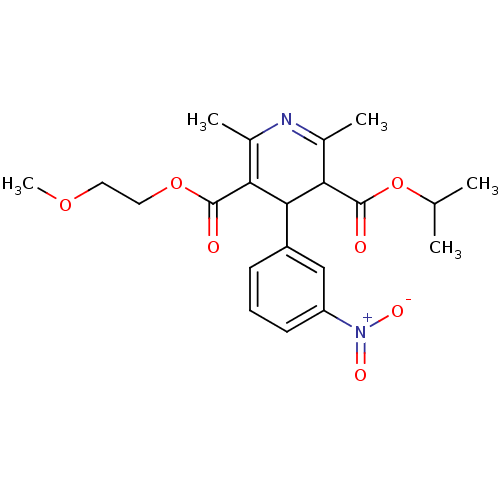
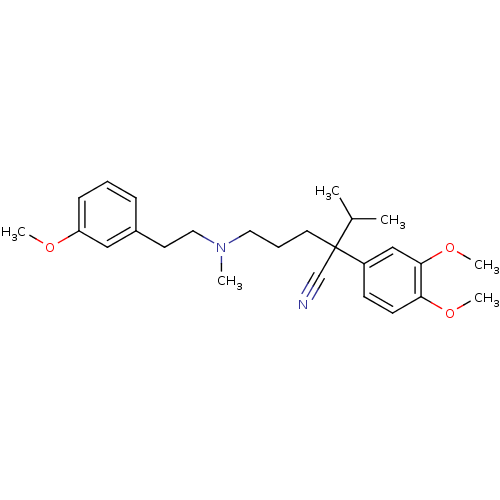
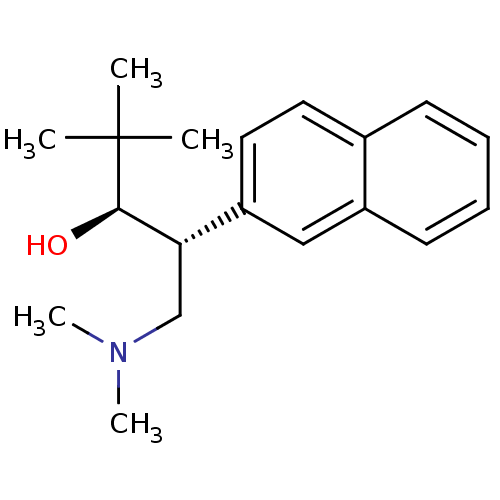
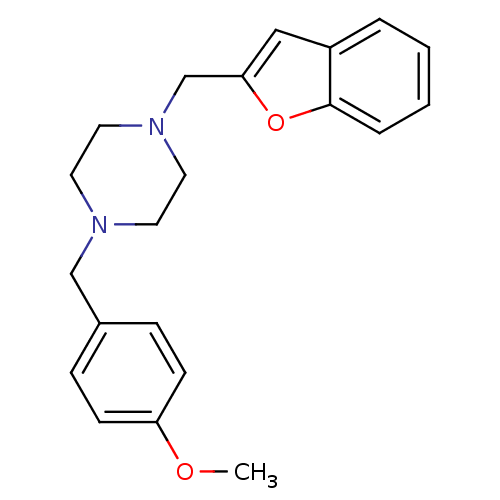
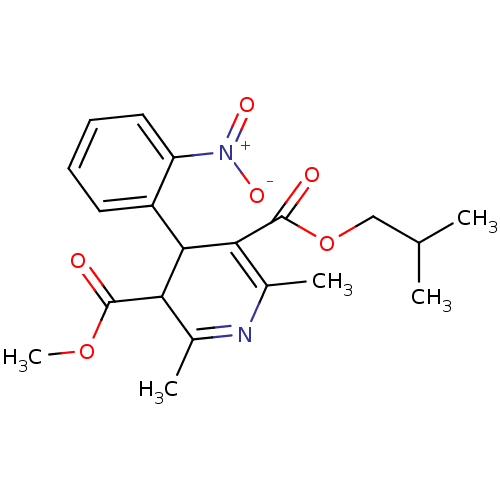
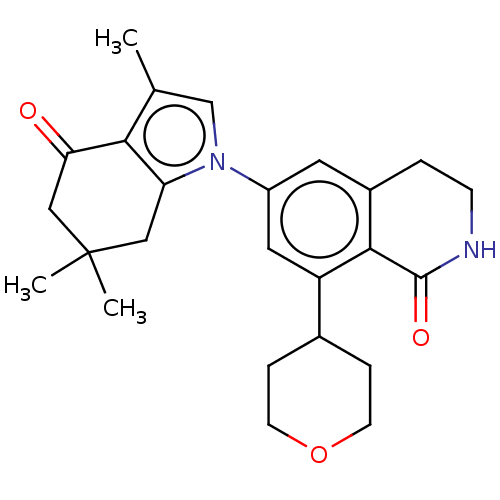
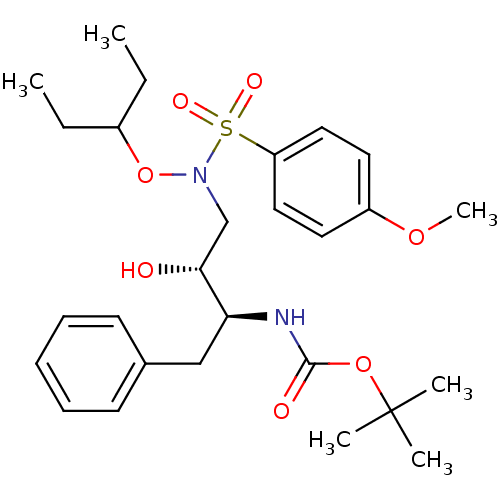
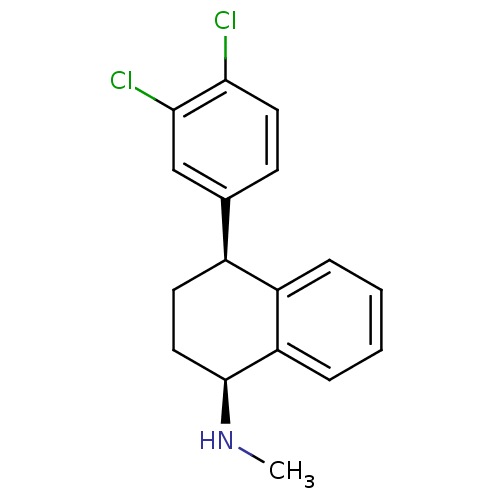
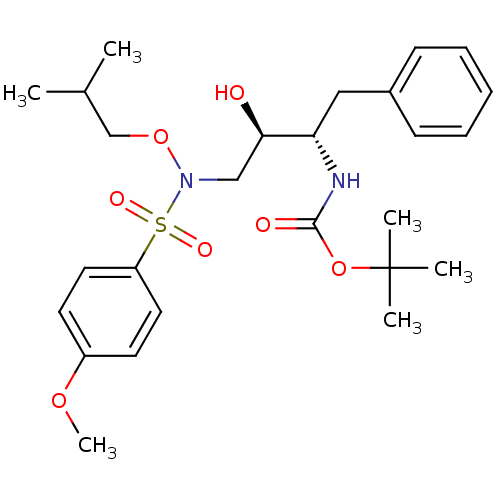
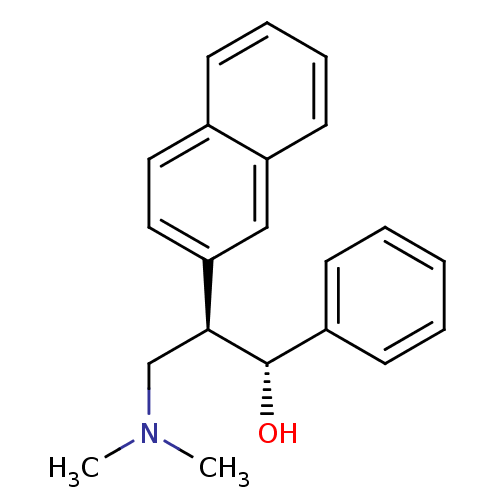
TargetDimer of Gag-Pol polyprotein [501-599,Q508K,L534I,L564I,C568A,C596A](Human immunodeficiency virus type 1)
Glaxosmithkline
Glaxosmithkline
Affinity DataKi: <0.00500nM IC50: 3nMAssay Description:The Ki values were determined using fluorogenic substrate, 2-(aminobenzoyl)-Thr-Ile-Nle-Phe(p-NO2)-Gln-ArgNH2. A standard curve relating changes in f...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [501-599,Q508K,L534I,L564I,C568A,C596A](Human immunodeficiency virus type 1)
Glaxosmithkline
Glaxosmithkline
Affinity DataKi: <0.00500nM IC50: 5nMAssay Description:The Ki values were determined using fluorogenic substrate, 2-(aminobenzoyl)-Thr-Ile-Nle-Phe(p-NO2)-Gln-ArgNH2. A standard curve relating changes in f...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [501-599,Q508K,L534I,L564I,C568A,C596A](Human immunodeficiency virus type 1)
Glaxosmithkline
Glaxosmithkline
Affinity DataKi: <0.00500nM ΔG°: <-64.5kJ/mole IC50: 6nMpH: 6.8 T: 2°CAssay Description:The Ki values were determined using fluorogenic substrate, 2-(aminobenzoyl)-Thr-Ile-Nle-Phe(p-NO2)-Gln-ArgNH2. A standard curve relating changes in f...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [501-599,Q508K,L534I,L564I,C568A,C596A](Human immunodeficiency virus type 1)
Glaxosmithkline
Glaxosmithkline
Affinity DataKi: <0.00500nM IC50: 7nMAssay Description:The Ki values were determined using fluorogenic substrate, 2-(aminobenzoyl)-Thr-Ile-Nle-Phe(p-NO2)-Gln-ArgNH2. A standard curve relating changes in f...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [501-599,Q508K,L534I,L564I,C568A,C596A](Human immunodeficiency virus type 1)
Glaxosmithkline
