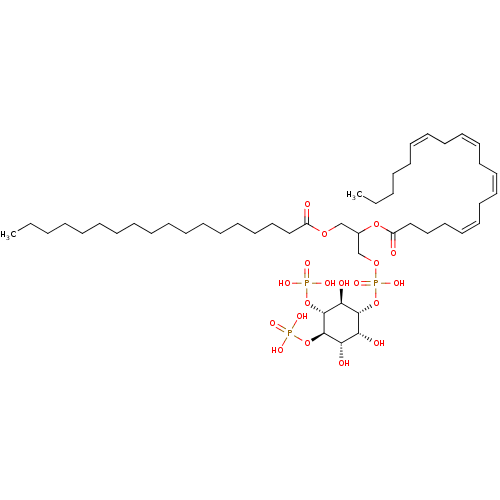
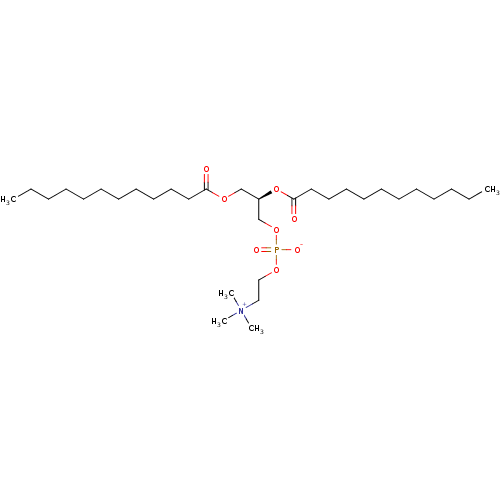
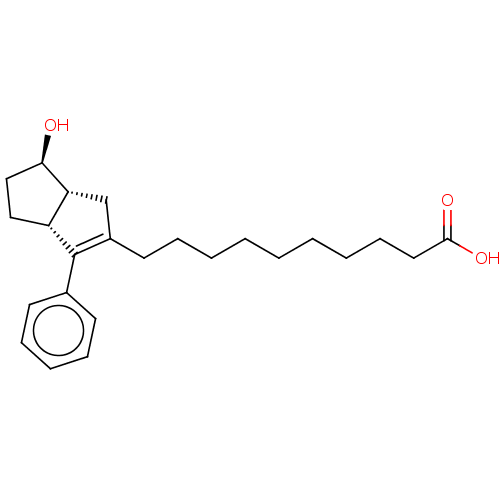
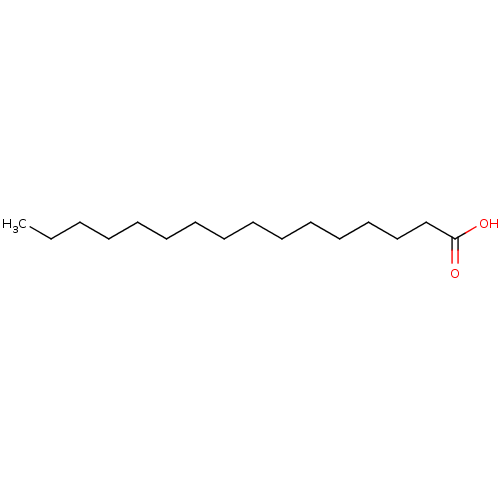
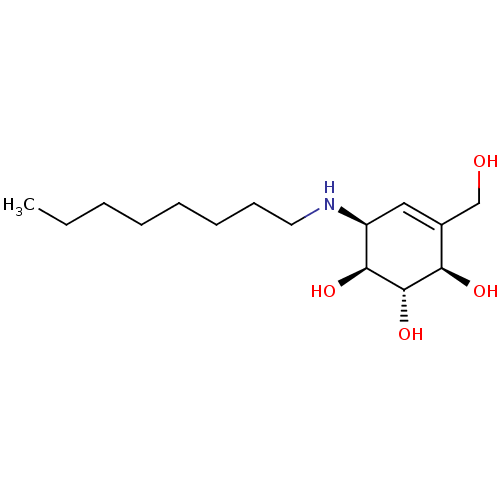
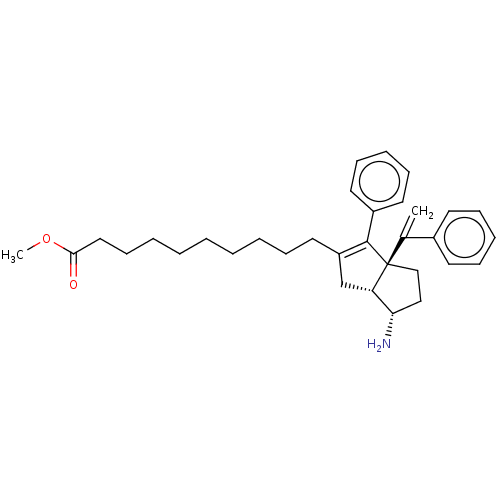
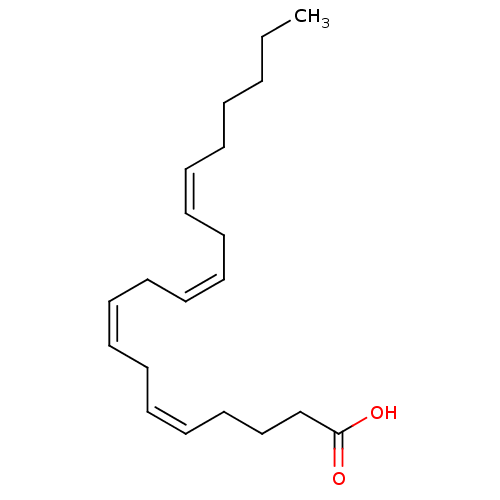
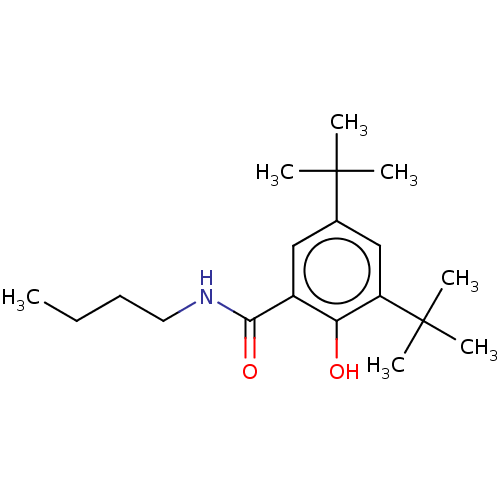
TargetNuclear receptor subfamily 5 group A member 2(Homo sapiens (Human))
Emory University School Of Medicine
Curated by ChEMBL
Emory University School Of Medicine
Curated by ChEMBL
Affinity DataKi: 0.150nMAssay Description:Binding affinity to N-terminal 6-His tagged human LRH-1 LBD (299 to 541 residues) expressed in Escherichia coli BL21-(DE3) assessed as inhibition con...More data for this Ligand-Target Pair
TargetNuclear receptor subfamily 5 group A member 2(Homo sapiens (Human))
Emory University School Of Medicine
Curated by ChEMBL
Emory University School Of Medicine
Curated by ChEMBL
Affinity DataKi: 1.30nMAssay Description:Binding affinity to N-terminal 6-His tagged human LRH-1 LBD (299 to 541 residues) expressed in Escherichia coli BL21-(DE3) assessed as inhibition con...More data for this Ligand-Target Pair
TargetNuclear receptor subfamily 5 group A member 2(Homo sapiens (Human))
Emory University School Of Medicine
Curated by ChEMBL
Emory University School Of Medicine
Curated by ChEMBL
Affinity DataKi: 2.10nMAssay Description:Displacement of 6N-FAM from human LRH1 LBD (299 to 541 residues) expressed in Escherichia coli BL21 pLysS by competitive binding based fluorescence p...More data for this Ligand-Target Pair
TargetNuclear receptor subfamily 5 group A member 2(Homo sapiens (Human))
Emory University School Of Medicine
Curated by ChEMBL
Emory University School Of Medicine
Curated by ChEMBL
Affinity DataKi: 2.70nMAssay Description:Displacement of 6N-FAM from LRH1 LBD (299 to 541 residues) expressed in Escherichia coli BL21 pLysS by competitive binding based fluorescence polariz...More data for this Ligand-Target Pair
TargetNuclear receptor subfamily 5 group A member 2(Homo sapiens (Human))
Emory University School Of Medicine
Curated by ChEMBL
Emory University School Of Medicine
Curated by ChEMBL
Affinity DataKi: 5.30nMAssay Description:Binding affinity to N-terminal 6-His tagged human LRH-1 LBD (299 to 541 residues) expressed in Escherichia coli BL21-(DE3) assessed as inhibition con...More data for this Ligand-Target Pair
TargetNuclear receptor subfamily 5 group A member 2(Homo sapiens (Human))
Emory University School Of Medicine
Curated by ChEMBL
Emory University School Of Medicine
Curated by ChEMBL
Affinity DataKi: 18nMAssay Description:Displacement of 6N-FAM from human LRH1 LBD (299 to 541 residues) expressed in Escherichia coli BL21 pLysS by competitive binding based fluorescence p...More data for this Ligand-Target Pair
