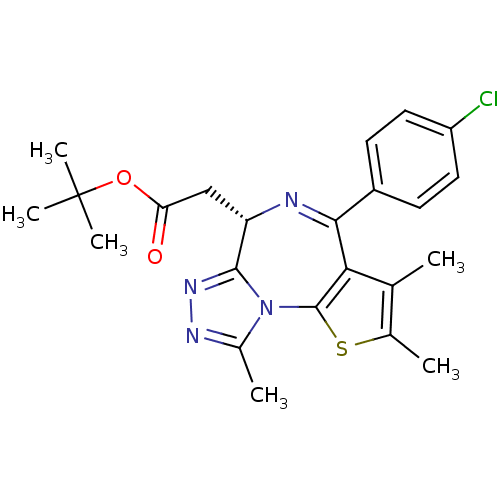
BDBM50365262 (+)-JQ1::(S)-JQ1 (1)::CHEMBL1957266::JQ1::US10124009, Compound (S)-JQ1::US10202360, Example JQ-1::US10308662, Compound JQ-1::US10407441, Compound (S)-JQ1::US10617680, Example JQ-1::US10881668, Compound JQ1::US10925881, Name (S)-JQ1::US11020380, Example JQ-1::US11078188, Example (+)-JQ1::US11279703, TABLE 6.180::US11427593, Compound JQ1::US11466034, Example (+)-JQ1::US9320741, (S)-JQ1::US9695172, JQ1
SMILES Cc1nnc2[C@H](CC(=O)OC(C)(C)C)N=C(c3c(C)c(C)sc3-n12)c1ccc(Cl)cc1
InChI Key InChIKey=DNVXATUJJDPFDM-KRWDZBQOSA-N
Activity Spreadsheet -- Enzyme Inhibition Constant Data from BindingDB
 Found 300 hits for monomerid = 50365262
Found 300 hits for monomerid = 50365262
Affinity DataKi: 5.30nMAssay Description:Binding affinity to human BRD4 BD1 (44 to 168 residues) by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetBromodomain-containing protein 3(Homo sapiens (Human))
University Of Michigan Comprehensive Cancer Center
Curated by ChEMBL
University Of Michigan Comprehensive Cancer Center
Curated by ChEMBL
Affinity DataKi: 6.60nMAssay Description:Inhibition of FAM-labeled ZBA248 binding to recombinant N-terminal His6-tagged BRD3 bromodomain 1 (24 to 144 residues) (unknown origin) expressed in ...More data for this Ligand-Target Pair
TargetBromodomain-containing protein 3(Homo sapiens (Human))
University Of Michigan Comprehensive Cancer Center
Curated by ChEMBL
University Of Michigan Comprehensive Cancer Center
Curated by ChEMBL
Affinity DataKi: 6.60nMAssay Description:Displacement of FAM-labeled ZBA248 from BRD3 BD1 (24 to 144 amino acid residues) (unknown origin) expressed in Rosetta2 DE3 cells after 30 mins by fl...More data for this Ligand-Target Pair
Affinity DataKi: 7.60nMAssay Description:Displacement of FAM-labeled ZBA248 from BRD4 BD1 (44 to 168 amino acid residues) (unknown origin) expressed in Rosetta2 DE3 cells after 30 mins by fl...More data for this Ligand-Target Pair
TargetBromodomain-containing protein 3(Homo sapiens (Human))
University Of Michigan Comprehensive Cancer Center
Curated by ChEMBL
University Of Michigan Comprehensive Cancer Center
Curated by ChEMBL
Affinity DataKi: 8.90nMAssay Description:Inhibition of FAM-labeled ZBA248 binding to recombinant N-terminal His6-tagged BRD3 bromodomain 2 (306 to 417residues) (unknown origin) expressed in ...More data for this Ligand-Target Pair
TargetBromodomain-containing protein 3(Homo sapiens (Human))
University Of Michigan Comprehensive Cancer Center
Curated by ChEMBL
University Of Michigan Comprehensive Cancer Center
Curated by ChEMBL
Affinity DataKi: 8.90nMAssay Description:Displacement of FAM-labeled ZBA248 from BRD3 BD2 (306 to 417 amino acid residues) (unknown origin) expressed in Rosetta2 DE3 cells after 30 mins by f...More data for this Ligand-Target Pair
Affinity DataKi: 11nMAssay Description:Displacement of FAM-labeled ZBA248 from BRD4 BD2 (333 to 460 amino acid residues) (unknown origin) expressed in Rosetta2 DE3 cells after 30 mins by f...More data for this Ligand-Target Pair
Affinity DataKi: 12nMAssay Description:Inhibition of FAM-labeled ZBA248 binding to recombinant N-terminal His6-tagged BRD4 bromodomain 2 (333 to 460 residues) (unknown origin) expressed in...More data for this Ligand-Target Pair
Affinity DataKi: 12nMAssay Description:Inhibition of FAM-labeled ZBA248 binding to recombinant human N-terminal His6-tagged BRD4 bromodomain 2 (333 to 460 residues) expressed in Rosetta2 D...More data for this Ligand-Target Pair