Glaxosmithkline
Affinity DataKi: <0.00500nM IC50: 7nMAssay Description:The Ki values were determined using fluorogenic substrate, 2-(aminobenzoyl)-Thr-Ile-Nle-Phe(p-NO2)-Gln-ArgNH2. A standard curve relating changes in f...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [501-599,Q508K,L534I,L564I,C568A,C596A](Human immunodeficiency virus type 1)
Glaxosmithkline
Glaxosmithkline
Affinity DataKi: <0.00500nM IC50: 9nMAssay Description:The Ki values were determined using fluorogenic substrate, 2-(aminobenzoyl)-Thr-Ile-Nle-Phe(p-NO2)-Gln-ArgNH2. A standard curve relating changes in f...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [501-599,Q508K,L534I,L564I,C568A,C596A](Human immunodeficiency virus type 1)
Glaxosmithkline
Glaxosmithkline
Affinity DataKi: 0.00500nM IC50: 11nMAssay Description:The Ki values were determined using fluorogenic substrate, 2-(aminobenzoyl)-Thr-Ile-Nle-Phe(p-NO2)-Gln-ArgNH2. A standard curve relating changes in f...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [501-599,Q508K,L534I,L564I,C568A,C596A](Human immunodeficiency virus type 1)
Glaxosmithkline
Glaxosmithkline
Affinity DataKi: <0.00500nM IC50: 12nMAssay Description:The Ki values were determined using fluorogenic substrate, 2-(aminobenzoyl)-Thr-Ile-Nle-Phe(p-NO2)-Gln-ArgNH2. A standard curve relating changes in f...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [501-599,Q508K,L534I,L564I,C568A,C596A](Human immunodeficiency virus type 1)
Glaxosmithkline
Glaxosmithkline
Affinity DataKi: <0.00500nM IC50: 13nMAssay Description:The Ki values were determined using fluorogenic substrate, 2-(aminobenzoyl)-Thr-Ile-Nle-Phe(p-NO2)-Gln-ArgNH2. A standard curve relating changes in f...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [501-599,Q508K,L534I,L564I,C568A,C596A](Human immunodeficiency virus type 1)
Glaxosmithkline
Glaxosmithkline
Affinity DataKi: <0.00500nM IC50: 17nMAssay Description:The Ki values were determined using fluorogenic substrate, 2-(aminobenzoyl)-Thr-Ile-Nle-Phe(p-NO2)-Gln-ArgNH2. A standard curve relating changes in f...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [501-599,Q508K,L534I,L564I,C568A,C596A](Human immunodeficiency virus type 1)