TargetNuclear receptor subfamily 5 group A member 2(Homo sapiens (Human))
Emory University School Of Medicine
Curated by ChEMBL
Emory University School Of Medicine
Curated by ChEMBL
Affinity DataKi: 38nMAssay Description:Displacement of 6N-FAM from human LRH1 LBD (299 to 541 residues) expressed in Escherichia coli BL21 pLysS by competitive binding based fluorescence p...More data for this Ligand-Target Pair
TargetNuclear receptor subfamily 5 group A member 2(Homo sapiens (Human))
Emory University School Of Medicine
Curated by ChEMBL
Emory University School Of Medicine
Curated by ChEMBL
Affinity DataKi: 56nMAssay Description:Displacement of 6N-FAM from human LRH1 LBD (299 to 541 residues) expressed in Escherichia coli BL21 pLysS by competitive binding based fluorescence p...More data for this Ligand-Target Pair
TargetNuclear receptor subfamily 5 group A member 2(Homo sapiens (Human))
Emory University School Of Medicine
Curated by ChEMBL
Emory University School Of Medicine
Curated by ChEMBL
Affinity DataKi: 64nMAssay Description:Displacement of 6N-FAM from human LRH1 LBD (299 to 541 residues) expressed in Escherichia coli BL21 pLysS by competitive binding based fluorescence p...More data for this Ligand-Target Pair
TargetNuclear receptor subfamily 5 group A member 2(Homo sapiens (Human))
Emory University School Of Medicine
Curated by ChEMBL
Emory University School Of Medicine
Curated by ChEMBL
Affinity DataKi: 81nMAssay Description:Displacement of 6N-FAM from human LRH1 LBD (299 to 541 residues) expressed in Escherichia coli BL21 pLysS by competitive binding based fluorescence p...More data for this Ligand-Target Pair
TargetNuclear receptor subfamily 5 group A member 2(Homo sapiens (Human))
Emory University School Of Medicine
Curated by ChEMBL
Emory University School Of Medicine
Curated by ChEMBL
Affinity DataKi: 280nMAssay Description:Displacement of 6N-FAM from human LRH1 LBD (299 to 541 residues) expressed in Escherichia coli BL21 pLysS by competitive binding based fluorescence p...More data for this Ligand-Target Pair
TargetNuclear receptor subfamily 5 group A member 2(Homo sapiens (Human))
Emory University School Of Medicine
Curated by ChEMBL
Emory University School Of Medicine
Curated by ChEMBL
Affinity DataKi: 374nMAssay Description:Displacement of 6N-FAM from human LRH1 LBD (299 to 541 residues) expressed in Escherichia coli BL21 pLysS by competitive binding based fluorescence p...More data for this Ligand-Target Pair
Affinity DataKi: 418nMAssay Description:Displacement of 6N-FAM from SF1 LBD (218 to 461 residues) (unknown origin) expressed in Escherichia coli BL21 pLysS by competitive binding based fluo...More data for this Ligand-Target Pair
Affinity DataKi: 670nM ΔG°: -35.8kJ/molepH: 8.2 T: 2°CAssay Description:In brief, both wild-type and mutant hFABP5 were expressed and purified to homogeneity as described above and dialyzed in PBS (pH 8.2). Binding affini...More data for this Ligand-Target Pair