TargetBromodomain-containing protein 2(Homo sapiens (Human))
University Of Michigan Comprehensive Cancer Center
Curated by ChEMBL
University Of Michigan Comprehensive Cancer Center
Curated by ChEMBL
Affinity DataKi: 13nMAssay Description:Inhibition of FAM-labeled ZBA248 binding to recombinant N-terminal His6-tagged BRD2 bromodomain 2 (349 to 460 residues) (unknown origin) expressed in...More data for this Ligand-Target Pair
TargetBromodomain-containing protein 2(Homo sapiens (Human))
University Of Michigan Comprehensive Cancer Center
Curated by ChEMBL
University Of Michigan Comprehensive Cancer Center
Curated by ChEMBL
Affinity DataKi: 13nMAssay Description:Inhibition of FAM-labeled ZBA248 binding to recombinant N-terminal His6-tagged BRD2 bromodomain 1 (72 to 205 residues) (unknown origin) expressed in ...More data for this Ligand-Target Pair
TargetBromodomain-containing protein 2(Homo sapiens (Human))
University Of Michigan Comprehensive Cancer Center
Curated by ChEMBL
University Of Michigan Comprehensive Cancer Center
Curated by ChEMBL
Affinity DataKi: 13nMAssay Description:Displacement of FAM-labeled ZBA248 from BRD2 BD1 (72 to 205 amino acid residues) (unknown origin) expressed in Rosetta2 DE3 cells after 30 mins by fl...More data for this Ligand-Target Pair
TargetBromodomain-containing protein 2(Homo sapiens (Human))
University Of Michigan Comprehensive Cancer Center
Curated by ChEMBL
University Of Michigan Comprehensive Cancer Center
Curated by ChEMBL
Affinity DataKi: 13nMAssay Description:Displacement of FAM-labeled ZBA248 from BRD2 BD2 (349 to 460 amino acid residues) (unknown origin) expressed in Rosetta2 DE3 cells after 30 mins by f...More data for this Ligand-Target Pair
Affinity DataKi: 15nMAssay Description:Inhibition of FAM-labeled ZBA248 binding to recombinant human N-terminal His6-tagged BRD4 bromodomain 1 (44 to 168 residues) expressed in Rosetta2 DE...More data for this Ligand-Target Pair
Affinity DataKi: 15nMAssay Description:Inhibition of FAM-labeled ZBA248 binding to recombinant N-terminal His6-tagged BRD4 bromodomain 1 (44 to 168 residues) (unknown origin) expressed in ...More data for this Ligand-Target Pair
Affinity DataKi: 77nMAssay Description:The compound was tested for the inhibition of fibrinogen receptorMore data for this Ligand-Target Pair
Affinity DataIC50: 77nMAssay Description:Binding affinity to first bromodomain of BRD4More data for this Ligand-Target Pair
Affinity DataKd: 39.5nMpH: 7.4Assay Description:ITC experiments were carried out on anAuto-ITC200 titration microcalorimeter (Malvern Instruments). All experimentswere carried out at 25 �C while st...More data for this Ligand-Target Pair
Affinity DataIC50: 18nMpH: 7.4 T: 2°CAssay Description:BRD4(1): Assays were performed as described previously with minor modifications from the manufacturer's protocol (PerkinElmer, USA). All reagents wer...More data for this Ligand-Target Pair
Affinity DataIC50: 14nMpH: 7.4 T: 2°CAssay Description:BRD4(2): Assays were performed as described previously with minor modifications from the manufacturer's protocol (PerkinElmer, USA). All reagents wer...More data for this Ligand-Target Pair
Affinity DataKd: 14.5nMpH: 7.5 T: 2°CAssay Description:For KD determinations, 10 nM JQ1-TAMRA probe was incubated in 384-well smallvolume black microtiter plates with serial (1:2) dilutions of each BRD4 v...More data for this Ligand-Target Pair