Glaxosmithkline
Glaxosmithkline
Affinity DataKi: <0.00500nM IC50: 20nMAssay Description:The Ki values were determined using fluorogenic substrate, 2-(aminobenzoyl)-Thr-Ile-Nle-Phe(p-NO2)-Gln-ArgNH2. A standard curve relating changes in f...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [501-599,Q508K,L534I,L564I,C568A,C596A](Human immunodeficiency virus type 1)
Glaxosmithkline
Glaxosmithkline
Affinity DataKi: <0.00500nM ΔG°: <-64.5kJ/mole IC50: 24nMpH: 6.8 T: 2°CAssay Description:The Ki values were determined using fluorogenic substrate, 2-(aminobenzoyl)-Thr-Ile-Nle-Phe(p-NO2)-Gln-ArgNH2. A standard curve relating changes in f...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [501-599,Q508K,L534I,L564I,C568A,C596A](Human immunodeficiency virus type 1)
Glaxosmithkline
Glaxosmithkline
Affinity DataKi: 0.0180nM IC50: 8nMAssay Description:The Ki values were determined using fluorogenic substrate, 2-(aminobenzoyl)-Thr-Ile-Nle-Phe(p-NO2)-Gln-ArgNH2. A standard curve relating changes in f...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [501-599,Q508K,L534I,L564I,C568A,C596A](Human immunodeficiency virus type 1)
Glaxosmithkline
Glaxosmithkline
Affinity DataKi: 0.0400nM ΔG°: -59.3kJ/mole IC50: 150nMpH: 6.8 T: 2°CAssay Description:The Ki values were determined using fluorogenic substrate, 2-(aminobenzoyl)-Thr-Ile-Nle-Phe(p-NO2)-Gln-ArgNH2. A standard curve relating changes in f...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [501-599,Q508K,L534I,L564I,C568A,C596A](Human immunodeficiency virus type 1)
Glaxosmithkline
Glaxosmithkline
Affinity DataKi: 0.0900nM ΔG°: -57.3kJ/mole IC50: 203nMpH: 6.8 T: 2°CAssay Description:The Ki values were determined using fluorogenic substrate, 2-(aminobenzoyl)-Thr-Ile-Nle-Phe(p-NO2)-Gln-ArgNH2. A standard curve relating changes in f...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [501-599,Q508K,L534I,L564I,C568A,C596A](Human immunodeficiency virus type 1)
