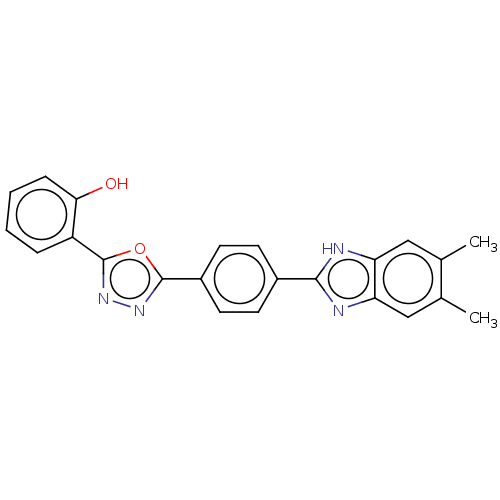
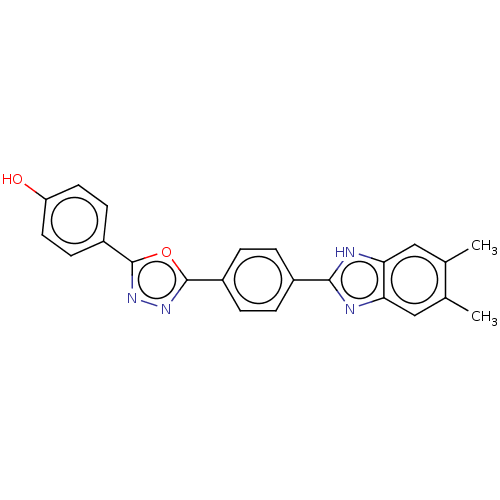
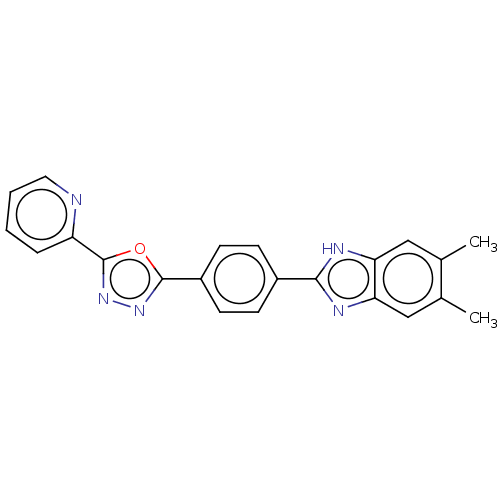
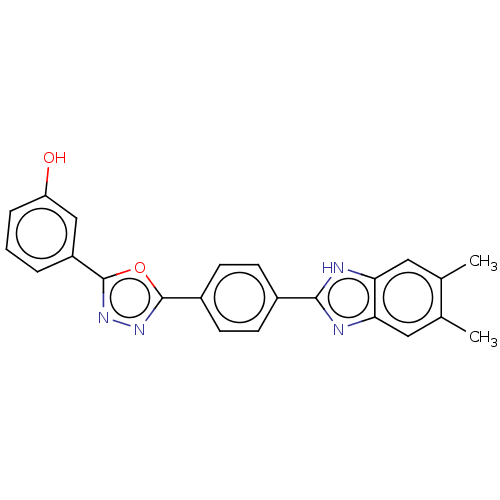
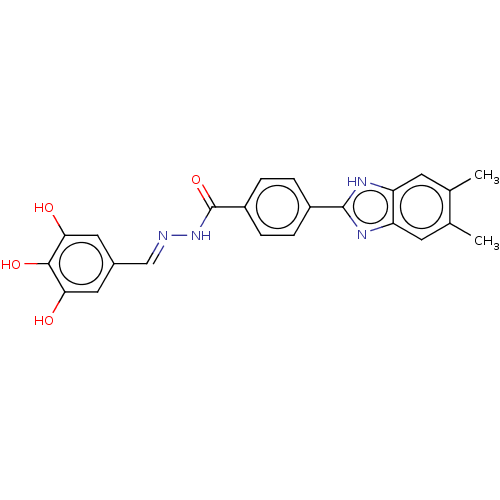