Affinity DataKi: 690nM ΔG°: -35.8kJ/molepH: 8.2 T: 2°CAssay Description:In brief, both wild-type and mutant hFABP5 were expressed and purified to homogeneity as described above and dialyzed in PBS (pH 8.2). Binding affini...More data for this Ligand-Target Pair
Affinity DataKi: 700nM ΔG°: -35.7kJ/molepH: 8.2 T: 2°CAssay Description:In brief, both wild-type and mutant hFABP5 were expressed and purified to homogeneity as described above and dialyzed in PBS (pH 8.2). Binding affini...More data for this Ligand-Target Pair
Affinity DataKi: 850nMAssay Description:Displacement of 6N-FAM from SF1 LBD (218 to 461 residues) (unknown origin) expressed in Escherichia coli BL21 pLysS by competitive binding based fluo...More data for this Ligand-Target Pair
Affinity DataKi: 1.10E+3nM ΔG°: -34.6kJ/molepH: 8.2 T: 2°CAssay Description:In brief, both wild-type and mutant hFABP5 were expressed and purified to homogeneity as described above and dialyzed in PBS (pH 8.2). Binding affini...More data for this Ligand-Target Pair
Affinity DataKi: 1.24E+3nM ΔG°: -34.3kJ/molepH: 8.2 T: 2°CAssay Description:In brief, both wild-type and mutant hFABP5 were expressed and purified to homogeneity as described above and dialyzed in PBS (pH 8.2). Binding affini...More data for this Ligand-Target Pair
Affinity DataKi: 1.26E+3nM ΔG°: -34.2kJ/molepH: 8.2 T: 2°CAssay Description:In brief, both wild-type and mutant hFABP5 were expressed and purified to homogeneity as described above and dialyzed in PBS (pH 8.2). Binding affini...More data for this Ligand-Target Pair
Affinity DataKi: 1.36E+3nM ΔG°: -34.0kJ/molepH: 8.2 T: 2°CAssay Description:In brief, both wild-type and mutant hFABP5 were expressed and purified to homogeneity as described above and dialyzed in PBS (pH 8.2). Binding affini...More data for this Ligand-Target Pair
Affinity DataKi: 1.40E+3nMAssay Description:Displacement of 6N-FAM from SF1 LBD (218 to 461 residues) (unknown origin) expressed in Escherichia coli BL21 pLysS by competitive binding based fluo...More data for this Ligand-Target Pair
Affinity DataKi: 1.64E+3nM ΔG°: -33.6kJ/molepH: 8.2 T: 2°CAssay Description:In brief, both wild-type and mutant hFABP5 were expressed and purified to homogeneity as described above and dialyzed in PBS (pH 8.2). Binding affini...More data for this Ligand-Target Pair
Affinity DataKi: 1.78E+3nM ΔG°: -33.4kJ/molepH: 8.2 T: 2°CAssay Description:In brief, both wild-type and mutant hFABP5 were expressed and purified to homogeneity as described above and dialyzed in PBS (pH 8.2). Binding affini...More data for this Ligand-Target Pair