Glaxosmithkline
Glaxosmithkline
Affinity DataKi: 0.190nM ΔG°: -55.5kJ/mole IC50: 738nMpH: 6.8 T: 2°CAssay Description:The Ki values were determined using fluorogenic substrate, 2-(aminobenzoyl)-Thr-Ile-Nle-Phe(p-NO2)-Gln-ArgNH2. A standard curve relating changes in f...More data for this Ligand-Target Pair
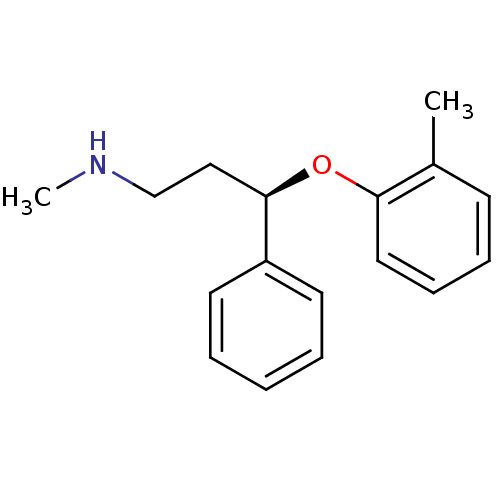
Affinity DataKi: 0.700nMAssay Description:Inhibition of [3H]- NE reuptake into rat hippocampal synaptosomesMore data for this Ligand-Target Pair
TargetSodium-dependent serotonin transporter(Rattus norvegicus (rat))
Hong Kong University Of Science And Technology
Curated by ChEMBL
Hong Kong University Of Science And Technology
Curated by ChEMBL
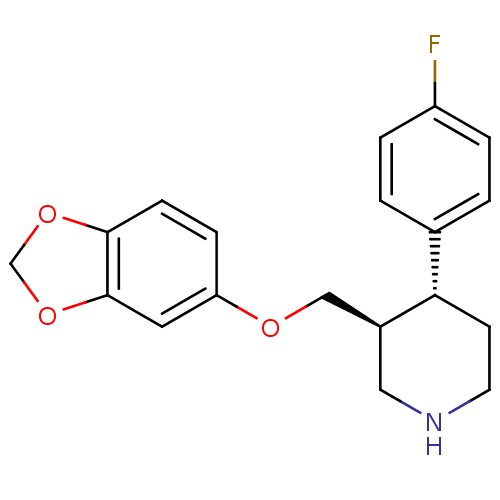
Affinity DataKi: 0.730nMAssay Description:Inhibition of [3H]5-HT reuptake into rat frontal cortex synaptosomesMore data for this Ligand-Target Pair
TargetSodium-dependent dopamine transporter(Homo sapiens (Human))
The University Of Sydney
Curated by ChEMBL
The University Of Sydney
Curated by ChEMBL
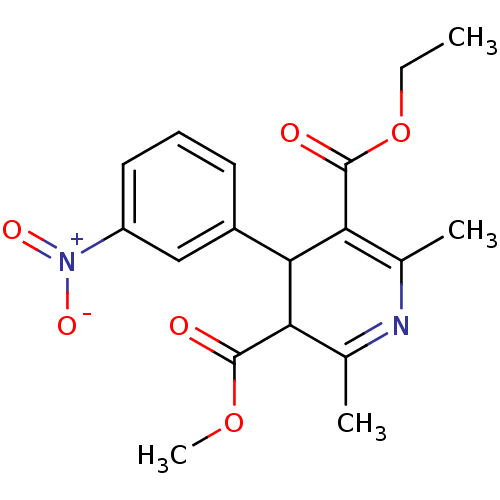
Affinity DataKi: 1.20nMAssay Description:Displacement of [3H]WIN-35428 form human DAT expressed in HEK cell membranesMore data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [501-599,Q508K,L534I,L564I,C568A,C596A](Human immunodeficiency virus type 1)
Glaxosmithkline
Glaxosmithkline
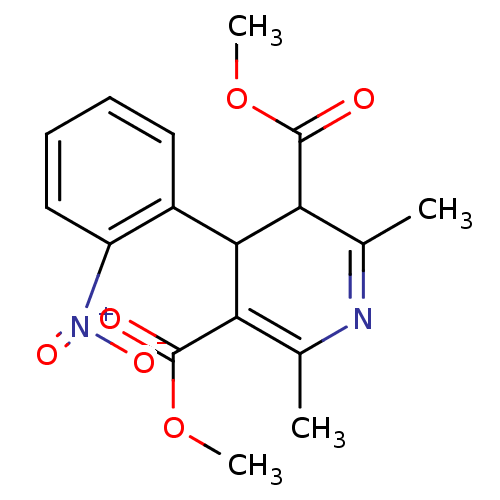
Affinity DataKi: 1.60nM ΔG°: -50.2kJ/molepH: 6.8 T: 2°CAssay Description:The Ki values were determined using fluorogenic substrate, 2-(aminobenzoyl)-Thr-Ile-Nle-Phe(p-NO2)-Gln-ArgNH2. A standard curve relating changes in f...More data for this Ligand-Target Pair
TargetVoltage-dependent L-type calcium channel subunit alpha-1C(RAT)
Johns Hopkins University
Curated by PDSP Ki Database
Johns Hopkins University
Curated by PDSP Ki Database
TargetVoltage-dependent L-type calcium channel subunit alpha-1C(RAT)
Johns Hopkins University
Curated by PDSP Ki Database