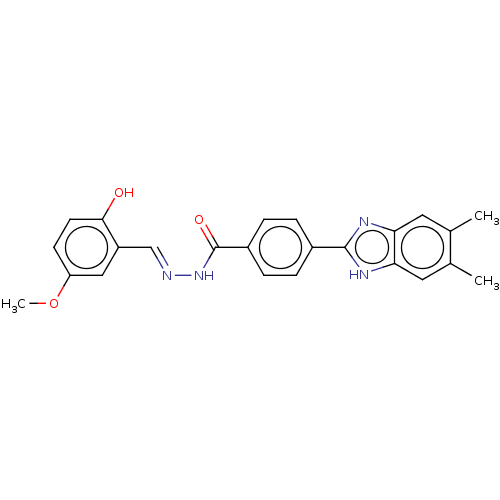
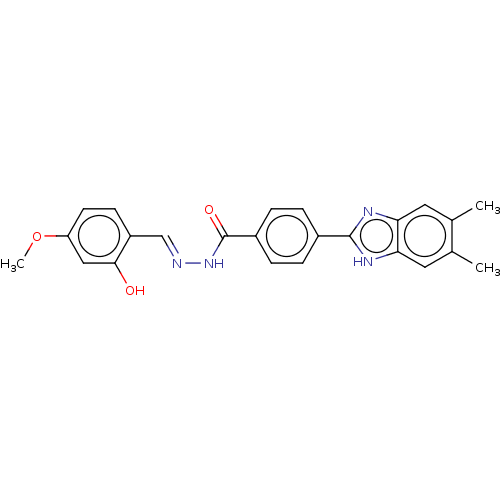
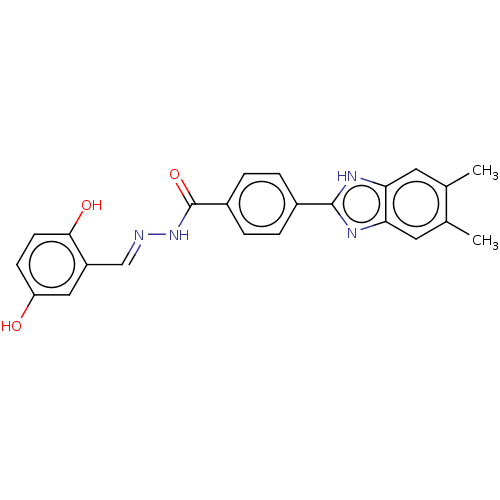
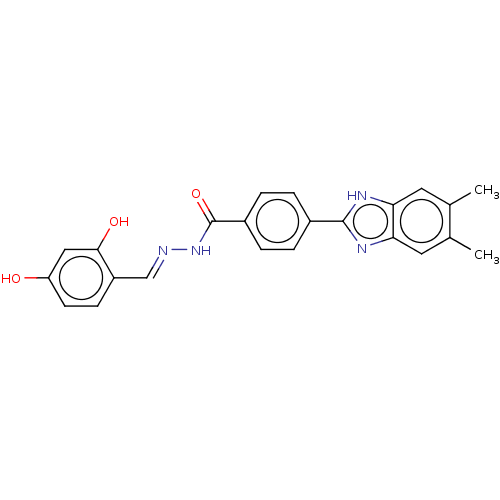
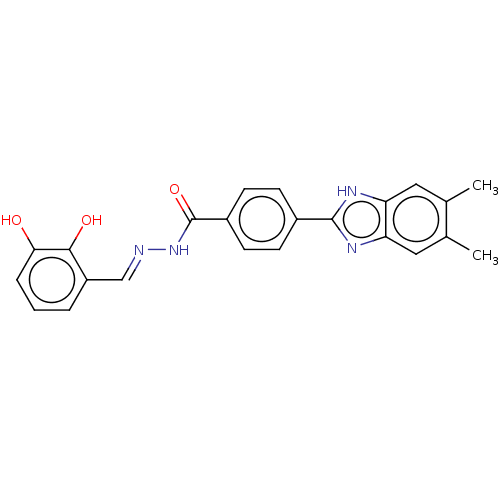
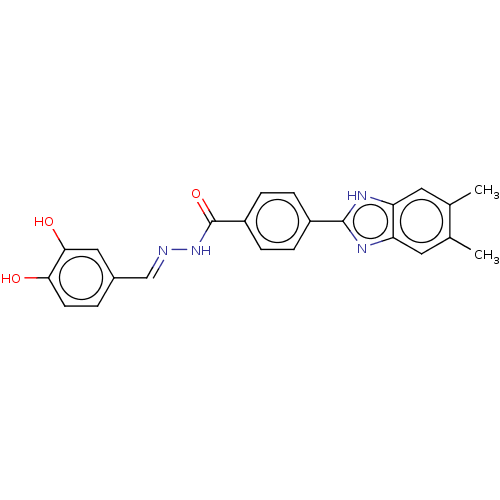
