, 10% glycerol, 0....More data for this Ligand-Target Pair
Affinity DataKi: >0.316nMAssay Description:Displacement of [3H]-aldosterone to human mineralocorticoid receptor LBD by radiometric binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 0.316nMAssay Description:Displacement of [3H]-aldosterone to human mineralocorticoid receptor LBD by radiometric binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 0.340nM ΔG°: -54.9kJ/molepH: 5.0 T: 2°CAssay Description:Inhibitor compounds, prepared at 3× the desired final concentration in 1×BACE assay buffer (20 mM sodium acetate pH 5.0, 10% glycerol, 0....More data for this Ligand-Target Pair
Affinity DataKi: 0.350nM ΔG°: -54.9kJ/molepH: 5.0 T: 2°CAssay Description:Inhibitor compounds, prepared at 3× the desired final concentration in 1×BACE assay buffer (20 mM sodium acetate pH 5.0, 10% glycerol, 0....More data for this Ligand-Target Pair
Affinity DataKi: 0.350nM ΔG°: -54.9kJ/molepH: 5.0 T: 2°CAssay Description:Inhibitor compounds, prepared at 3× the desired final concentration in 1×BACE assay buffer (20 mM sodium acetate pH 5.0, 10% glycerol, 0....More data for this Ligand-Target Pair
Affinity DataKi: 0.370nM ΔG°: -54.7kJ/molepH: 5.0 T: 2°CAssay Description:Inhibitor IC50s at purified human autoBACE-2 were determined in a time-resolved endpoint proteolysis assay that measures hydrolysis of the QSY7-EISEV...More data for this Ligand-Target Pair
Affinity DataKi: 0.370nM ΔG°: -54.7kJ/molepH: 5.0 T: 2°CAssay Description:Inhibitor IC50s at purified human autoBACE-2 were determined in a time-resolved endpoint proteolysis assay that measures hydrolysis of the QSY7-EISEV...More data for this Ligand-Target Pair
Affinity DataKi: 0.410nM ΔG°: -54.5kJ/molepH: 5.0 T: 2°CAssay Description:Inhibitor compounds, prepared at 3× the desired final concentration in 1×BACE assay buffer (20 mM sodium acetate pH 5.0, 10% glycerol, 0....More data for this Ligand-Target Pair
Affinity DataKi: 0.450nM ΔG°: -54.2kJ/molepH: 5.0 T: 2°CAssay Description:Inhibitor IC50s at purified human autoBACE-2 were determined in a time-resolved endpoint proteolysis assay that measures hydrolysis of the QSY7-EISEV...More data for this Ligand-Target Pair
Affinity DataKi: 0.470nM ΔG°: -54.1kJ/molepH: 5.0 T: 2°CAssay Description:Inhibitor IC50s at purified human autoBACE-2 were determined in a time-resolved endpoint proteolysis assay that measures hydrolysis of the QSY7-EISEV...More data for this Ligand-Target Pair
Affinity DataKi: 0.5nM ΔG°: -54.0kJ/molepH: 5.0 T: 2°CAssay Description:Varying concentrations of inhibitors at 3× the final desired concentration in a volume of 10 μl are preincubated with purified human BACE1 c...More data for this Ligand-Target Pair
Affinity DataKi: 0.501nMAssay Description:Displacement of [3H]-aldosterone to human mineralocorticoid receptor LBD by radiometric binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 0.600nMAssay Description:Inhibition of renin (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 0.630nM ΔG°: -53.4kJ/molepH: 5.0 T: 2°CAssay Description:Inhibitor compounds, prepared at 3× the desired final concentration in 1×BACE assay buffer (20 mM sodium acetate pH 5.0, 10% glycerol, 0....More data for this Ligand-Target Pair
Affinity DataKi: 0.631nMAssay Description:Displacement of [3H]-aldosterone to human mineralocorticoid receptor LBD by radiometric binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 0.690nM ΔG°: -53.2kJ/molepH: 5.0 T: 2°CAssay Description:Inhibitor compounds, prepared at 3× the desired final concentration in 1×BACE assay buffer (20 mM sodium acetate pH 5.0, 10% glycerol, 0....More data for this Ligand-Target Pair