Johns Hopkins University
Curated by PDSP Ki Database
TargetVoltage-dependent L-type calcium channel subunit alpha-1C(RAT)
Johns Hopkins University
Curated by PDSP Ki Database
Johns Hopkins University
Curated by PDSP Ki Database
TargetVoltage-dependent L-type calcium channel subunit alpha-1C(RAT)
Johns Hopkins University
Curated by PDSP Ki Database
Johns Hopkins University
Curated by PDSP Ki Database
TargetSodium-dependent serotonin transporter(Rattus norvegicus (rat))
Hong Kong University Of Science And Technology
Curated by ChEMBL
Hong Kong University Of Science And Technology
Curated by ChEMBL
Affinity DataKi: 2.60nMAssay Description:Inhibition of [3H]5-HT reuptake into rat frontal cortex synaptosomesMore data for this Ligand-Target Pair
Affinity DataKi: 2.60nMAssay Description:Displacement of [3H]pentazocine from sigma1 receptor in rat brain homogenateMore data for this Ligand-Target Pair
Affinity DataKi: 2.70nMAssay Description:Displacement of [3H]pentazocine from sigma1 receptor in rat brain homogenateMore data for this Ligand-Target Pair
TargetVoltage-dependent L-type calcium channel subunit alpha-1C(RAT)
Johns Hopkins University
Curated by PDSP Ki Database
Johns Hopkins University
Curated by PDSP Ki Database
TargetDimer of Gag-Pol polyprotein [501-599,Q508K,L534I,L564I,C568A,C596A](Human immunodeficiency virus type 1)
Glaxosmithkline
Glaxosmithkline
Affinity DataKi: 2.80nM ΔG°: -48.8kJ/molepH: 6.8 T: 2°CAssay Description:The Ki values were determined using fluorogenic substrate, 2-(aminobenzoyl)-Thr-Ile-Nle-Phe(p-NO2)-Gln-ArgNH2. A standard curve relating changes in f...More data for this Ligand-Target Pair
Affinity DataKi: 3nMAssay Description:Displacement of FITC-geldanamycin from recombinant HSP90beta (unknown origin) after 2 hrs by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 3nMAssay Description:Displacement of FITC-geldanamycin from recombinant HSP90alpha (unknown origin) after 2 hrs by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [501-599,Q508K,L534I,L564I,C568A,C596A](Human immunodeficiency virus type 1)