Affinity DataKi: 2.17E+3nM ΔG°: -32.9kJ/molepH: 8.2 T: 2°CAssay Description:In brief, both wild-type and mutant hFABP5 were expressed and purified to homogeneity as described above and dialyzed in PBS (pH 8.2). Binding affini...More data for this Ligand-Target Pair
TargetNuclear receptor subfamily 5 group A member 2(Homo sapiens (Human))
Emory University School Of Medicine
Curated by ChEMBL
Emory University School Of Medicine
Curated by ChEMBL
Affinity DataKi: 2.40E+3nMAssay Description:Displacement of 6N-FAM from human LRH1 LBD (299 to 541 residues) expressed in Escherichia coli BL21 pLysS by competitive binding based fluorescence p...More data for this Ligand-Target Pair
Affinity DataKi: 2.59E+3nM ΔG°: -32.4kJ/molepH: 8.2 T: 2°CAssay Description:In brief, both wild-type and mutant hFABP5 were expressed and purified to homogeneity as described above and dialyzed in PBS (pH 8.2). Binding affini...More data for this Ligand-Target Pair
Affinity DataKi: 3.06E+3nM ΔG°: -32.0kJ/molepH: 8.2 T: 2°CAssay Description:In brief, both wild-type and mutant hFABP5 were expressed and purified to homogeneity as described above and dialyzed in PBS (pH 8.2). Binding affini...More data for this Ligand-Target Pair
Affinity DataKi: 3.30E+3nMAssay Description:Displacement of 6N-FAM from SF1 LBD (218 to 461 residues) (unknown origin) expressed in Escherichia coli BL21 pLysS by competitive binding based fluo...More data for this Ligand-Target Pair
Affinity DataKi: 3.47E+3nM ΔG°: -31.7kJ/molepH: 8.2 T: 2°CAssay Description:In brief, both wild-type and mutant hFABP5 were expressed and purified to homogeneity as described above and dialyzed in PBS (pH 8.2). Binding affini...More data for this Ligand-Target Pair
Affinity DataKi: 5.01E+3nM ΔG°: -30.8kJ/molepH: 8.2 T: 2°CAssay Description:In brief, both wild-type and mutant hFABP5 were expressed and purified to homogeneity as described above and dialyzed in PBS (pH 8.2). Binding affini...More data for this Ligand-Target Pair
TargetNuclear receptor subfamily 5 group A member 2(Homo sapiens (Human))
Emory University School Of Medicine
Curated by ChEMBL
Emory University School Of Medicine
Curated by ChEMBL
Affinity DataKi: 7.00E+3nMAssay Description:Displacement of 6N-FAM from human LRH1 LBD (299 to 541 residues) expressed in Escherichia coli BL21 pLysS by competitive binding based fluorescence p...More data for this Ligand-Target Pair
Affinity DataKi: 7.80E+3nMAssay Description:Displacement of 6N-FAM from SF1 LBD (218 to 461 residues) (unknown origin) expressed in Escherichia coli BL21 pLysS by competitive binding based fluo...More data for this Ligand-Target Pair
Affinity DataKi: 9.37E+3nM ΔG°: -29.2kJ/molepH: 8.2 T: 2°CAssay Description:In brief, both wild-type and mutant hFABP5 were expressed and purified to homogeneity as described above and dialyzed in PBS (pH 8.2). Binding affini...More data for this Ligand-Target Pair
