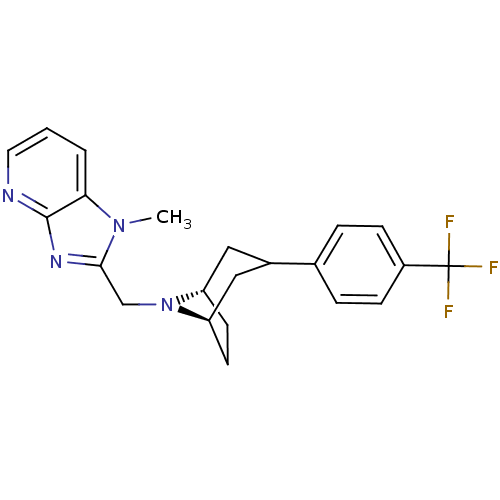
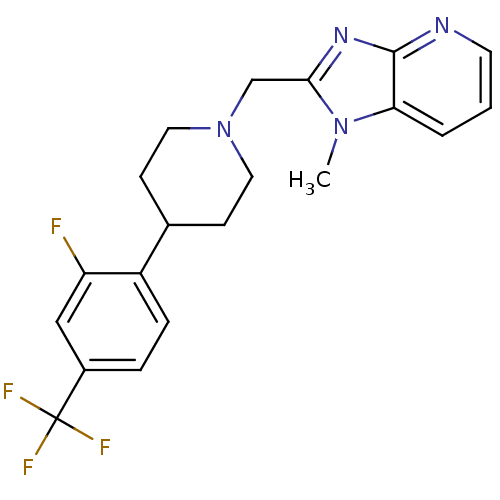
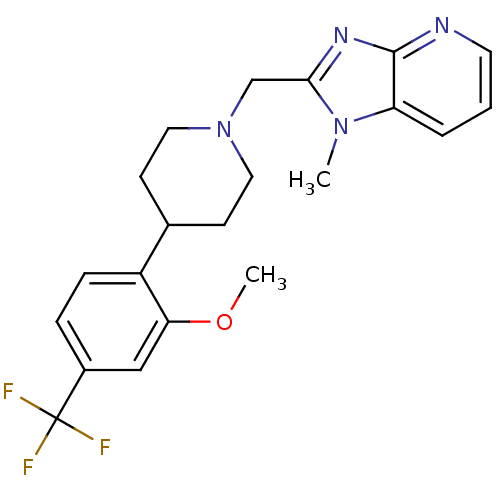
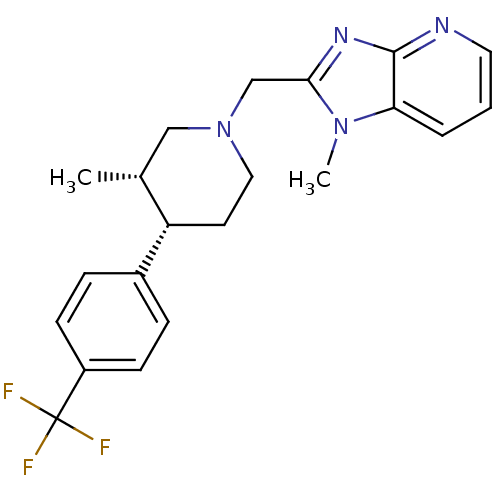
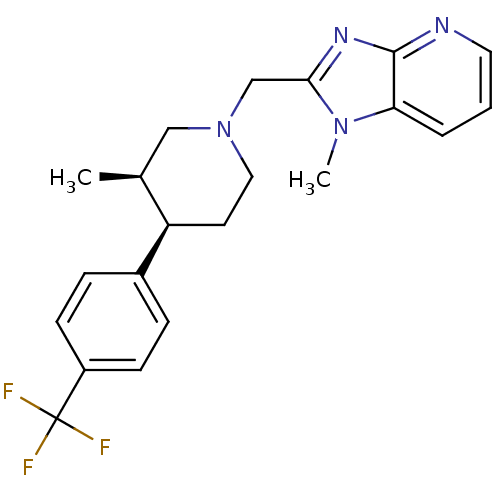
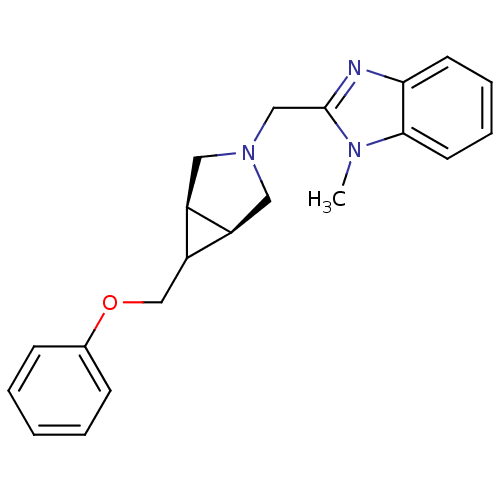
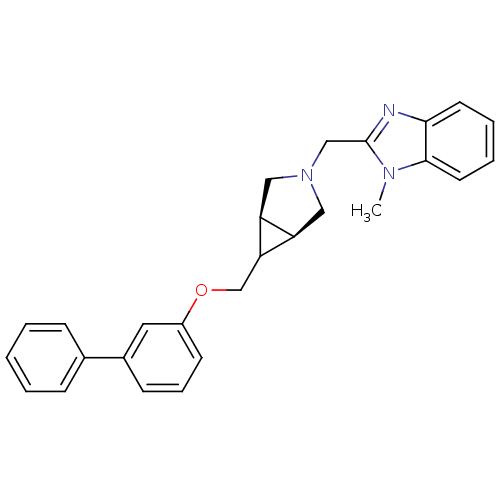
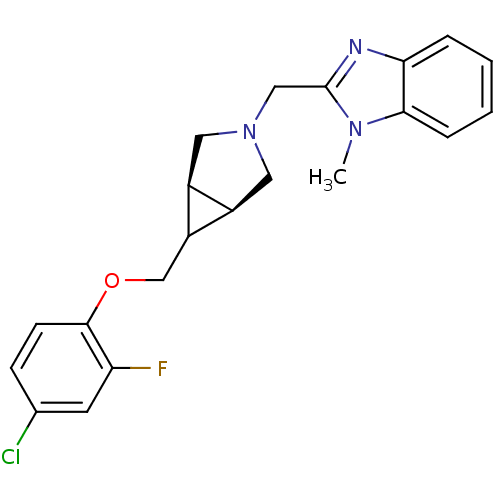
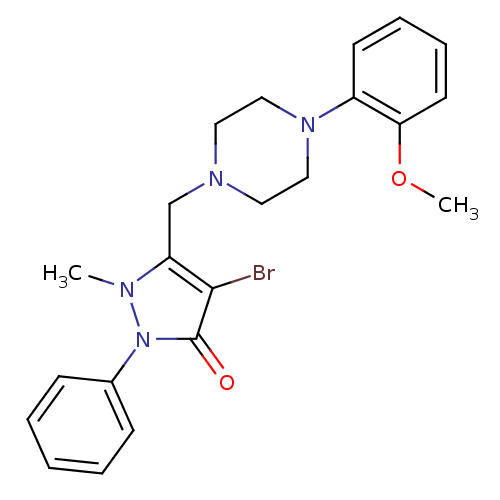
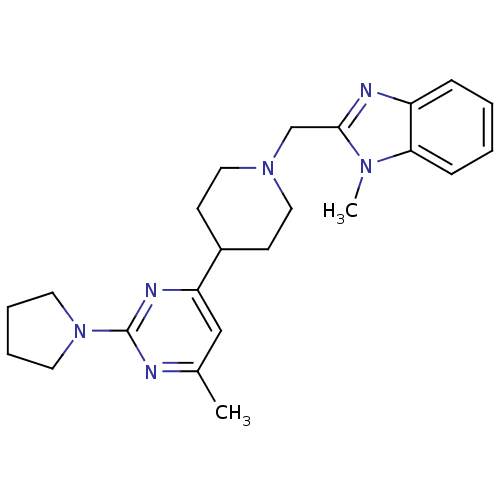
Affinity DataKi: 24nMAssay Description:Displacement of [3H]spiperone from dopamine D2 receptor in rat corpus striatum by beta plate scintillation countingMore data for this Ligand-Target Pair
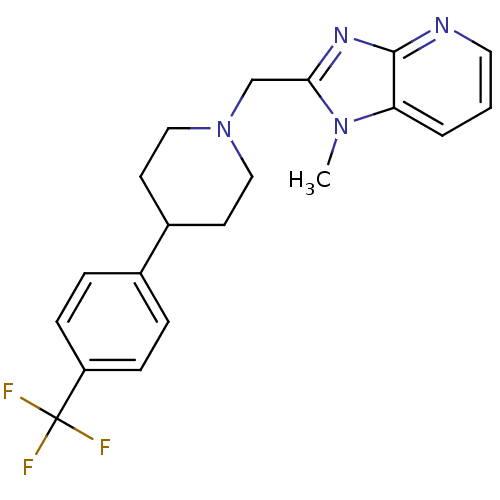
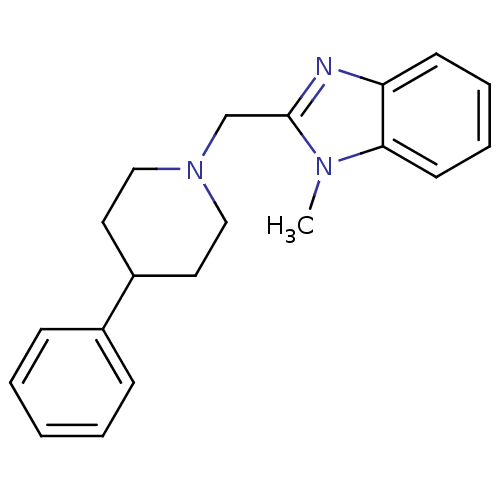
Affinity DataKi: 91nMAssay Description:Displacement of [3H]spiperone from dopamine D2 receptor in rat corpus striatum by beta plate scintillation countingMore data for this Ligand-Target Pair
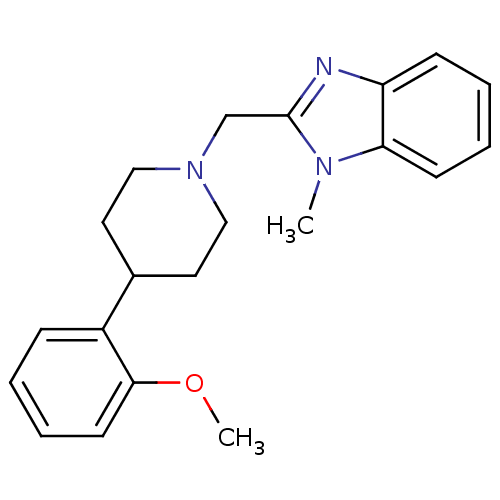
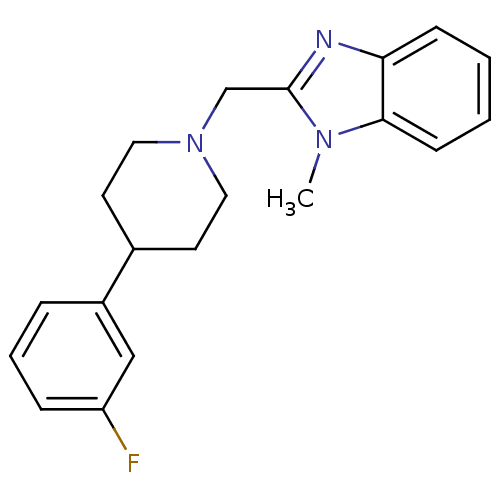
Affinity DataKi: 93nMAssay Description:Displacement of [3H]spiperone from dopamine D2 receptor in rat corpus striatum by beta plate scintillation countingMore data for this Ligand-Target Pair
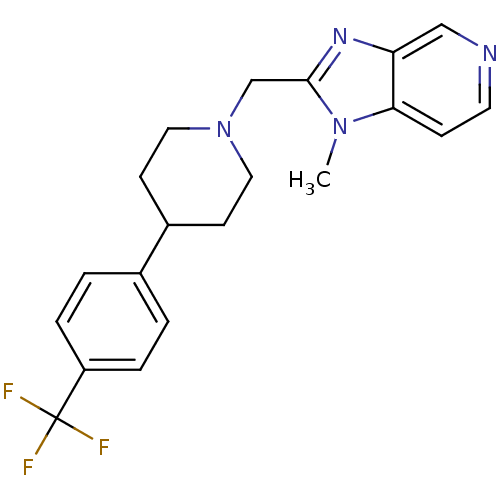
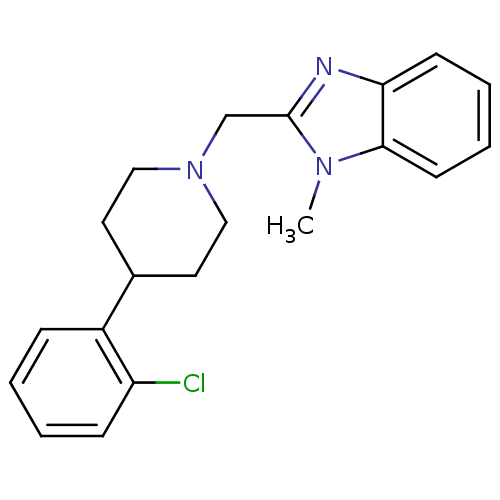
Affinity DataKi: 355nMAssay Description:Displacement of [3H]spiperone from dopamine D2 receptor in rat corpus striatum by beta plate scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 404nMAssay Description:Displacement of [3H]spiperone from dopamine D2 receptor in rat corpus striatum by beta plate scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 426nMAssay Description:Displacement of [3H]spiperone from dopamine D2 receptor in rat corpus striatum by beta plate scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 561nMAssay Description:Displacement of [3H]spiperone from dopamine D2 receptor in rat corpus striatum by beta plate scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 589nMAssay Description:Displacement of [3H]spiperone from