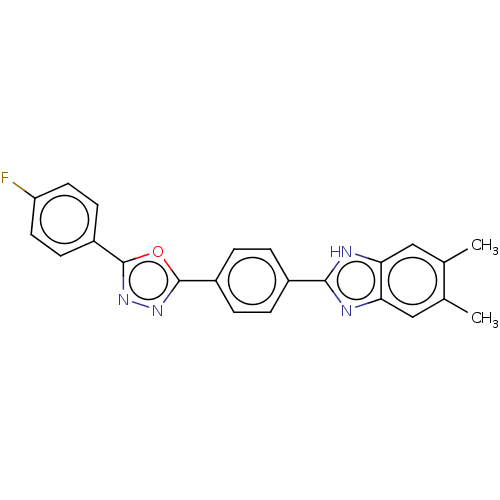
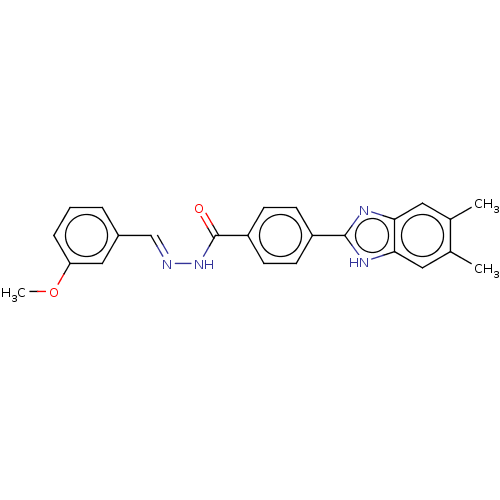
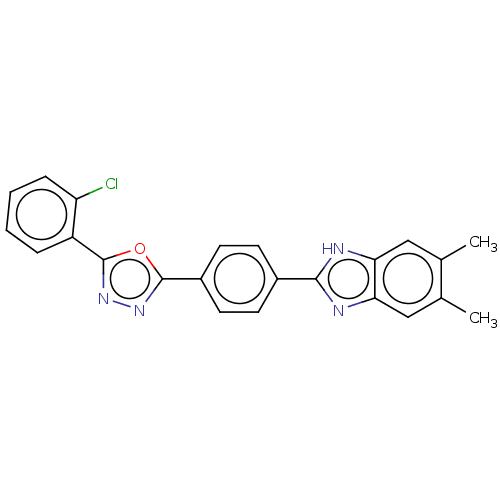
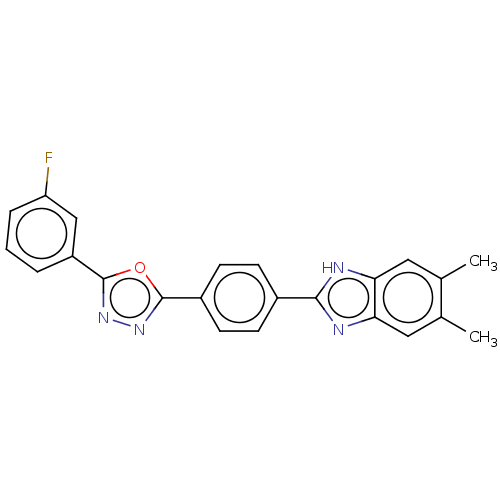
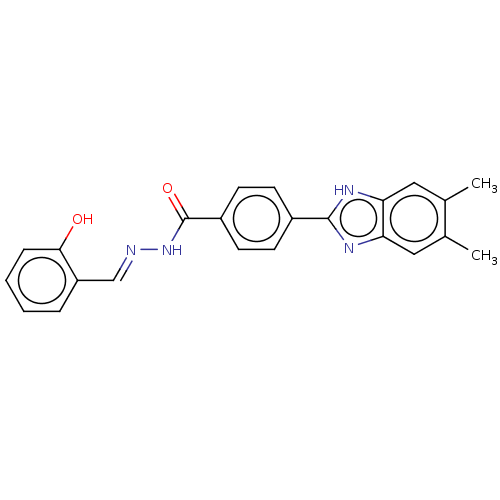
