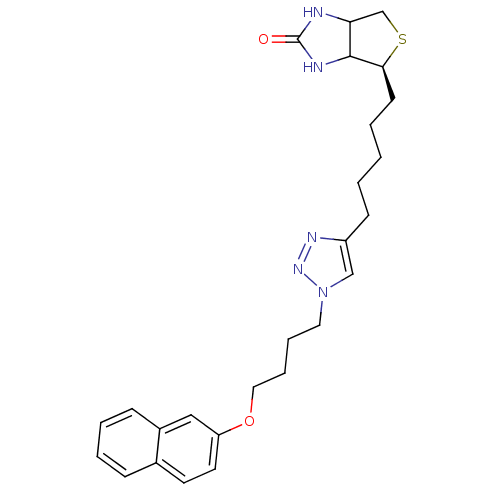
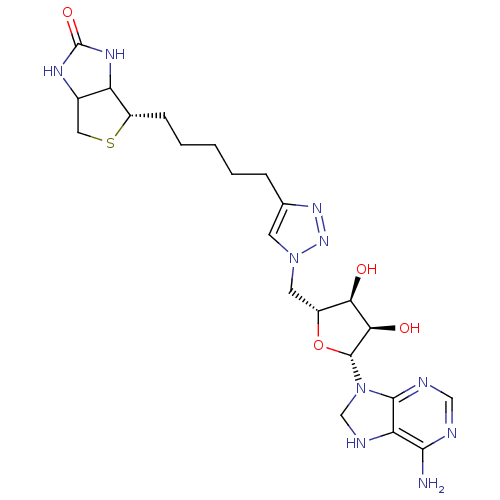
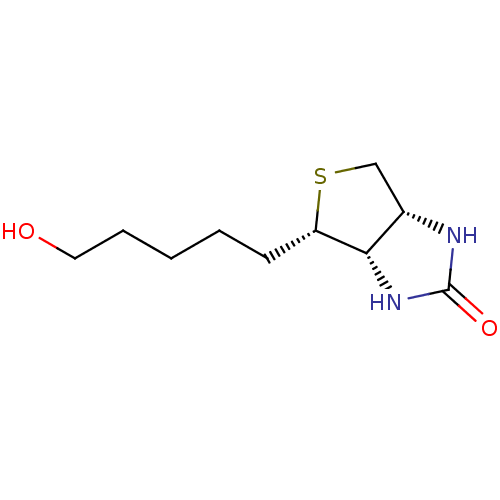
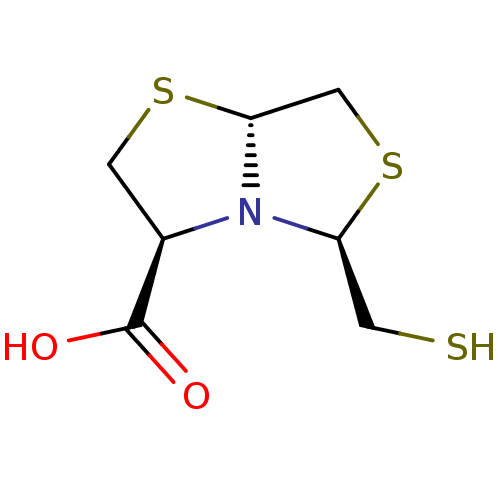
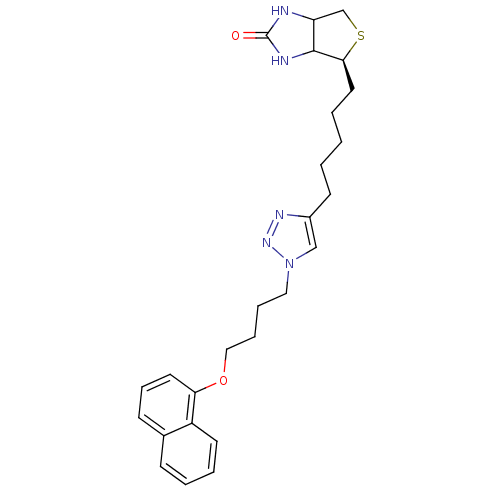
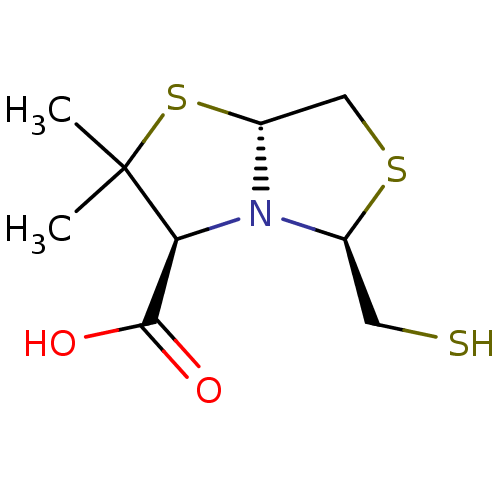
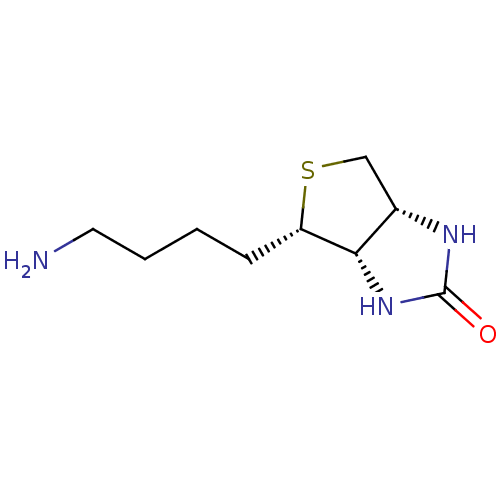
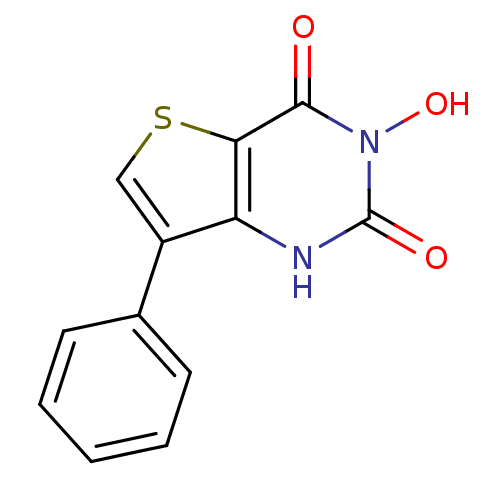
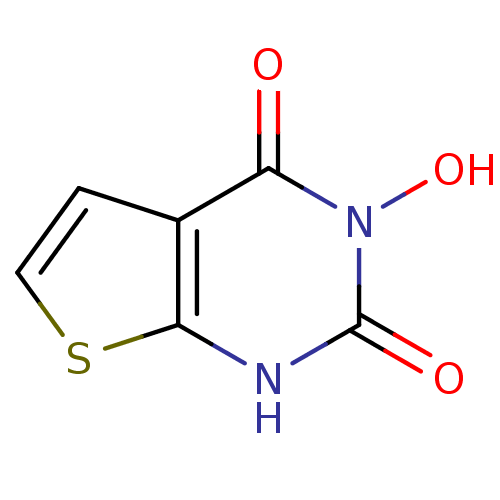
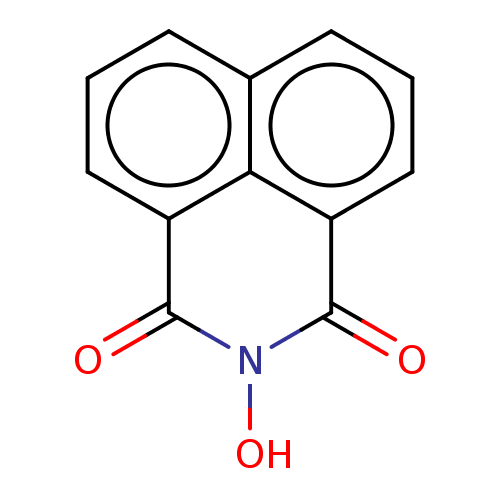
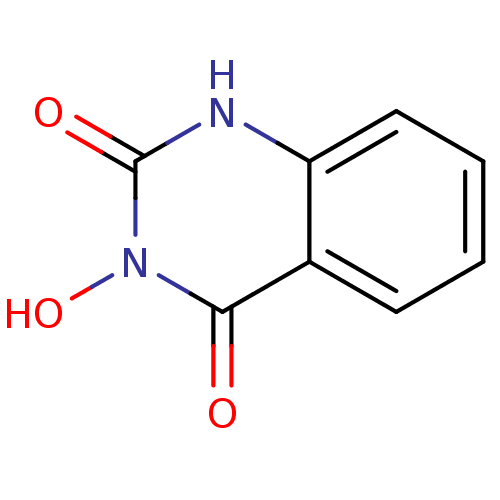
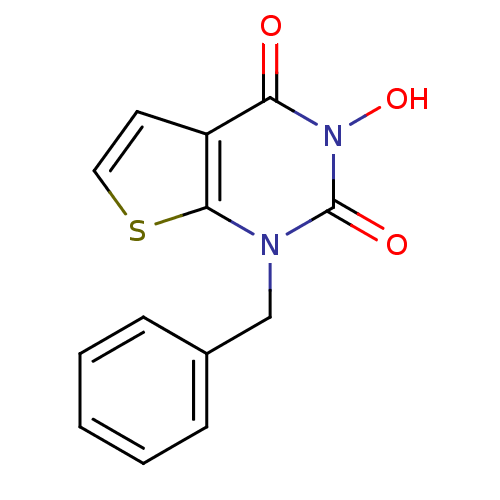
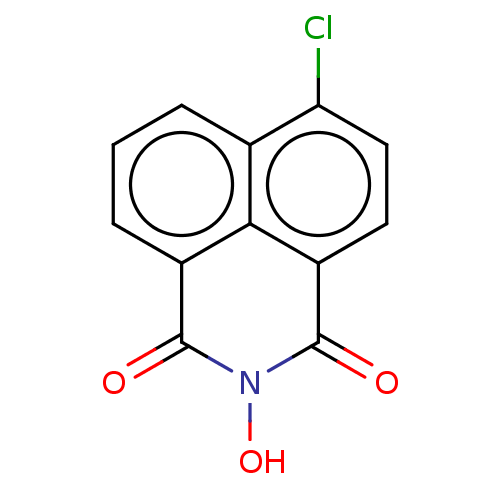
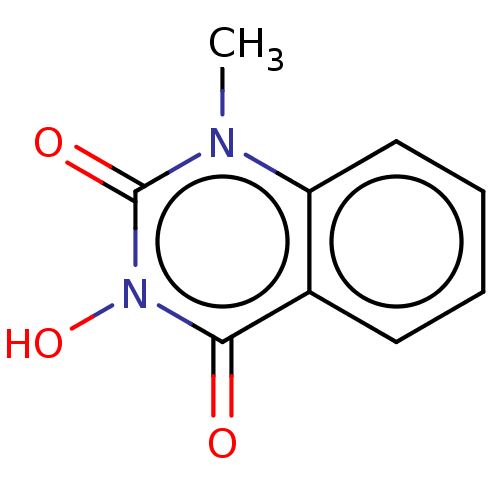
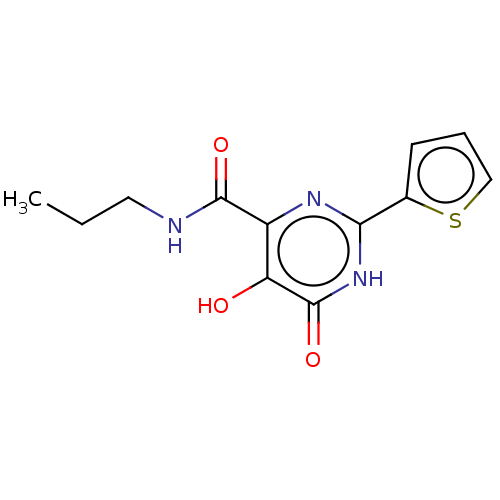
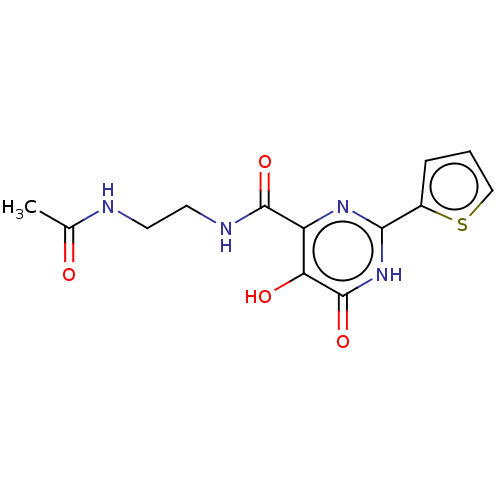
TargetAdenosine receptor A3(Homo sapiens (Human))
University Of California San Francisco
Curated by ChEMBL
University Of California San Francisco
Curated by ChEMBL
Affinity DataKi: 2nMAssay Description:Inhibitory constant against human adenosne A3 receptorMore data for this Ligand-Target Pair
TargetAdenosine receptor A1(Homo sapiens (Human))
University Of California San Francisco
Curated by ChEMBL
University Of California San Francisco
Curated by ChEMBL
Affinity DataKi: 3nMAssay Description:Inhibitory constant aganist human adenosine A1 receptorMore data for this Ligand-Target Pair
Affinity DataKi: 90nMAssay Description:In vitro enzyme inhibition using biotin protein ligase.More data for this Ligand-Target Pair
Affinity DataKi: 140nMAssay Description:Displacement of [3H]-biotin from human BPL after 20 mins by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 200nMAssay Description:Displacement of [3H]-biotin from human BPL after 20 mins by scintillation countingMore data for this Ligand-Target Pair
TargetAdenosine receptor A1(Homo sapiens (Human))
University Of California San Francisco
Curated by ChEMBL
University Of California San Francisco
Curated by ChEMBL
Affinity DataKi: 590nMAssay Description:Inhibitory constant aganist human adenosine A1 receptorMore data for this Ligand-Target Pair
