Affinity DataKi: 0.0750nMAssay Description:The binding affinity of the compound was measured on histamine H1 receptor using [3H]- mepyramine as radioligand.More data for this Ligand-Target Pair
Affinity DataKi: 0.150nMAssay Description:Binding affinity was evaluated by determining in vitro displacement of [3H]-8-OH-DPAT from the central 5-hydroxytryptamine 1A receptor recognition si...More data for this Ligand-Target Pair
Affinity DataKi: 0.190nMAssay Description:The binding affinity was measured on 5-hydroxytryptamine 3 receptor using [3H]GR-65630 as radioligand.More data for this Ligand-Target Pair
Affinity DataKi: 0.190nMAssay Description:The binding affinity of the compound was measured on alpha2-adrenergic receptor using [3H]- clonidine as radioligand.More data for this Ligand-Target Pair
Affinity DataKi: 0.230nMAssay Description:Displacement of [3H]GR-65630 binding to 5-hydroxytryptamine 3 receptor in rat brain cortical membranesMore data for this Ligand-Target Pair
Affinity DataKi: 0.25nMAssay Description:Displacement of [3H]- GR-65,630 binding to 5-hydroxytryptamine 3 receptor in rat brain cortical membranesMore data for this Ligand-Target Pair
Affinity DataKi: 0.280nMAssay Description:The binding affinity of the compound was measured on glycine receptor using [3H]- strychnine as radioligand.More data for this Ligand-Target Pair
Affinity DataKi: 0.300nMAssay Description:Binding affinity was evaluated by determining in vitro displacement of [3H]-8-OH-DPAT from the central 5-hydroxytryptamine 1A receptor recognition si...More data for this Ligand-Target Pair
Affinity DataKi: 0.320nMAssay Description:Binding affinity was evaluated by determining in vitro displacement of [3H]-8-OH-DPAT from the central 5-hydroxytryptamine 1A receptor recognition si...More data for this Ligand-Target Pair
Affinity DataKi: 0.330nMAssay Description:The binding affinity of the compound was measured on muscarine M1 receptor using [3H]- pirenzepine as radioligand.More data for this Ligand-Target Pair
Affinity DataKi: 0.640nMAssay Description:The binding affinity of the compound was measured on dopamine receptor D1 using [3H]- dopamine as radioligand.More data for this Ligand-Target Pair
Affinity DataKi: 0.640nMAssay Description:The binding affinity of the compound was measured on 5-hydroxytryptamine 1C receptor using [3H]- serotonin as radioligand.More data for this Ligand-Target Pair
Affinity DataKi: 0.75nMAssay Description:Displacement of [3H]- GR-65,630 binding to 5-hydroxytryptamine 3 receptor in rat brain cortical membranesMore data for this Ligand-Target Pair