Affinity DataKi: 3.47E+4nMAssay Description:Displacement of 6N-FAM from SF1 LBD (218 to 461 residues) (unknown origin) expressed in Escherichia coli BL21 pLysS by competitive binding based fluo...More data for this Ligand-Target Pair
Affinity DataIC50: 30nMAssay Description:Inhibition of Gal4-fused human SF1 LBD (198 to 462 residues) expressed in CHO-K1 cellsMore data for this Ligand-Target Pair
Affinity DataKd: 6.57E+3nMpH: 8.2 T: 2°CAssay Description:In brief, both wild-type and mutant hFABP5 were expressed and purified to homogeneity as described above and dialyzed in PBS (pH 8.2). Binding affini...More data for this Ligand-Target Pair
Affinity DataKd: 1.26E+4nMpH: 8.2 T: 2°CAssay Description:In brief, both wild-type and mutant hFABP5 were expressed and purified to homogeneity as described above and dialyzed in PBS (pH 8.2). Binding affini...More data for this Ligand-Target Pair
Affinity DataKd: 6.10E+3nMpH: 8.2 T: 2°CAssay Description:In brief, both wild-type and mutant hFABP5 were expressed and purified to homogeneity as described above and dialyzed in PBS (pH 8.2). Binding affini...More data for this Ligand-Target Pair
TargetNuclear receptor subfamily 5 group A member 2(Homo sapiens (Human))
Emory University School Of Medicine
Curated by ChEMBL
Emory University School Of Medicine
Curated by ChEMBL
Affinity DataEC50: 1.30E+4nMAssay Description:Agonist activity at recombinant full length human LRH1 expressed in HeLa cells after 24 hrs by dual-glo luciferase reporter gene assayMore data for this Ligand-Target Pair
TargetNuclear receptor subfamily 5 group A member 2(Homo sapiens (Human))
Emory University School Of Medicine
Curated by ChEMBL
Emory University School Of Medicine
Curated by ChEMBL
Affinity DataEC50: >1.00E+5nMAssay Description:Agonist activity at recombinant full length human LRH1 expressed in HeLa cells after 24 hrs by dual-glo luciferase reporter gene assayMore data for this Ligand-Target Pair
TargetNuclear receptor subfamily 5 group A member 2(Homo sapiens (Human))
Emory University School Of Medicine
Curated by ChEMBL
Emory University School Of Medicine
Curated by ChEMBL
Affinity DataEC50: 1.10E+3nMAssay Description:Agonist activity at recombinant full length human LRH1 expressed in HeLa cells after 24 hrs by dual-glo luciferase reporter gene assayMore data for this Ligand-Target Pair
TargetNuclear receptor subfamily 5 group A member 2(Homo sapiens (Human))
Emory University School Of Medicine
Curated by ChEMBL
Emory University School Of Medicine
Curated by ChEMBL
Affinity DataEC50: 5.00E+3nMAssay Description:Agonist activity at recombinant full length human LRH1 expressed in HeLa cells after 24 hrs by dual-glo luciferase reporter gene assayMore data for this Ligand-Target Pair
TargetNuclear receptor subfamily 5 group A member 2(Homo sapiens (Human))
Emory University School Of Medicine