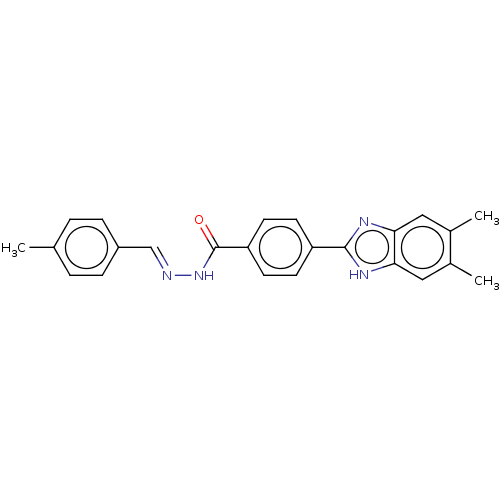
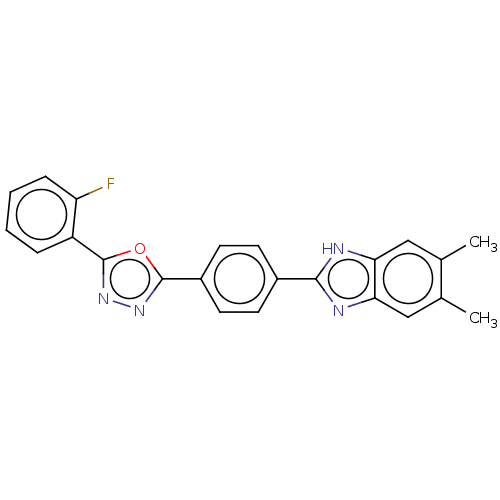
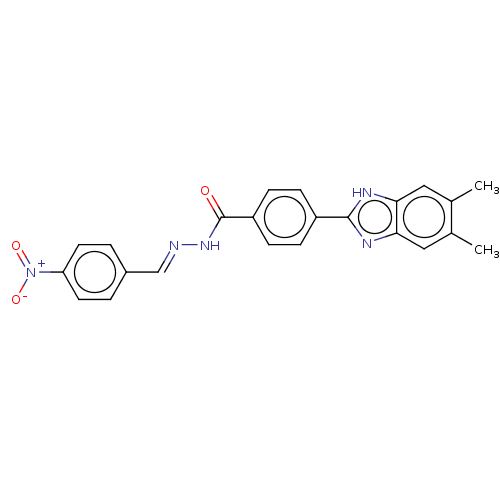
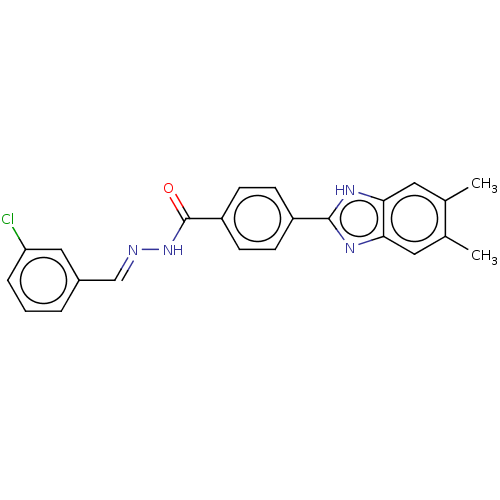
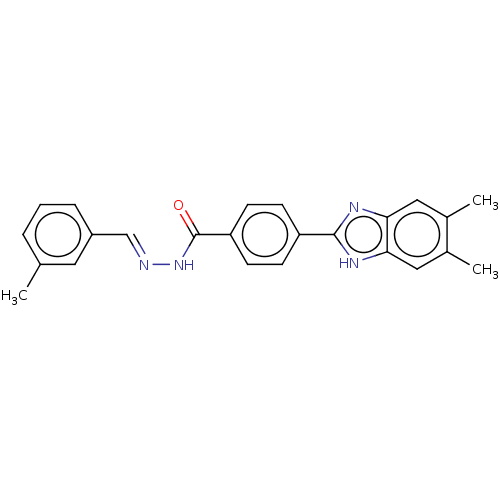