TargetAdenosine receptor A2a(Homo sapiens (Human))
University Of California San Francisco
Curated by ChEMBL
University Of California San Francisco
Curated by ChEMBL
Affinity DataKi: 680nMAssay Description:Inhibitory constant against human adenosine A2a receptorMore data for this Ligand-Target Pair
Affinity DataKi: 1.17E+3nMAssay Description:In vitro enzyme inhibition using biotin protein ligase.More data for this Ligand-Target Pair
Affinity DataKi: 1.17E+3nMAssay Description:In vitro enzyme inhibition using biotin protein ligase.More data for this Ligand-Target Pair
Affinity DataKi: 3.50E+3nMAssay Description:Displacement of [3H]-biotin from human BPL after 20 mins by scintillation countingMore data for this Ligand-Target Pair
TargetMetallo-beta-lactamase VIM-2(Pseudomonas aeruginosa (g-Proteobacteria))
Louis Stokes Cleveland Veterans Affairs Medical Center
Louis Stokes Cleveland Veterans Affairs Medical Center
Affinity DataKi: 3.70E+3nMAssay Description:The Ki for each inhibitor was determined by direct competition assays under steady-state conditions. The initial velocity was measured in the presenc...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase VIM-2(Pseudomonas aeruginosa (g-Proteobacteria))
Louis Stokes Cleveland Veterans Affairs Medical Center
Louis Stokes Cleveland Veterans Affairs Medical Center
Affinity DataKi: 3.80E+3nMAssay Description:The Ki for each inhibitor was determined by direct competition assays under steady-state conditions. The initial velocity was measured in the presenc...More data for this Ligand-Target Pair
Affinity DataKi: 3.85E+3nMAssay Description:Displacement of [3H]-biotin from human BPL after 20 mins by scintillation countingMore data for this Ligand-Target Pair
TargetVIM-24(Klebsiella pneumoniae (Enterobacteria))
Louis Stokes Cleveland Veterans Affairs Medical Center
Louis Stokes Cleveland Veterans Affairs Medical Center