Curated by ChEMBL
Emory University School Of Medicine
Curated by ChEMBL
Affinity DataEC50: 400nMAssay Description:Agonist activity at recombinant full length human LRH1 expressed in HeLa cells after 24 hrs by dual-glo luciferase reporter gene assayMore data for this Ligand-Target Pair
TargetNuclear receptor subfamily 5 group A member 2(Homo sapiens (Human))
Emory University School Of Medicine
Curated by ChEMBL
Emory University School Of Medicine
Curated by ChEMBL
Affinity DataEC50: 900nMAssay Description:Agonist activity at human full length LRH1 transfected in human HeLa cells incubated for 24 hrs by renilla luciferase reporter gene assayMore data for this Ligand-Target Pair
TargetNuclear receptor subfamily 5 group A member 2(Homo sapiens (Human))
Emory University School Of Medicine
Curated by ChEMBL
Emory University School Of Medicine
Curated by ChEMBL
Affinity DataEC50: 100nMAssay Description:Agonist activity at human full length LRH1 transfected in human HeLa cells incubated for 24 hrs by renilla luciferase reporter gene assayMore data for this Ligand-Target Pair
TargetNuclear receptor subfamily 5 group A member 2(Homo sapiens (Human))
Emory University School Of Medicine
Curated by ChEMBL
Emory University School Of Medicine
Curated by ChEMBL
Affinity DataEC50: 1.30E+4nMAssay Description:Agonist activity at human full length LRH1 transfected in human HeLa cells incubated for 24 hrs by renilla luciferase reporter gene assayMore data for this Ligand-Target Pair
TargetNuclear receptor subfamily 5 group A member 2(Homo sapiens (Human))
Emory University School Of Medicine
Curated by ChEMBL
Emory University School Of Medicine
Curated by ChEMBL
Affinity DataEC50: 15nMAssay Description:Agonist activity at human full length LRH1 transfected in human HeLa cells incubated for 24 hrs by renilla luciferase reporter gene assayMore data for this Ligand-Target Pair
TargetNuclear receptor subfamily 5 group A member 2(Homo sapiens (Human))
Emory University School Of Medicine
Curated by ChEMBL
Emory University School Of Medicine
Curated by ChEMBL
Affinity DataEC50: 1.00E+3nMAssay Description:Agonist activity at human full length LRH1 transfected in human HeLa cells incubated for 24 hrs by renilla luciferase reporter gene assayMore data for this Ligand-Target Pair
TargetNuclear receptor subfamily 5 group A member 2(Homo sapiens (Human))
Emory University School Of Medicine
Curated by ChEMBL
Emory University School Of Medicine
Curated by ChEMBL
Affinity DataEC50: 800nMAssay Description:Agonist activity at human full length LRH1 transfected in human HeLa cells incubated for 24 hrs by renilla luciferase reporter gene assayMore data for this Ligand-Target Pair

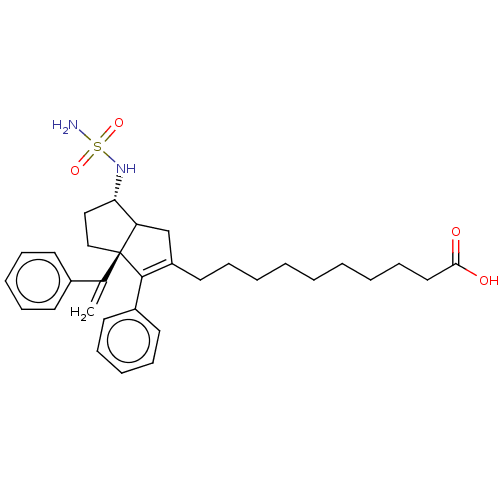
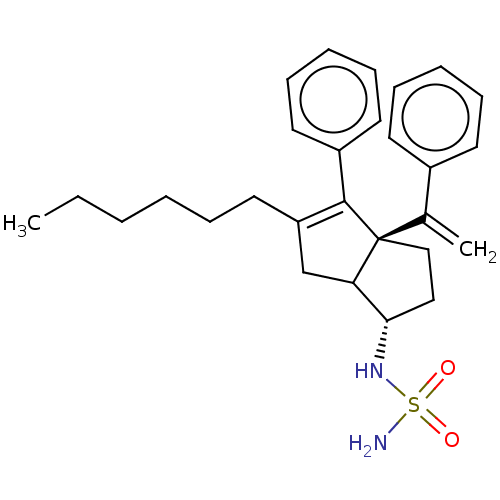



 3D Structure (crystal)
3D Structure (crystal)