dopamine D2 receptor in rat corpus striatum by beta plate scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 671nMAssay Description:Displacement of [3H]spiperone from dopamine D2 receptor in rat corpus striatum by beta plate scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 1.18E+3nMAssay Description:Displacement of [3H]spiperone from dopamine D2 receptor in rat corpus striatum by beta plate scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 1.62E+3nMAssay Description:Displacement of [3H]spiperone from dopamine D2 receptor in rat corpus striatum by beta plate scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 1.78E+3nMAssay Description:Displacement of [3H]spiperone from dopamine D2 receptor in rat corpus striatum by beta plate scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 2.43E+3nMAssay Description:Displacement of [3H]spiperone from dopamine D2 receptor in rat corpus striatum by beta plate scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: >2.80E+3nMAssay Description:Displacement of [3H]spiperone from dopamine D2 receptor in rat corpus striatum by beta plate scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: >2.80E+3nMAssay Description:Displacement of [3H]spiperone from dopamine D2 receptor in rat corpus striatum by beta plate scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: >2.80E+3nMAssay Description:Displacement of [3H]spiperone from dopamine D2 receptor in rat corpus striatum by beta plate scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: >2.80E+3nMAssay Description:Displacement of [3H]spiperone from dopamine D2 receptor in rat corpus striatum by beta plate scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: >4.68E+3nMAssay Description:Displacement of [3H]spiperone from dopamine D2 receptor in rat corpus striatum by beta plate scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: >5.26E+3nMAssay Description:Displacement of [3H]spiperone from dopamine D2 receptor in rat corpus striatum by beta plate scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: >5.26E+3nMAssay Description:Displacement of [3H]spiperone from dopamine D2 receptor in rat corpus striatum by beta plate scintillation countingMore data for this Ligand-Target Pair