Affinity DataKi: 0.770nMAssay Description:Displacement of [3H]- GR-65,630 binding to 5-hydroxytryptamine 3 receptor in rat brain cortical membranesMore data for this Ligand-Target Pair
Affinity DataKi: 0.780nMAssay Description:Displacement of [3H]- GR-65,630 binding to 5-hydroxytryptamine 3 receptor in rat brain cortical membranesMore data for this Ligand-Target Pair
Affinity DataKi: 0.800nMAssay Description:Binding affinity was evaluated by determining in vitro displacement of [3H]-8-OH-DPAT from the central 5-hydroxytryptamine 1A receptor recognition si...More data for this Ligand-Target Pair
Affinity DataKi: 1nMAssay Description:Binding affinity was evaluated by determining in vitro displacement of [3H]-8-OH-DPAT from the central 5-hydroxytryptamine 1A receptor recognition si...More data for this Ligand-Target Pair
Affinity DataKi: 1nMAssay Description:Binding affinity was evaluated by determining in vitro displacement of [3H]-8-OH-DPAT from the central 5-hydroxytryptamine 1A receptor recognition si...More data for this Ligand-Target Pair
Affinity DataKi: 1nMAssay Description:Binding affinity was evaluated by determining in vitro displacement of [3H]-8-OH-DPAT from the central 5-hydroxytryptamine 1A receptor recognition si...More data for this Ligand-Target Pair
Affinity DataKi: 1.10nMAssay Description:Binding affinity was evaluated by determining in vitro displacement of [3H]-8-OH-DPAT from the central 5-hydroxytryptamine 1A receptor recognition si...More data for this Ligand-Target Pair
Affinity DataKi: 1.10nMAssay Description:The binding affinity of the compound was measured on mu-opiate receptor using [3H]- naloxone as radioligand.More data for this Ligand-Target Pair
Affinity DataKi: 1.10nMAssay Description:The binding affinity of the compound was measured on kappa-opiate receptor using [3H]- EKC as radioligand.More data for this Ligand-Target Pair
Affinity DataKi: 1.10nMAssay Description:The binding affinity of the compound was measured on 5-hydroxytryptamine 3 receptor using [3H]- GR-65,630 as radioligand.More data for this Ligand-Target Pair
Affinity DataKi: 1.30nMAssay Description:Displacement of [3H]GR-65630 binding to 5-hydroxytryptamine 3 receptor in rat brain cortical membranesMore data for this Ligand-Target Pair
Affinity DataKi: 1.30nMAssay Description:Binding affinity was evaluated by determining in vitro displacement of [3H]-8-OH-DPAT from the central 5-hydroxytryptamine 1A receptor recognition si...More data for this Ligand-Target Pair