Affinity DataKi: 0.730nM ΔG°: -53.0kJ/molepH: 5.0 T: 2°CAssay Description:Inhibitor compounds, prepared at 3× the desired final concentration in 1×BACE assay buffer (20 mM sodium acetate pH 5.0, 10% glycerol, 0....More data for this Ligand-Target Pair
Affinity DataKi: 0.794nMAssay Description:Displacement of [3H]-aldosterone to human mineralocorticoid receptor LBD by radiometric binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 0.870nM ΔG°: -52.6kJ/molepH: 5.0 T: 2°CAssay Description:Varying concentrations of inhibitors at 3× the final desired concentration in a volume of 10 μl are preincubated with purified human BACE1 c...More data for this Ligand-Target Pair
Affinity DataKi: 0.880nMAssay Description:High inhibition constant against [3H]-spiperone binding to human Dopamine receptor D3 expressed in CHO cellsMore data for this Ligand-Target Pair
Affinity DataKi: 0.949nMpH: 5.01Assay Description:The protocol that was used to determine the recited values isdescribed as follows.BACE1 HTRF FRET AssayReagentsNa+-Acetate pH 5.01% Brij-35GlycerolDi...More data for this Ligand-Target Pair
Affinity DataKi: 1nM ΔG°: -52.2kJ/molepH: 5.0 T: 2°CAssay Description:A homogeneous time-resolved FRET assay can be used to determine IC50 values for inhibitors of the soluble human BACE1 catalytic domain. This assay mo...More data for this Ligand-Target Pair
Affinity DataKi: 1nM ΔG°: -52.2kJ/molepH: 5.0 T: 2°CAssay Description:Inhibitor compounds, prepared at 3× the desired final concentration in 1×BACE assay buffer (20 mM sodium acetate pH 5.0, 10% glycerol, 0....More data for this Ligand-Target Pair
Affinity DataKi: 1nM ΔG°: -52.2kJ/molepH: 5.0 T: 2°CAssay Description:A homogeneous time-resolved FRET assay can be used to determine IC50 values for inhibitors of the soluble human BACE1 catalytic domain. This assay mo...More data for this Ligand-Target Pair
Affinity DataKi: 1nM ΔG°: -52.2kJ/molepH: 5.0 T: 2°CAssay Description:Inhibitor IC50's at purified human autoBACE-2 are determined in a time-resolved endpoint proteolysis assay that measures hydrolysis of the QSY7-EISEV...More data for this Ligand-Target Pair
Affinity DataKi: 1nM ΔG°: -52.2kJ/molepH: 5.0 T: 2°CAssay Description:Inhibitor IC50's at purified human autoBACE-2 are determined in a time-resolved endpoint proteolysis assay that measures hydrolysis of the QSY7-EISEV...More data for this Ligand-Target Pair
Affinity DataKi: 1nM ΔG°: -52.2kJ/molepH: 5.0 T: 2°CAssay Description:Inhibitor IC50's at purified human autoBACE-2 are determined in a time-resolved endpoint proteolysis assay that measures hydrolysis of the QSY7-EISEV...More data for this Ligand-Target Pair
Affinity DataKi: 1nM ΔG°: -52.2kJ/molepH: 5.0 T: 2°CAssay Description:A homogeneous time-resolved FRET assay can be used to determine IC50 values for inhibitors of the soluble human BACE1 catalytic domain. This assay mo...More data for this Ligand-Target Pair
Affinity DataKi: 1nM ΔG°: -52.2kJ/molepH: 5.0 T: 2°CAssay Description:A homogeneous time-resolved FRET assay can be used to determine IC50 values for inhibitors of the soluble human BACE1 catalytic domain. This assay mo...More data for this Ligand-Target Pair
Affinity DataKi: 1nM ΔG°: -52.2kJ/molepH: 5.0 T: 2°CAssay Description:A homogeneous time-resolved FRET assay can be used to determine IC50 values for inhibitors of the soluble human BACE1 catalytic domain. This assay mo...More data for this Ligand-Target Pair