Affinity DataKd: 835nMpH: 7.5 T: 2°CAssay Description:For KD determinations, 10 nM JQ1-TAMRA probe was incubated in 384-well smallvolume black microtiter plates with serial (1:2) dilutions of each BRD4 v...More data for this Ligand-Target Pair
Affinity DataKd: 229nMpH: 7.5 T: 2°CAssay Description:For KD determinations, 10 nM JQ1-TAMRA probe was incubated in 384-well smallvolume black microtiter plates with serial (1:2) dilutions of each BRD4 v...More data for this Ligand-Target Pair
Affinity DataKd: 85.3nMpH: 7.5 T: 2°CAssay Description:For KD determinations, 10 nM JQ1-TAMRA probe was incubated in 384-well smallvolume black microtiter plates with serial (1:2) dilutions of each BRD4 v...More data for this Ligand-Target Pair
Affinity DataKd: 475nMpH: 7.5 T: 2°CAssay Description:For KD determinations, 10 nM JQ1-TAMRA probe was incubated in 384-well smallvolume black microtiter plates with serial (1:2) dilutions of each BRD4 v...More data for this Ligand-Target Pair
Affinity DataKd: 11.4nMpH: 7.5 T: 2°CAssay Description:For KD determinations, 10 nM JQ1-TAMRA probe was incubated in 384-well smallvolume black microtiter plates with serial (1:2) dilutions of each BRD4 v...More data for this Ligand-Target Pair
Affinity DataKd: 448nMpH: 7.5 T: 2°CAssay Description:For KD determinations, 10 nM JQ1-TAMRA probe was incubated in 384-well smallvolume black microtiter plates with serial (1:2) dilutions of each BRD4 v...More data for this Ligand-Target Pair
Affinity DataKd: 56.6nMpH: 7.5 T: 2°CAssay Description:For KD determinations, 10 nM JQ1-TAMRA probe was incubated in 384-well smallvolume black microtiter plates with serial (1:2) dilutions of each BRD4 v...More data for this Ligand-Target Pair
Affinity DataKd: 15.8nMpH: 7.5 T: 2°CAssay Description:For KD determinations, 10 nM JQ1-TAMRA probe was incubated in 384-well smallvolume black microtiter plates with serial (1:2) dilutions of each BRD4 v...More data for this Ligand-Target Pair
Affinity DataKd: 415nMpH: 7.5 T: 2°CAssay Description:For KD determinations, 10 nM JQ1-TAMRA probe was incubated in 384-well smallvolume black microtiter plates with serial (1:2) dilutions of each BRD4 v...More data for this Ligand-Target Pair
Affinity DataKd: 268nMpH: 7.5 T: 2°CAssay Description:For KD determinations, 10 nM JQ1-TAMRA probe was incubated in 384-well smallvolume black microtiter plates with serial (1:2) dilutions of each BRD4 v...More data for this Ligand-Target Pair
Affinity DataKd: 1nMpH: 7.5Assay Description:In a typical TSA, 0.4 μg of BRD4 BD1 protein were mixed with 5� Sypro Orange (Molecular Probe) and adjusted to a final volume of 5 μl with ...More data for this Ligand-Target Pair
Affinity DataKd: 13.7nMpH: 7.5Assay Description:In a typical TSA, 0.4 μg of BRD4 BD1 protein were mixed with 5� Sypro Orange (Molecular Probe) and adjusted to a final volume of 5 μl with ...More data for this Ligand-Target Pair
Affinity DataKd: 3.64nMpH: 7.5Assay Description:In a typical TSA, 0.4 μg of BRD4 BD1 protein were mixed with 5� Sypro Orange (Molecular Probe) and adjusted to a final volume of 5 μl with ...More data for this Ligand-Target Pair
Affinity DataKd: 2.14nMpH: 7.5Assay Description:In a typical TSA, 0.4 μg of BRD4 BD1 protein were mixed with 5� Sypro Orange (Molecular Probe) and adjusted to a final volume of 5 μl with ...More data for this Ligand-Target Pair
Affinity DataKd: 10.5nMpH: 7.5Assay Description:In a typical TSA, 0.4 μg of BRD4 BD1 protein were mixed with 5� Sypro Orange (Molecular Probe) and adjusted to a final volume of 5 μl with ...More data for this Ligand-Target Pair