Glaxosmithkline
Glaxosmithkline
Affinity DataKi: 3.20nM ΔG°: -48.5kJ/molepH: 6.8 T: 2°CAssay Description:The Ki values were determined using fluorogenic substrate, 2-(aminobenzoyl)-Thr-Ile-Nle-Phe(p-NO2)-Gln-ArgNH2. A standard curve relating changes in f...More data for this Ligand-Target Pair
TargetSodium-dependent serotonin transporter(Rattus norvegicus (rat))
Hong Kong University Of Science And Technology
Curated by ChEMBL
Hong Kong University Of Science And Technology
Curated by ChEMBL
Affinity DataKi: 3.40nMAssay Description:Inhibition of [3H]5-HT reuptake into rat frontal cortex synaptosomesMore data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [501-599,Q508K,L534I,L564I,C568A,C596A](Human immunodeficiency virus type 1)
Glaxosmithkline
Glaxosmithkline
Affinity DataKi: 4nM ΔG°: -47.9kJ/molepH: 6.8 T: 2°CAssay Description:The Ki values were determined using fluorogenic substrate, 2-(aminobenzoyl)-Thr-Ile-Nle-Phe(p-NO2)-Gln-ArgNH2. A standard curve relating changes in f...More data for this Ligand-Target Pair
TargetSodium-dependent serotonin transporter(Rattus norvegicus (rat))
Hong Kong University Of Science And Technology
Curated by ChEMBL
Hong Kong University Of Science And Technology
Curated by ChEMBL
Affinity DataKi: 4.10nMAssay Description:Inhibition of [3H]5-HT reuptake into rat frontal cortex synaptosomesMore data for this Ligand-Target Pair
Affinity DataKi: 5nMAssay Description:Displacement of FITC-geldanamycin from recombinant HSP90alpha (unknown origin) after 2 hrs by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 5nMAssay Description:Displacement of FITC-geldanamycin from recombinant HSP90beta (unknown origin) after 2 hrs by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 5nMAssay Description:Displacement of FITC-geldanamycin from recombinant HSP90beta (unknown origin) after 2 hrs by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 5.20nMAssay Description:Displacement of [3H]pentazocine from sigma1 receptor in rat brain homogenateMore data for this Ligand-Target Pair
