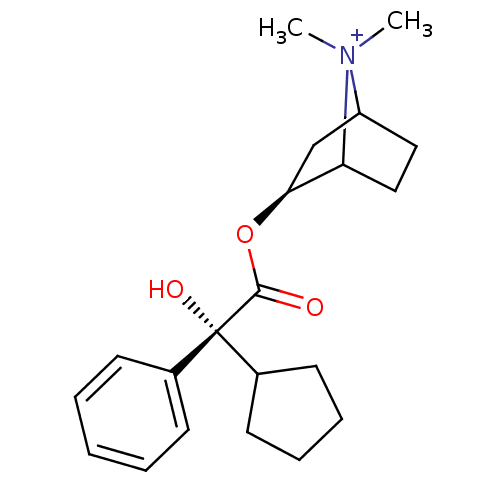
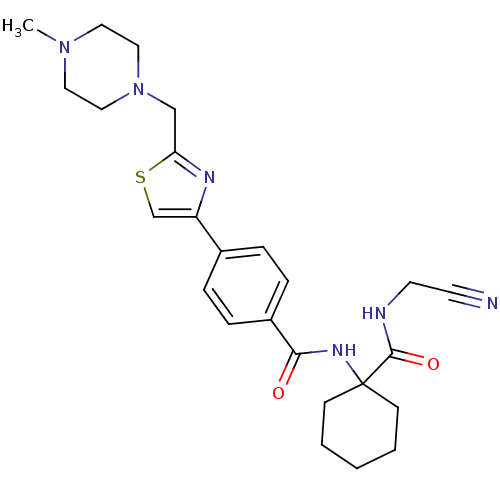
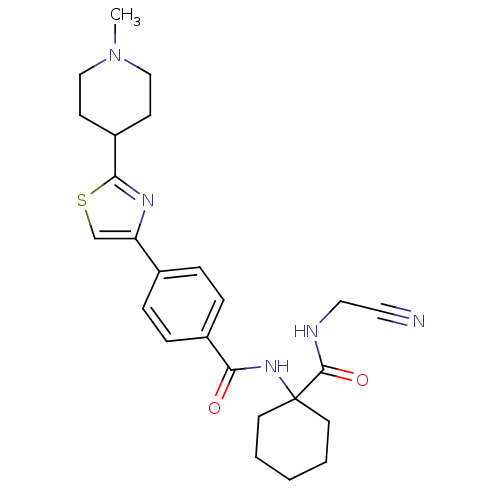
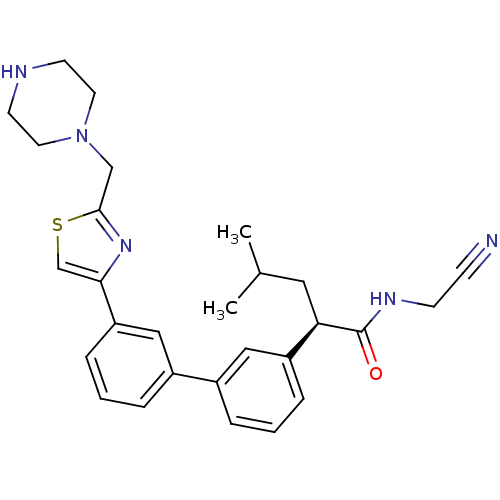
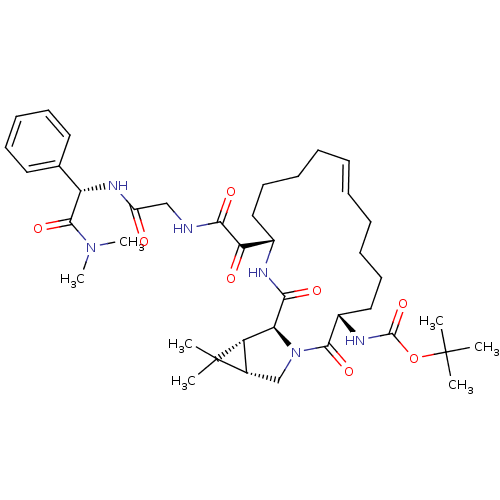
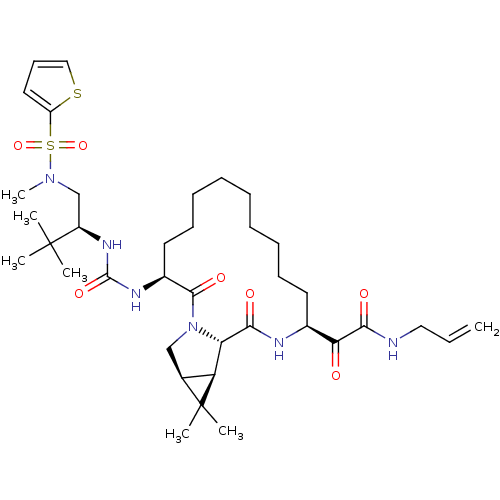
Affinity DataKi: 0.160nM ΔG°: -55.3kJ/mole IC50: 0.230nMT: 2°CAssay Description:Radioligand binding studies were carried out with M3 receptor cell homogenates as described (Peralta et al., The EMBO Journal 6, 3923-3929, (1987)). ...More data for this Ligand-Target Pair
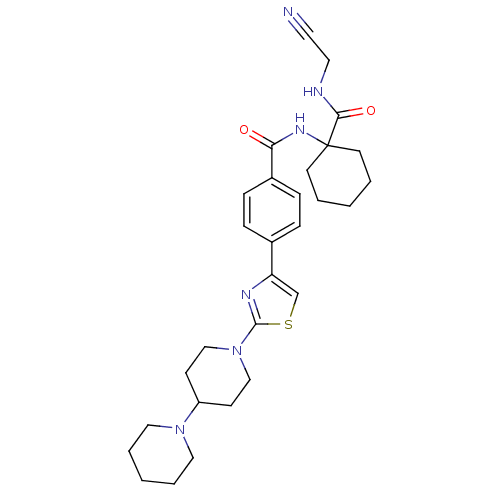
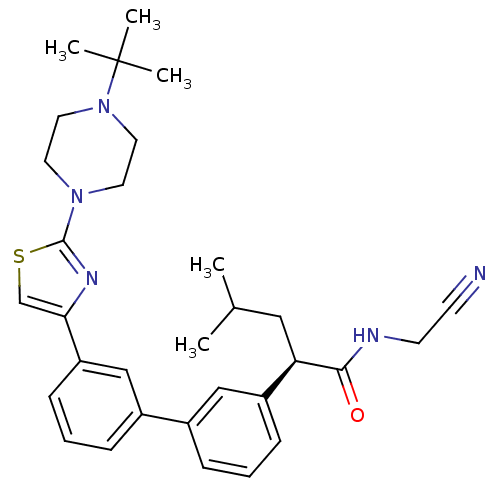
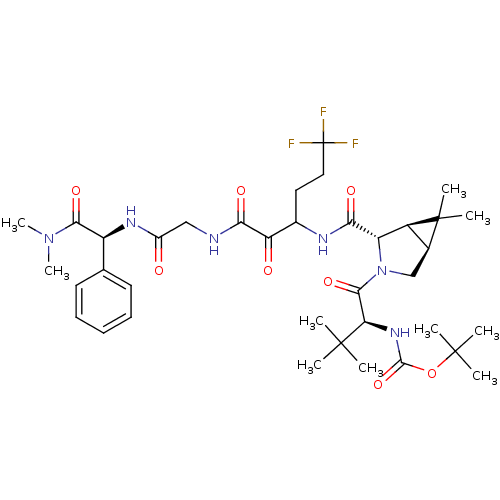
Affinity DataKi: 0.180nM ΔG°: -55.1kJ/mole IC50: 0.25nMT: 2°CAssay Description:Radioligand binding studies were carried out with M3 receptor cell homogenates as described (Peralta et al., The EMBO Journal 6, 3923-3929, (1987)). ...More data for this Ligand-Target Pair
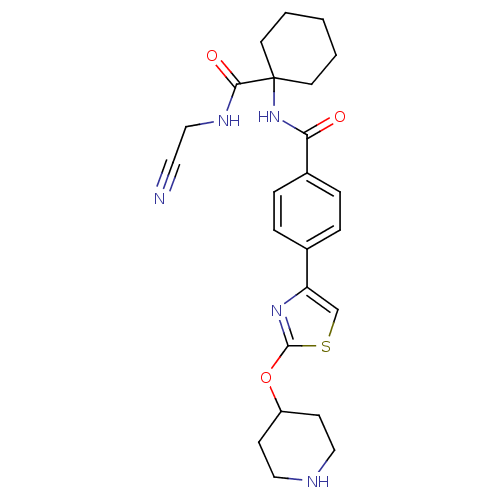
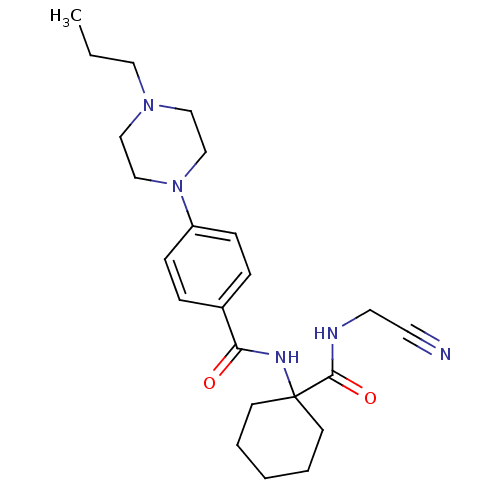
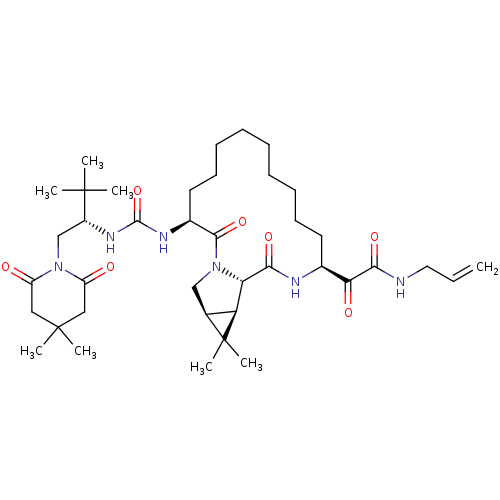
Affinity DataKi: 0.180nM ΔG°: -55.1kJ/mole IC50: 0.25nMT: 2°CAssay Description:Radioligand binding studies were carried out with M3 receptor cell homogenates as described (Peralta et al., The EMBO Journal 6, 3923-3929, (1987)). ...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
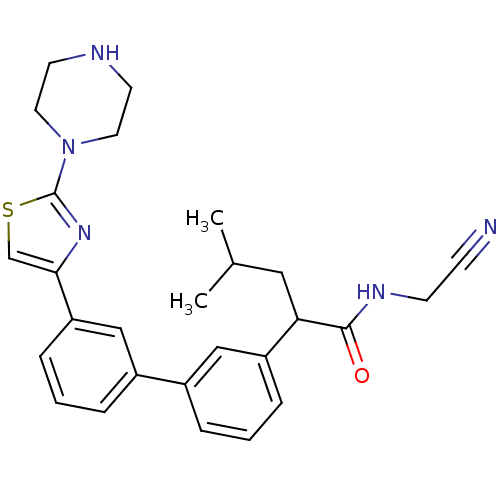
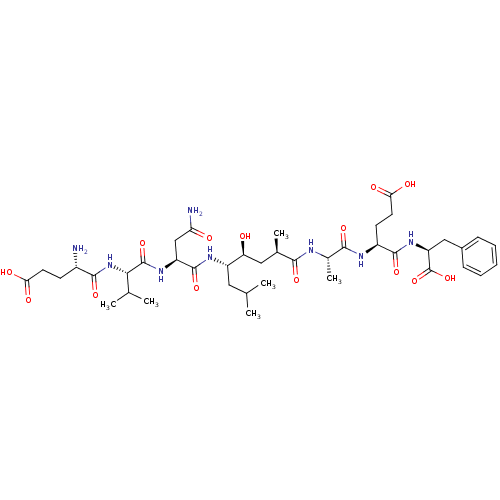