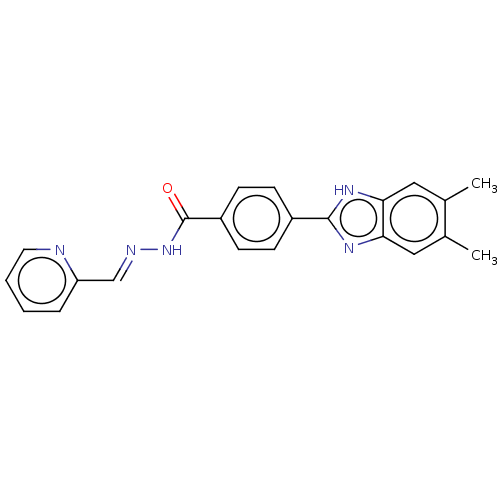
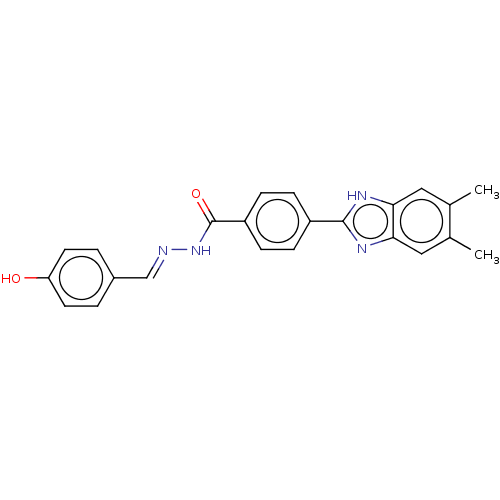
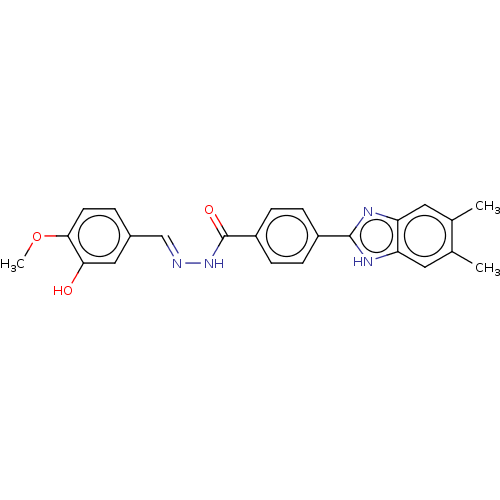
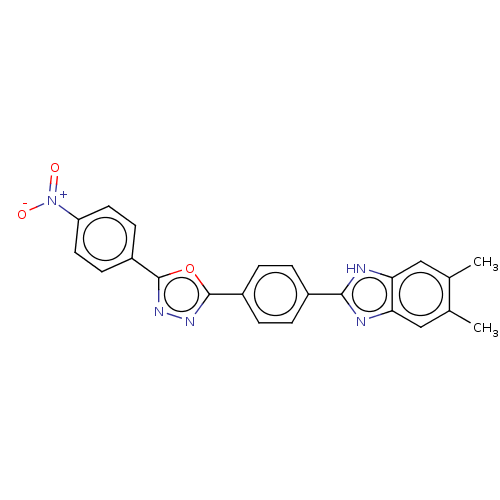
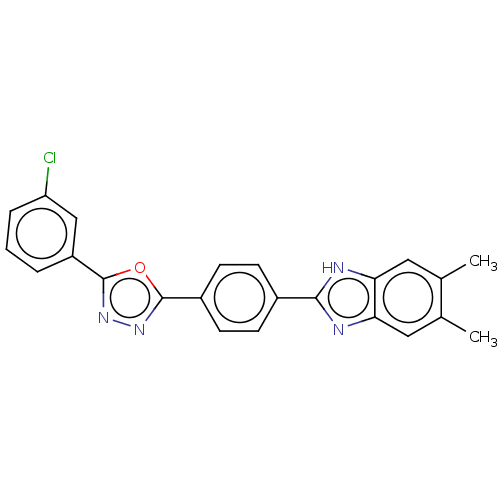
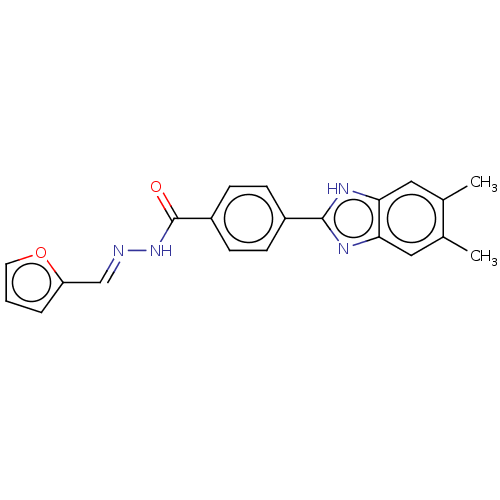