Affinity DataEC50: 511nMAssay Description:Activity at rat mGluR2 receptor expressed in HEK cells by FLIPR assay in presence of glutamateMore data for this Ligand-Target Pair
Affinity DataEC50: 30nMAssay Description:Activity at rat mGluR2 receptor expressed in HEK cells by FLIPR assay in presence of glutamateMore data for this Ligand-Target Pair
Affinity DataEC50: 20nMAssay Description:Activity at rat mGluR2 receptor expressed in HEK cells by FLIPR assay in presence of glutamateMore data for this Ligand-Target Pair
Affinity DataEC50: 251nMAssay Description:Activity at rat mGluR2 receptor expressed in HEK cells by FLIPR assay in presence of glutamateMore data for this Ligand-Target Pair
Affinity DataEC50: 2.12E+3nMAssay Description:Activity at rat mGluR2 receptor expressed in HEK cells by FLIPR assay in presence of glutamateMore data for this Ligand-Target Pair
Affinity DataEC50: 419nMAssay Description:Activity at rat mGluR2 receptor expressed in HEK cells by FLIPR assay in presence of glutamateMore data for this Ligand-Target Pair
Affinity DataEC50: 64nMAssay Description:Activity at rat mGluR2 receptor expressed in HEK cells by FLIPR assay in presence of glutamateMore data for this Ligand-Target Pair
Affinity DataEC50: 86nMAssay Description:Activity at rat mGluR2 receptor expressed in HEK cells by FLIPR assay in presence of glutamateMore data for this Ligand-Target Pair
Affinity DataEC50: 24nMAssay Description:Activity at rat mGluR2 receptor expressed in HEK cells by FLIPR assay in presence of glutamateMore data for this Ligand-Target Pair
Affinity DataEC50: >1.00E+4nMAssay Description:Activity at rat mGluR2 receptor expressed in HEK cells by FLIPR assay in presence of glutamateMore data for this Ligand-Target Pair