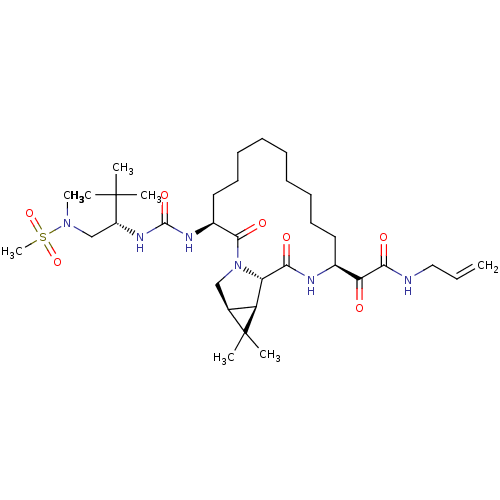
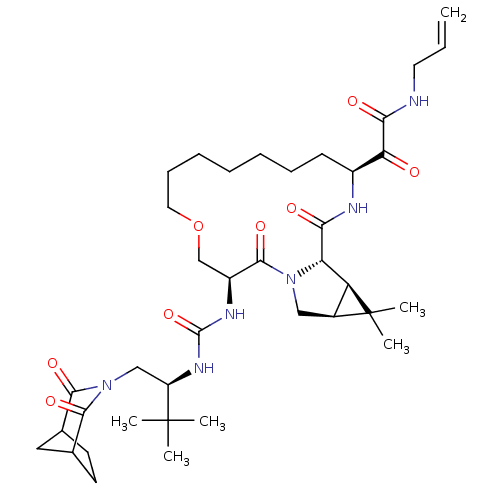
Affinity DataKi: 0.200nM ΔG°: -56.3kJ/molepH: 6.5 T: 2°CAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
Affinity DataKi: 0.200nMAssay Description:Inhibitory constant against rabbit cathepsin K using Z-Phe-Arg-AMC substrateMore data for this Ligand-Target Pair
Affinity DataKi: 0.220nM ΔG°: -54.6kJ/mole IC50: 0.300nMT: 2°CAssay Description:Radioligand binding studies were carried out with M3 receptor cell homogenates as described (Peralta et al., The EMBO Journal 6, 3923-3929, (1987)). ...More data for this Ligand-Target Pair
Affinity DataKi: 0.25nMAssay Description:Inhibitory constant against rabbit cathepsin K using Z-Phe-Arg-AMC substrateMore data for this Ligand-Target Pair
Affinity DataKi: 0.290nMAssay Description:Inhibitory constant against rabbit cathepsin K using Z-Phe-Arg-AMC substrateMore data for this Ligand-Target Pair
Affinity DataKi: 0.450nMAssay Description:Inhibitory constant against rabbit cathepsin K using Z-Phe-Arg-AMC substrateMore data for this Ligand-Target Pair
Affinity DataKi: 0.470nMAssay Description:Inhibitory constant against rabbit cathepsin K using Z-Phe-Arg-AMC substrateMore data for this Ligand-Target Pair
Affinity DataKi: 0.480nMAssay Description:Inhibitory constant against rabbit cathepsin K using Z-Phe-Arg-AMC substrateMore data for this Ligand-Target Pair