Affinity DataKi: 4.90E+3nMAssay Description:The Ki for each inhibitor was determined by direct competition assays under steady-state conditions. The initial velocity was measured in the presenc...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase VIM-2(Pseudomonas aeruginosa (g-Proteobacteria))
Louis Stokes Cleveland Veterans Affairs Medical Center
Louis Stokes Cleveland Veterans Affairs Medical Center
Affinity DataKi: 5.40E+3nMAssay Description:The Ki for each inhibitor was determined by direct competition assays under steady-state conditions. The initial velocity was measured in the presenc...More data for this Ligand-Target Pair
TargetVIM-24(Klebsiella pneumoniae (Enterobacteria))
Louis Stokes Cleveland Veterans Affairs Medical Center
Louis Stokes Cleveland Veterans Affairs Medical Center
Affinity DataKi: 6.00E+3nMAssay Description:The Ki for each inhibitor was determined by direct competition assays under steady-state conditions. The initial velocity was measured in the presenc...More data for this Ligand-Target Pair
Affinity DataKi: 6.43E+3nMAssay Description:Displacement of [3H]-biotin from human BPL after 20 mins by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: >1.00E+4nMAssay Description:In vitro enzyme inhibition using biotin protein ligase.More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+4nMAssay Description:In vitro enzyme inhibition using biotin protein ligase.More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+4nMAssay Description:In vitro enzyme inhibition using biotin protein ligase.More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+4nMAssay Description:In vitro enzyme inhibition using biotin protein ligase.More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+4nMAssay Description:In vitro enzyme inhibition using biotin protein ligase.More data for this Ligand-Target Pair
TargetVIM-24(Klebsiella pneumoniae (Enterobacteria))
Louis Stokes Cleveland Veterans Affairs Medical Center