Affinity DataEC50: >1.00E+4nMAssay Description:Activity at rat mGluR2 receptor expressed in HEK cells by FLIPR assay in presence of glutamateMore data for this Ligand-Target Pair
Affinity DataEC50: 9.75E+3nMAssay Description:Activity at rat mGluR2 receptor expressed in HEK cells by FLIPR assay in presence of glutamateMore data for this Ligand-Target Pair
Affinity DataEC50: 3.02E+3nMAssay Description:Activity at rat mGluR2 receptor expressed in HEK cells by FLIPR assay in presence of glutamateMore data for this Ligand-Target Pair
Affinity DataEC50: 967nMAssay Description:Activity at rat mGluR2 receptor expressed in HEK cells by FLIPR assay in presence of glutamateMore data for this Ligand-Target Pair
Affinity DataEC50: 113nMAssay Description:Activity at rat mGluR2 receptor expressed in HEK cells by FLIPR assay in presence of glutamateMore data for this Ligand-Target Pair
Affinity DataEC50: 7nMAssay Description:Activity at rat mGluR2 receptor expressed in HEK cells by FLIPR assay in presence of glutamateMore data for this Ligand-Target Pair
Affinity DataEC50: 26nMAssay Description:Activity at rat mGluR2 receptor expressed in HEK cells by FLIPR assay in presence of glutamateMore data for this Ligand-Target Pair
Affinity DataEC50: 26nMAssay Description:Activity at rat mGluR2 receptor expressed in HEK cells by FLIPR assay in presence of glutamateMore data for this Ligand-Target Pair
Affinity DataEC50: 8nMAssay Description:Activity at rat mGluR2 receptor expressed in HEK cells by FLIPR assay in presence of glutamateMore data for this Ligand-Target Pair
Affinity DataEC50: 85nMAssay Description:Activity at rat mGluR2 receptor expressed in HEK cells by FLIPR assay in presence of glutamateMore data for this Ligand-Target Pair
Affinity DataEC50: 26nMAssay Description:Activity at rat mGluR2 receptor expressed in HEK cells by FLIPR assay in presence of glutamateMore data for this Ligand-Target Pair