Affinity DataKi: 0.5nMAssay Description:Inhibitory constant against rabbit cathepsin K using Z-Phe-Arg-AMC substrateMore data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
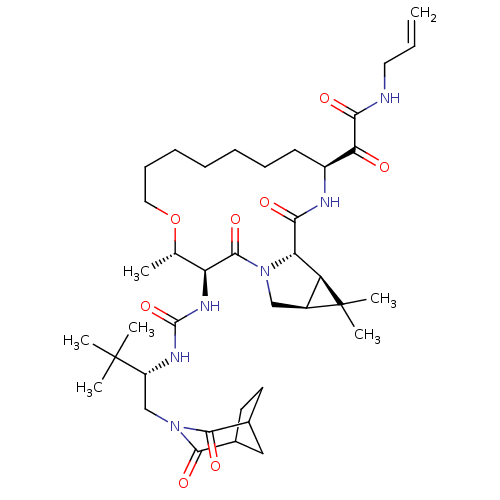
Affinity DataKi: 0.5nM ΔG°: -54.0kJ/molepH: 6.5 T: 2°CAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
Affinity DataKi: 0.570nMAssay Description:Inhibitory constant against rabbit cathepsin K using Z-Phe-Arg-AMC substrateMore data for this Ligand-Target Pair
Affinity DataKi: 0.590nMAssay Description:Inhibitory constant against rabbit cathepsin K using Z-Phe-Arg-AMC substrateMore data for this Ligand-Target Pair
Affinity DataKi: 0.670nMAssay Description:Inhibitory constant against rabbit cathepsin K using Z-Phe-Arg-AMC substrateMore data for this Ligand-Target Pair
Affinity DataKi: 0.790nMAssay Description:Inhibitory constant against rabbit cathepsin K using Z-Phe-Arg-AMC substrateMore data for this Ligand-Target Pair
Affinity DataKi: 0.800nMAssay Description:Inhibition of human cathepsin KMore data for this Ligand-Target Pair
Affinity DataKi: 0.940nMAssay Description:Inhibitory constant against rabbit cathepsin K using Z-Phe-Arg-AMC substrateMore data for this Ligand-Target Pair
Affinity DataKi: 0.950nMAssay Description:Inhibitory constant against rabbit cathepsin K using Z-Phe-Arg-AMC substrateMore data for this Ligand-Target Pair