Affinity DataKi: 1.60nMAssay Description:The binding affinity of the compound was measured on imidazoline I2 receptor using [3H]- indazoxan as radioligand.More data for this Ligand-Target Pair
Affinity DataKi: 1.60nMAssay Description:The binding affinity was measured on muscarine M2 receptor using [3H]- N-Me-SCOPOL as radioligand.More data for this Ligand-Target Pair
Affinity DataKi: 1.60nMAssay Description:Displacement of [3H]- GR-65,630 binding to 5-hydroxytryptamine 3 receptor in rat brain cortical membranesMore data for this Ligand-Target Pair
Affinity DataKi: 1.60nMAssay Description:Binding affinity was evaluated by determining in vitro displacement of [3H]-8-OH-DPAT from the central 5-hydroxytryptamine 1A receptor recognition si...More data for this Ligand-Target Pair
Affinity DataKi: 1.70nMAssay Description:Binding affinity was evaluated by determining in vitro displacement of [3H]-8-OH-DPAT from the central 5-hydroxytryptamine 1A receptor recognition si...More data for this Ligand-Target Pair
Affinity DataKi: 1.80nMAssay Description:Binding affinity was evaluated by determining in vitro displacement of [3H]-8-OH-DPAT from the central 5-hydroxytryptamine 1A receptor recognition si...More data for this Ligand-Target Pair
Affinity DataKi: 2.5nMAssay Description:Binding affinity was evaluated by determining in vitro displacement of [3H]-8-OH-DPAT from the central 5-hydroxytryptamine 1A receptor recognition si...More data for this Ligand-Target Pair
Affinity DataKi: 3.10nMAssay Description:In vitro displacement of [3H]-spiperone from Dopamine receptor D2 binding site in rat striatum.More data for this Ligand-Target Pair
Affinity DataKi: 3.30nMAssay Description:In vitro displacement of [3H]-spiperone from Dopamine receptor D2 binding site in rat striatum.More data for this Ligand-Target Pair
Affinity DataKi: 3.30nMAssay Description:In vitro displacement of [3H]-spiperone from Dopamine receptor D2 binding site in rat striatum.More data for this Ligand-Target Pair
Affinity DataKi: 3.40nMAssay Description:In vitro displacement of [3H]-spiperone from Dopamine receptor D2 binding site in rat striatum.More data for this Ligand-Target Pair
Affinity DataKi: 3.40nMAssay Description:Binding affinity was evaluated by determining in vitro displacement of [3H]-8-OH-DPAT from the central 5-hydroxytryptamine 1A receptor recognition si...More data for this Ligand-Target Pair
Affinity DataKi: 4.10nMAssay Description:In vitro displacement of [3H]-spiperone from Dopamine receptor D2 binding site in rat striatum.More data for this Ligand-Target Pair