Affinity DataEC50: 951nMAssay Description:Antagonist activity at human dopamine D2 receptor expressed in CHO cells assessed as inhibition of quinpirole-induced effectMore data for this Ligand-Target Pair
Affinity DataEC50: 891nMAssay Description:Positive allosteric modulation of rat mGluR2 expressed in HEK293 cells co-expressing Galpha15 G protein assessed as increase in glutamate-induced int...More data for this Ligand-Target Pair
Affinity DataEC50: 4.79E+3nMAssay Description:Positive allosteric modulation of rat mGluR2 expressed in HEK293 cells co-expressing Galpha15 G protein assessed as increase in glutamate-induced int...More data for this Ligand-Target Pair
Affinity DataEC50: 87nMAssay Description:Positive allosteric modulation of rat mGluR2 expressed in HEK293 cells co-expressing Galpha15 G protein assessed as increase in glutamate-induced int...More data for this Ligand-Target Pair
Affinity DataEC50: 212nMAssay Description:Positive allosteric modulation of rat mGluR2 expressed in HEK293 cells co-expressing Galpha15 G protein assessed as increase in glutamate-induced int...More data for this Ligand-Target Pair
Affinity DataEC50: 22nMAssay Description:Positive allosteric modulation of rat mGluR2 expressed in HEK293 cells co-expressing Galpha15 G protein assessed as increase in glutamate-induced int...More data for this Ligand-Target Pair
Affinity DataEC50: 3.31E+3nMAssay Description:Positive allosteric modulation of rat mGluR2 expressed in HEK293 cells co-expressing Galpha15 G protein assessed as increase in glutamate-induced int...More data for this Ligand-Target Pair
Affinity DataEC50: 159nMAssay Description:Positive allosteric modulation of rat mGluR2 expressed in HEK293 cells co-expressing Galpha15 G protein assessed as increase in glutamate-induced int...More data for this Ligand-Target Pair
Affinity DataEC50: 144nMAssay Description:Positive allosteric modulation of rat mGluR2 expressed in HEK293 cells co-expressing Galpha15 G protein assessed as increase in glutamate-induced int...More data for this Ligand-Target Pair