Affinity DataKi: 1.30nMAssay Description:Inhibition of human cathepsin KMore data for this Ligand-Target Pair
Affinity DataKi: 1.30nMAssay Description:Inhibition of human cathepsin KMore data for this Ligand-Target Pair
Affinity DataKi: 1.40nMAssay Description:Inhibitory constant against rabbit cathepsin K using Z-Phe-Arg-AMC substrateMore data for this Ligand-Target Pair
Affinity DataKi: 1.40nMAssay Description:Inhibitory constant against rabbit cathepsin K using Z-Phe-Arg-AMC substrateMore data for this Ligand-Target Pair
Affinity DataKi: 1.5nM ΔG°: -49.9kJ/mole IC50: 2.10nMT: 2°CAssay Description:Radioligand binding studies were carried out with M3 receptor cell homogenates as described (Peralta et al., The EMBO Journal 6, 3923-3929, (1987)). ...More data for this Ligand-Target Pair
Affinity DataKi: 1.5nMAssay Description:Inhibitory constant against rabbit cathepsin K using Z-Phe-Arg-AMC substrateMore data for this Ligand-Target Pair
Affinity DataKi: 1.60nMAssay Description:Inhibition of human BACE1More data for this Ligand-Target Pair
Affinity DataKi: 1.60nMAssay Description:Inhibitory constant against rabbit cathepsin K using Z-Phe-Arg-AMC substrateMore data for this Ligand-Target Pair
Affinity DataKi: 1.60nMAssay Description:Inhibitory constant against rabbit cathepsin K using Z-Phe-Arg-AMC substrateMore data for this Ligand-Target Pair