Affinity DataKd: 1nMpH: 7.5Assay Description:In a typical TSA, 0.4 μg of BRD4 BD1 protein were mixed with 5� Sypro Orange (Molecular Probe) and adjusted to a final volume of 5 μl with ...More data for this Ligand-Target Pair
Affinity DataKd: 8.60nMpH: 7.5Assay Description:In a typical TSA, 0.4 μg of BRD4 BD1 protein were mixed with 5� Sypro Orange (Molecular Probe) and adjusted to a final volume of 5 μl with ...More data for this Ligand-Target Pair
Affinity DataKd: 1nMpH: 7.5Assay Description:In a typical TSA, 0.4 μg of BRD4 BD1 protein were mixed with 5� Sypro Orange (Molecular Probe) and adjusted to a final volume of 5 μl with ...More data for this Ligand-Target Pair
Affinity DataKd: 1.10nMpH: 7.5Assay Description:In a typical TSA, 0.4 μg of BRD4 BD1 protein were mixed with 5� Sypro Orange (Molecular Probe) and adjusted to a final volume of 5 μl with ...More data for this Ligand-Target Pair
Affinity DataKd: 2.40nMpH: 7.5Assay Description:In a typical TSA, 0.4 μg of BRD4 BD1 protein were mixed with 5� Sypro Orange (Molecular Probe) and adjusted to a final volume of 5 μl with ...More data for this Ligand-Target Pair
Affinity DataKd: 9.70nMpH: 7.5Assay Description:In a typical TSA, 0.4 μg of BRD4 BD1 protein were mixed with 5� Sypro Orange (Molecular Probe) and adjusted to a final volume of 5 μl with ...More data for this Ligand-Target Pair
Affinity DataKd: 106nMpH: 7.5 T: 2°CAssay Description:The experiments were performed in 384-well black small volume microtiter plates(Greiner) in a buffer composed of 50mM Hepes, pH 7.5 (Applichem), 50 m...More data for this Ligand-Target Pair
Affinity DataKd: 174nMpH: 7.5 T: 2°CAssay Description:The experiments were performed in 384-well black small volume microtiter plates(Greiner) in a buffer composed of 50mM Hepes, pH 7.5 (Applichem), 50 m...More data for this Ligand-Target Pair
Affinity DataKd: 42.8nMpH: 7.5 T: 2°CAssay Description:The experiments were performed in 384-well black small volume microtiter plates(Greiner) in a buffer composed of 50mM Hepes, pH 7.5 (Applichem), 50 m...More data for this Ligand-Target Pair
Affinity DataKd: 61.3nMpH: 7.5 T: 2°CAssay Description:The experiments were performed in 384-well black small volume microtiter plates(Greiner) in a buffer composed of 50mM Hepes, pH 7.5 (Applichem), 50 m...More data for this Ligand-Target Pair
Affinity DataKd: 44nMpH: 7.5 T: 2°CAssay Description:The experiments were performed in 384-well black small volume microtiter plates(Greiner) in a buffer composed of 50mM Hepes, pH 7.5 (Applichem), 50 m...More data for this Ligand-Target Pair
Affinity DataKd: 16.8nMpH: 7.5 T: 2°CAssay Description:The experiments were performed in 384-well black small volume microtiter plates(Greiner) in a buffer composed of 50mM Hepes, pH 7.5 (Applichem), 50 m...More data for this Ligand-Target Pair
Affinity DataKd: 112nMpH: 7.5 T: 2°CAssay Description:The experiments were performed in 384-well black small volume microtiter plates(Greiner) in a buffer composed of 50mM Hepes, pH 7.5 (Applichem), 50 m...More data for this Ligand-Target Pair
Affinity DataIC50: 44nMpH: 7.5 T: 2°CAssay Description:Assays were performed with minor modifications from the manufacturer's protocol (PerkinElmer, USA). All reagents were diluted in 50 mM HEPES, 150 mM ...More data for this Ligand-Target Pair
Activity Spreadsheet -- ITC Data from BindingDB
 Found 1 hit for monomerid = 50365262
Found 1 hit for monomerid = 50365262
ITC DataΔG°: -10.1kcal/mole −TΔS°: -3.31kcal/mole ΔH°: -6.80kcal/mole logk: 2.62E+7
T: 25.00°C
T: 25.00°C