Affinity DataKi: 4.5nMAssay Description:Binding affinity was evaluated by determining in vitro displacement of [3H]-8-OH-DPAT from the central 5-hydroxytryptamine 1A receptor recognition si...More data for this Ligand-Target Pair
Affinity DataKi: 4.90nMAssay Description:Binding affinity was evaluated by determining in vitro displacement of [3H]-8-OH-DPAT from the central 5-hydroxytryptamine 1A receptor recognition si...More data for this Ligand-Target Pair
Affinity DataKi: 5nMAssay Description:In vitro displacement of [3H]-spiperone from Dopamine receptor D2 binding site in rat striatum.More data for this Ligand-Target Pair
Affinity DataKi: 5.30nMAssay Description:In vitro displacement of [3H]-spiperone from Dopamine receptor D2 binding site in rat striatum.More data for this Ligand-Target Pair
Affinity DataKi: 6.40nMAssay Description:In vitro displacement of [3H]-spiperone from Dopamine receptor D2 binding site in rat striatum.More data for this Ligand-Target Pair
Affinity DataKi: 10nMAssay Description:In vitro displacement of [3H]-spiperone from Dopamine receptor D2 binding site in rat striatum.More data for this Ligand-Target Pair
Affinity DataKi: 13nMAssay Description:In vitro displacement of [3H]-spiperone from Dopamine receptor D2 binding site in rat striatum.More data for this Ligand-Target Pair
Affinity DataKi: 13nMAssay Description:Binding affinity was evaluated by determining in vitro displacement of [3H]-8-OH-DPAT from the central 5-hydroxytryptamine 1A receptor recognition si...More data for this Ligand-Target Pair
Affinity DataKi: 14nMAssay Description:In vitro displacement of [3H]-spiperone from Dopamine receptor D2 binding site in rat striatum.More data for this Ligand-Target Pair
Affinity DataKi: 15nMAssay Description:Binding affinity was evaluated by determining in vitro displacement of [3H]-8-OH-DPAT from the central 5-hydroxytryptamine 1A receptor recognition si...More data for this Ligand-Target Pair
Affinity DataKi: 16nMAssay Description:Binding affinity was evaluated by determining in vitro displacement of [3H]-8-OH-DPAT from the central 5-hydroxytryptamine 1A receptor recognition si...More data for this Ligand-Target Pair
Affinity DataKi: 17nMAssay Description:In vitro displacement of [3H]-spiperone from Dopamine receptor D2 binding site in rat striatum.More data for this Ligand-Target Pair


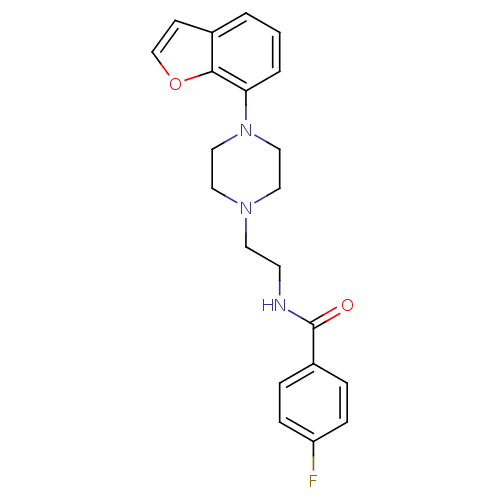
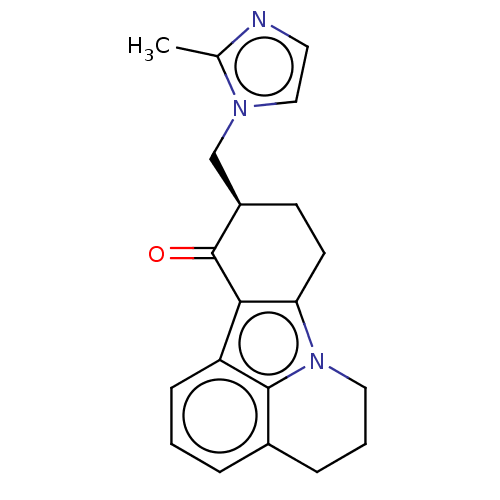


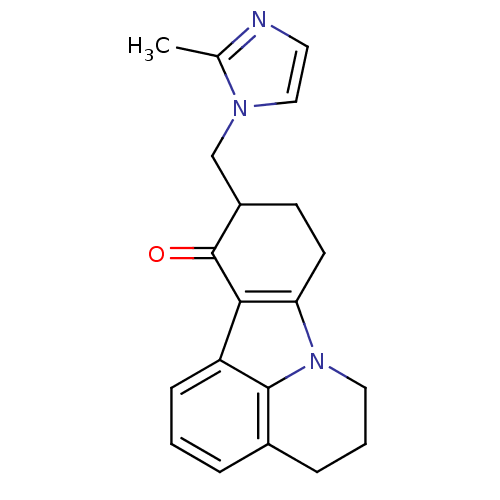
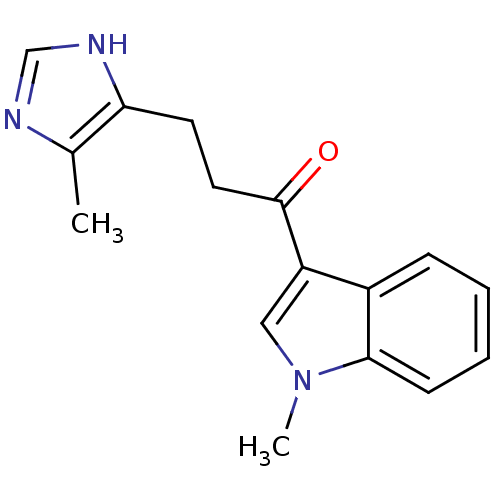


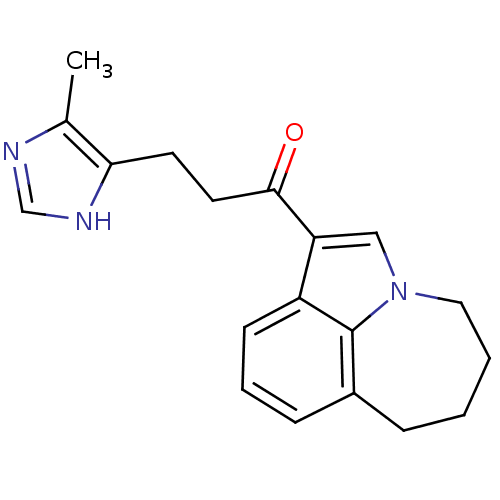
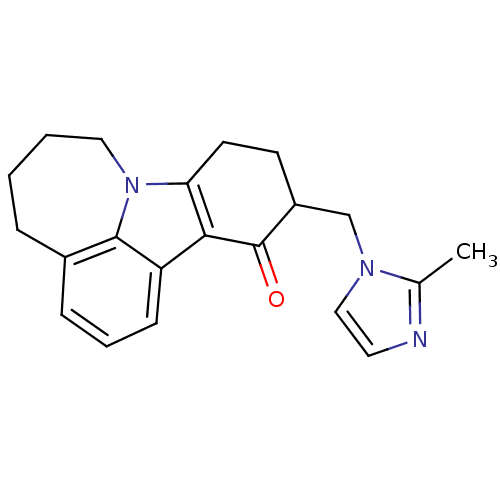

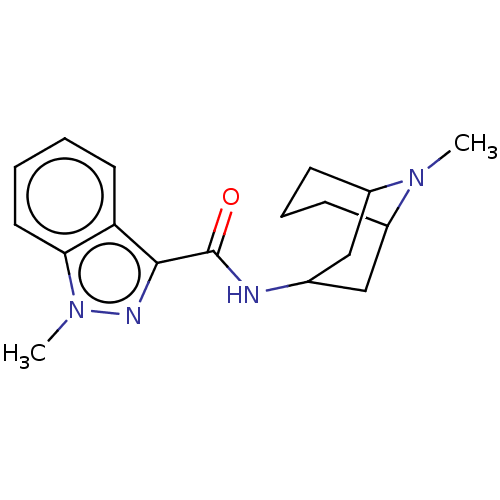
 3D Structure (crystal)
3D Structure (crystal)