Affinity DataKi: 2nMAssay Description:Inhibitory constant against rabbit cathepsin K using Z-Phe-Arg-AMC substrateMore data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
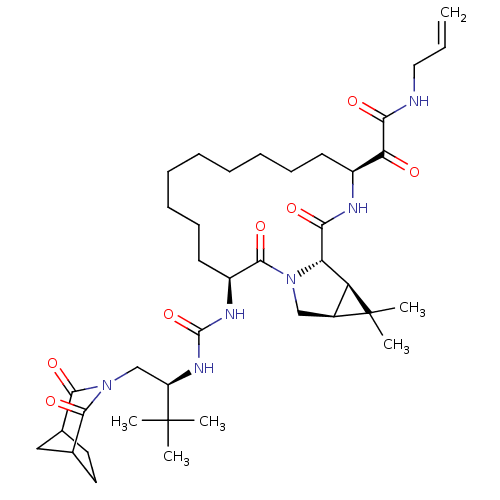
Affinity DataKi: 2nMAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 2nM ΔG°: -50.5kJ/molepH: 6.5 T: 2°CAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 2nM ΔG°: -50.5kJ/molepH: 6.5 T: 2°CAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 2nMAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
Affinity DataKi: 2.10nM ΔG°: -49.0kJ/mole IC50: 2.90nMT: 2°CAssay Description:Radioligand binding studies were carried out with M3 receptor cell homogenates as described (Peralta et al., The EMBO Journal 6, 3923-3929, (1987)). ...More data for this Ligand-Target Pair
Affinity DataKi: 2.30nMAssay Description:Inhibitory constant against rabbit cathepsin K using Z-Phe-Arg-AMC substrateMore data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 3nMAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 3nM ΔG°: -49.5kJ/molepH: 6.5 T: 2°CAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 3nM ΔG°: -49.5kJ/molepH: 6.5 T: 2°CAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 3nM ΔG°: -49.5kJ/molepH: 6.5 T: 2°CAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 3nMAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
Affinity DataKi: 3nMAssay Description:Inhibition of human cathepsin KMore data for this Ligand-Target Pair