Louis Stokes Cleveland Veterans Affairs Medical Center
Affinity DataKi: 1.10E+4nMAssay Description:The Ki for each inhibitor was determined by direct competition assays under steady-state conditions. The initial velocity was measured in the presenc...More data for this Ligand-Target Pair
Affinity DataKi: 1.20E+4nMAssay Description:Displacement of [3H]-biotin from human BPL after 20 mins by scintillation countingMore data for this Ligand-Target Pair
TargetVIM-24(Klebsiella pneumoniae (Enterobacteria))
Louis Stokes Cleveland Veterans Affairs Medical Center
Louis Stokes Cleveland Veterans Affairs Medical Center
Affinity DataKi: 1.20E+4nMAssay Description:The Ki for each inhibitor was determined by direct competition assays under steady-state conditions. The initial velocity was measured in the presenc...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase VIM-2(Pseudomonas aeruginosa (g-Proteobacteria))
Louis Stokes Cleveland Veterans Affairs Medical Center
Louis Stokes Cleveland Veterans Affairs Medical Center
Affinity DataKi: 1.40E+4nMAssay Description:The Ki for each inhibitor was determined by direct competition assays under steady-state conditions. The initial velocity was measured in the presenc...More data for this Ligand-Target Pair
Affinity DataKi: >2.00E+4nMAssay Description:Displacement of [3H]-biotin from human BPL after 20 mins by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: >2.00E+4nMAssay Description:Displacement of [3H]-biotin from human BPL after 20 mins by scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: >2.00E+4nMAssay Description:Displacement of [3H]-biotin from human BPL after 20 mins by scintillation countingMore data for this Ligand-Target Pair
