Reaction Details Report a problem with these data
Report a problem with these data
 Report a problem with these data
Report a problem with these dataTarget
G protein-activated inward rectifier potassium channel 1/4
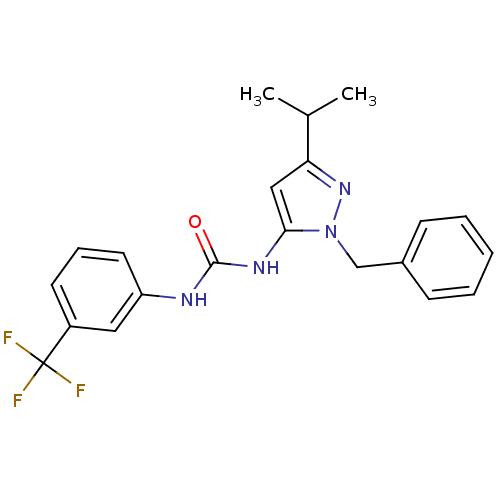
Ligand
BDBM50437968
Substrate
n/a
Meas. Tech.
ChEMBL_972874 (CHEMBL2411651)
EC50
3500±n/a nM
Citation
 Wen, W; Wu, W; Romaine, IM; Kaufmann, K; Du, Y; Sulikowski, GA; Weaver, CD; Lindsley, CW Discovery of 'molecular switches' within a GIRK activator scaffold that afford selective GIRK inhibitors. Bioorg Med Chem Lett 23:4562-6 (2013) [PubMed] Article
Wen, W; Wu, W; Romaine, IM; Kaufmann, K; Du, Y; Sulikowski, GA; Weaver, CD; Lindsley, CW Discovery of 'molecular switches' within a GIRK activator scaffold that afford selective GIRK inhibitors. Bioorg Med Chem Lett 23:4562-6 (2013) [PubMed] Article More Info.:
Target
Name:
G protein-activated inward rectifier potassium channel 1/4
Synonyms:
GIRK1/4 channels | Kir3.1/Kir3.4
Type:
PROTEIN
Mol. Mass.:
n/a
Description:
n/a
Components:
This complex has 2 components.
Component 1
Name:
G protein-activated inward rectifier potassium channel 1
Synonyms:
GIRK-1 | GIRK1 | Inward rectifier K(+) channel Kir3.1 | KCNJ3 | KCNJ3_HUMAN | Potassium channel, inwardly rectifying subfamily J member 3
Type:
PROTEIN
Mol. Mass.:
56610.74
Organism:
Homo sapiens (Human)
Description:
ChEMBL_107915
Residue:
501
Sequence:
MSALRRKFGDDYQVVTTSSSGSGLQPQGPGQDPQQQLVPKKKRQRFVDKNGRCNVQHGNLGSETSRYLSDLFTTLVDLKWRWNLFIFILTYTVAWLFMASMWWVIAYTRGDLNKAHVGNYTPCVANVYNFPSAFLFFIETEATIGYGYRYITDKCPEGIILFLFQSILGSIVDAFLIGCMFIKMSQPKKRAETLMFSEHAVISMRDGKLTLMFRVGNLRNSHMVSAQIRCKLLKSRQTPEGEFLPLDQLELDVGFSTGADQLFLVSPLTICHVIDAKSPFYDLSQRSMQTEQFEIVVILEGIVETTGMTCQARTSYTEDEVLWGHRFFPVISLEEGFFKVDYSQFHATFEVPTPPYSVKEQEEMLLMSSPLIAPAITNSKERHNSVECLDGLDDITTKLPSKLQKITGREDFPKKLLRMSSTTSEKAYSLGDLPMKLQRISSVPGNSEEKLVSKTTKMLSDPMSQSVADLPPKLQKMAGGAARMEGNLPAKLRKMNSDRFT
Component 2
Name:
G protein-activated inward rectifier potassium channel 4
Synonyms:
CIR | Cardiac inward rectifier | GIRK-4 | GIRK4 | Heart KATP channel | IRK-4 | Inward rectifier K(+) channel Kir3.4 | KATP-1 | KCNJ5 | KCNJ5_HUMAN | Potassium channel, inwardly rectifying subfamily J member 5
Type:
PROTEIN
Mol. Mass.:
47658.06
Organism:
Homo sapiens (Human)
Description:
ChEMBL_107914
Residue:
419
Sequence:
MAGDSRNAMNQDMEIGVTPWDPKKIPKQARDYVPIATDRTRLLAEGKKPRQRYMEKSGKCNVHHGNVQETYRYLSDLFTTLVDLKWRFNLLVFTMVYTVTWLFFGFIWWLIAYIRGDLDHVGDQEWIPCVENLSGFVSAFLFSIETETTIGYGFRVITEKCPEGIILLLVQAILGSIVNAFMVGCMFVKISQPKKRAETLMFSNNAVISMRDEKLCLMFRVGDLRNSHIVEASIRAKLIKSRQTKEGEFIPLNQTDINVGFDTGDDRLFLVSPLIISHEINQKSPFWEMSQAQLHQEEFEVVVILEGMVEATGMTCQARSSYMDTEVLWGHRFTPVLTLEKGFYEVDYNTFHDTYETNTPSCCAKELAEMKREGRLLQYLPSPPLLGGCAEAGLDAEAEQNEEDEPKGLGGSREARGSV