Affinity DataKi: 3.30nMAssay Description:Inhibitory constant against rabbit cathepsin K using Z-Phe-Arg-AMC substrateMore data for this Ligand-Target Pair
Affinity DataKi: 3.5nMAssay Description:Inhibition of human cathepsin KMore data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 3.80nM ΔG°: -48.9kJ/molepH: 6.5 T: 2°CAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 4nMAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
Affinity DataKi: 4nMAssay Description:Binding affinity against cathepsin KMore data for this Ligand-Target Pair
Affinity DataKi: 4.80nMAssay Description:Inhibition of human cathepsin KMore data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 5nM ΔG°: -48.2kJ/molepH: 6.5 T: 2°CAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus (HCV genotype 1a, isolate H))
Schering-Plough Research Institute
Schering-Plough Research Institute
Affinity DataKi: 5nM ΔG°: -48.2kJ/molepH: 6.5 T: 2°CAssay Description:Proteolytic cleavage of the ester linkage between the Nva (L-norvaline) and the chromophore (PAP) was monitored for change in absorbance at 370 nm. I...More data for this Ligand-Target Pair

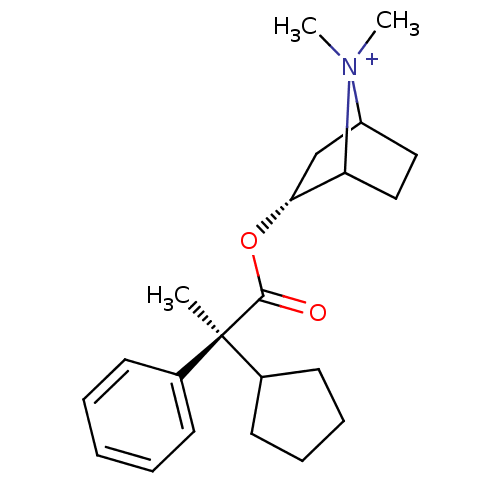
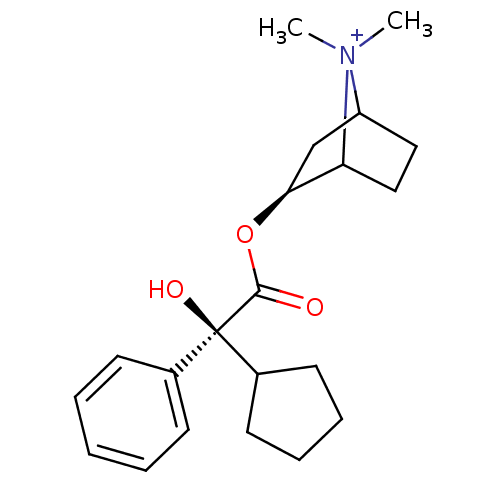
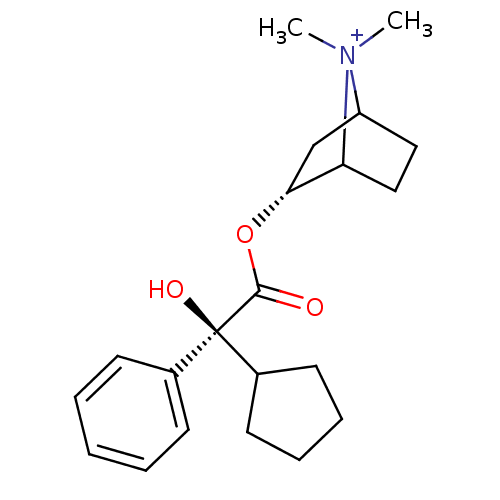
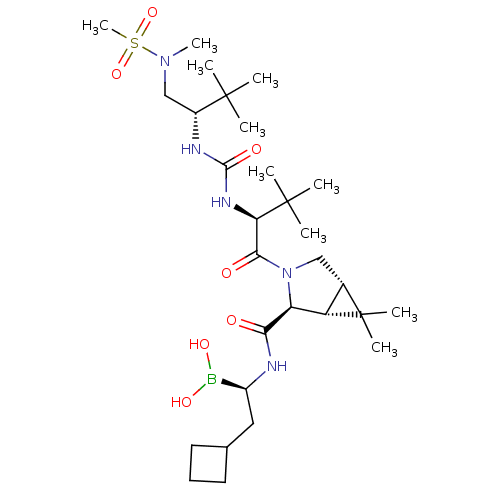
 3D Structure (crystal)
3D Structure (crystal)