Affinity DataIC50: 3nMAssay Description:Inhibition of FEN1 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 9nMAssay Description:Inhibition of FEN1 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 38nMAssay Description:Inhibition of FEN1 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 57nMAssay Description:Inhibition of FEN1 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 230nMAssay Description:Inhibition of FEN1 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 300nMAssay Description:Inhibition of FEN1 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 300nMAssay Description:Inhibition of FEN1 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 630nMAssay Description:Inhibition of FEN1 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 680nMAssay Description:Inhibition of FEN1 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 700nMAssay Description:Inhibition of FEN1 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 1.10E+3nMAssay Description:Inhibition of FEN1 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 1.30E+3nMAssay Description:Inhibition of FEN1 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+3nMAssay Description:Inhibition of FEN1 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 2.10E+3nMAssay Description:Inhibition of FEN1 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 2.10E+3nMAssay Description:Inhibition of FEN1 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 2.10E+3nMAssay Description:Inhibition of FEN1 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 3.50E+3nMAssay Description:Inhibition of FEN1 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 4.00E+3nMAssay Description:Inhibition of FEN1 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 4.10E+3nMAssay Description:Inhibition of FEN1 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 4.20E+3nMAssay Description:Inhibition of FEN1 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+3nMAssay Description:Inhibition of FEN1 (unknown origin)More data for this Ligand-Target Pair


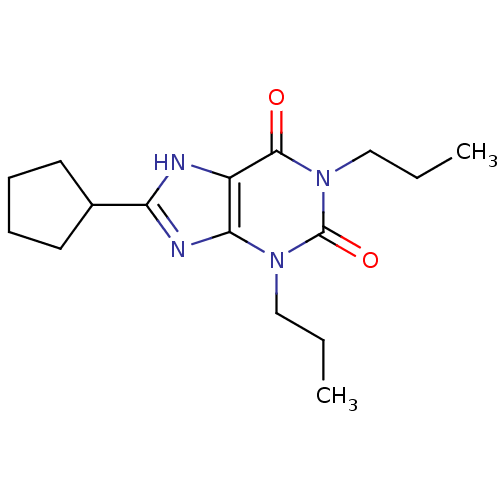





 3D Structure (crystal)
3D Structure (crystal)