Affinity DataKi: 1nM ΔG°: -52.2kJ/molepH: 5.0 T: 2°CAssay Description:A homogeneous time-resolved FRET assay can be used to determine IC50 values for inhibitors of the soluble human BACE1 catalytic domain. This assay mo...More data for this Ligand-Target Pair
Affinity DataKi: 1nM ΔG°: -52.2kJ/molepH: 5.0 T: 2°CAssay Description:A homogeneous time-resolved FRET assay can be used to determine IC50 values for inhibitors of the soluble human BACE1 catalytic domain. This assay mo...More data for this Ligand-Target Pair
Affinity DataKi: 1nM ΔG°: -52.2kJ/molepH: 5.0 T: 2°CAssay Description:A homogeneous time-resolved FRET assay can be used to determine IC50 values for inhibitors of the soluble human BACE1 catalytic domain. This assay mo...More data for this Ligand-Target Pair
Affinity DataKi: 1.05nMpH: 5.01Assay Description:The protocol that was used to determine the recited values isdescribed as follows.BACE1 HTRF FRET AssayReagentsNa+-Acetate pH 5.01% Brij-35GlycerolDi...More data for this Ligand-Target Pair
Affinity DataKi: 1.10nM ΔG°: -52.0kJ/molepH: 5.0 T: 2°CAssay Description:Varying concentrations of inhibitors at 3× the final desired concentration in a volume of 10 μl are preincubated with purified human BACE1 c...More data for this Ligand-Target Pair
Affinity DataKi: 1.10nM ΔG°: -52.0kJ/molepH: 5.0 T: 2°CAssay Description:Inhibitor compounds, prepared at 3× the desired final concentration in 1×BACE assay buffer (20 mM sodium acetate pH 5.0, 10% glycerol, 0....More data for this Ligand-Target Pair
Affinity DataKi: 1.10nM ΔG°: -52.0kJ/molepH: 5.0 T: 2°CAssay Description:Inhibitor compounds, prepared at 3× the desired final concentration in 1×BACE assay buffer (20 mM sodium acetate pH 5.0, 10% glycerol, 0....More data for this Ligand-Target Pair
Affinity DataKi: 1.20nM ΔG°: -51.8kJ/molepH: 5.0 T: 2°CAssay Description:Varying concentrations of inhibitors at 3× the final desired concentration in a volume of 10 μl are preincubated with purified human BACE1 c...More data for this Ligand-Target Pair
Affinity DataKi: 1.26nMpH: 5.01Assay Description:The protocol that was used to determine the recited values isdescribed as follows.BACE1 HTRF FRET AssayReagentsNa+-Acetate pH 5.01% Brij-35GlycerolDi...More data for this Ligand-Target Pair
Affinity DataKi: 1.30nM ΔG°: -51.6kJ/molepH: 5.0 T: 2°CAssay Description:Inhibitor compounds, prepared at 3× the desired final concentration in 1×BACE assay buffer (20 mM sodium acetate pH 5.0, 10% glycerol, 0....More data for this Ligand-Target Pair
Affinity DataKi: 1.30nMAssay Description:Binding affinity to recombinant human GR LBD by fluormone GS red-fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 1.40nM ΔG°: -51.4kJ/molepH: 5.0 T: 2°CAssay Description:Inhibitor compounds, prepared at 3× the desired final concentration in 1×BACE assay buffer (20 mM sodium acetate pH 5.0, 10% glycerol, 0....More data for this Ligand-Target Pair
Affinity DataKi: 1.60nM ΔG°: -51.0kJ/molepH: 5.0 T: 2°CAssay Description:Varying concentrations of inhibitors at 3× the final desired concentration in a volume of 10 μl are preincubated with purified human BACE1 c...More data for this Ligand-Target Pair
Affinity DataKi: 1.60nM ΔG°: -51.0kJ/molepH: 5.0 T: 2°CAssay Description:Inhibitor compounds, prepared at 3× the desired final concentration in 1×BACE assay buffer (20 mM sodium acetate pH 5.0, 10% glycerol, 0....More data for this Ligand-Target Pair

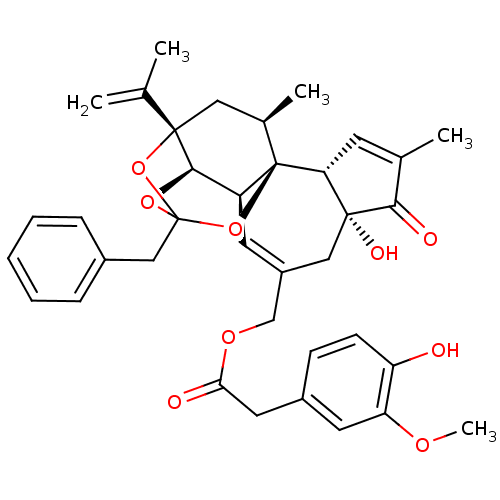
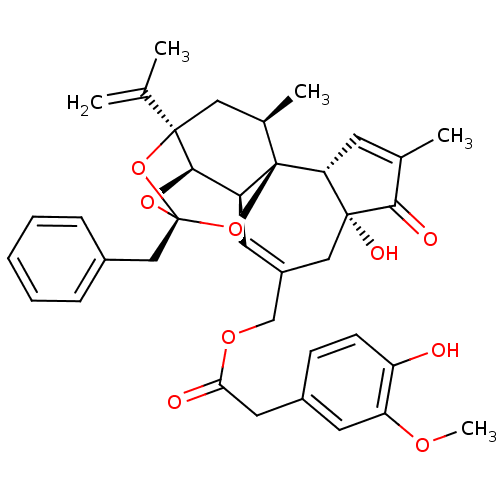
 3D Structure (crystal)
3D Structure (crystal)