Affinity DataKi: 6nMAssay Description:Displacement of FITC-geldanamycin from recombinant HSP90alpha (unknown origin) after 2 hrs by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetSodium-dependent dopamine transporter(Rattus norvegicus (rat))
Hong Kong University Of Science And Technology
Curated by ChEMBL
Hong Kong University Of Science And Technology
Curated by ChEMBL
Affinity DataKi: 6.20nMAssay Description:Inhibition of [3H]- DA reuptake into rat striatal synaptosomesMore data for this Ligand-Target Pair
TargetSodium-dependent dopamine transporter(Homo sapiens (Human))
The University Of Sydney
Curated by ChEMBL
The University Of Sydney
Curated by ChEMBL
Affinity DataKi: 7.70nMAssay Description:Inhibition of [3H]- NE reuptake into rat hippocampal synaptosomesMore data for this Ligand-Target Pair
TargetVoltage-dependent L-type calcium channel subunit alpha-1C(RAT)
Johns Hopkins University
Curated by PDSP Ki Database
Johns Hopkins University
Curated by PDSP Ki Database
Affinity DataKi: 9.10nMAssay Description:Displacement of [3H]pentazocine from sigma1 receptor in rat brain homogenateMore data for this Ligand-Target Pair
Affinity DataKi: 9.70nMAssay Description:Binding affinity to 5HT2BMore data for this Ligand-Target Pair
Affinity DataKi: 10nMAssay Description:Inhibition of [3H]- NE reuptake into rat hippocampal synaptosomesMore data for this Ligand-Target Pair
Affinity DataKi: 10nMAssay Description:Displacement of FITC-geldanamycin from recombinant HSP90alpha (unknown origin) after 2 hrs by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 10nMAssay Description:Displacement of FITC-geldanamycin from recombinant HSP90beta (unknown origin) after 2 hrs by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKi: 10.6nMAssay Description:Displacement of [3H]pentazocine from sigma1 receptor in rat brain homogenateMore data for this Ligand-Target Pair



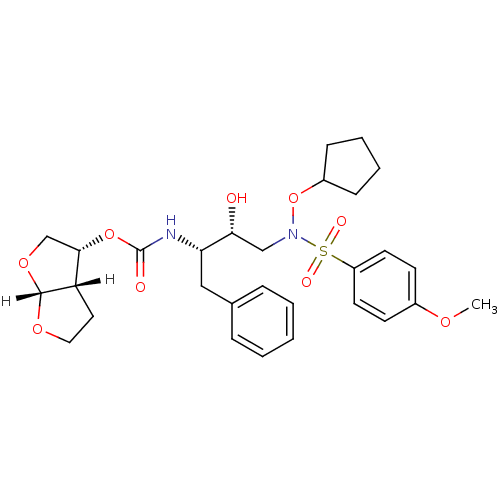










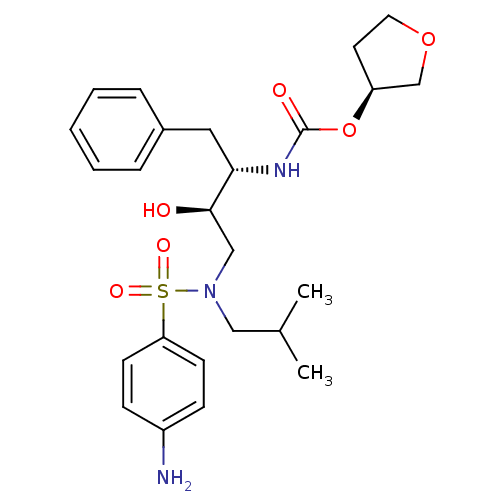
 3D Structure (crystal)
3D Structure (crystal)