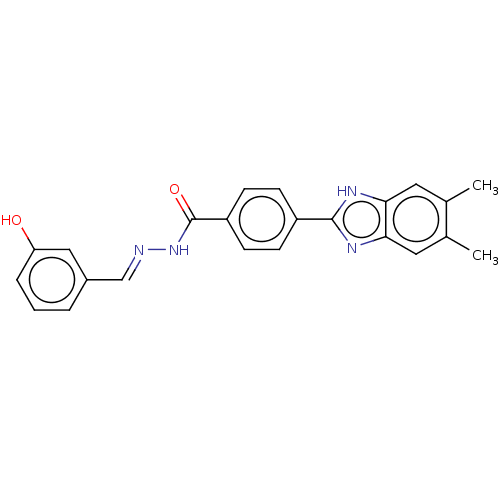
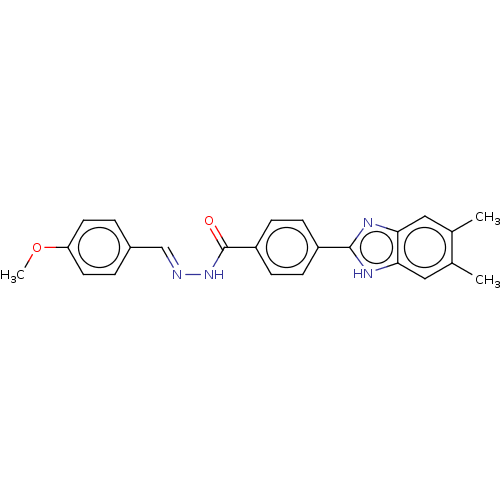
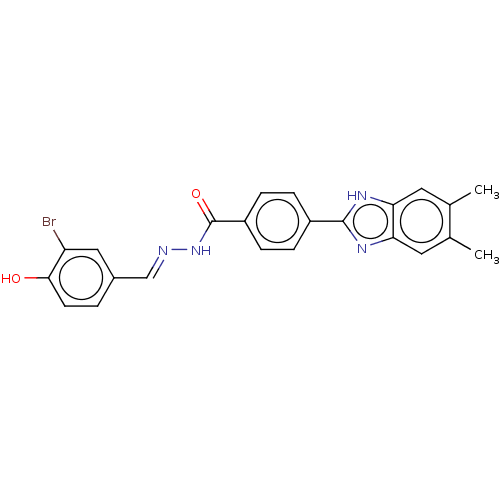
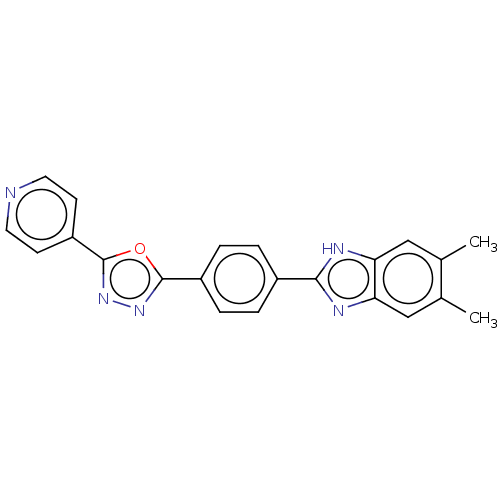
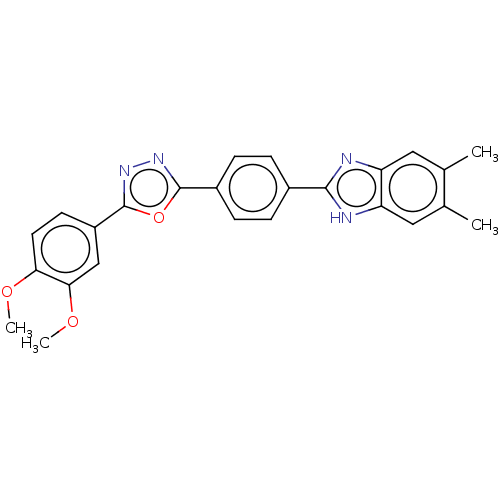
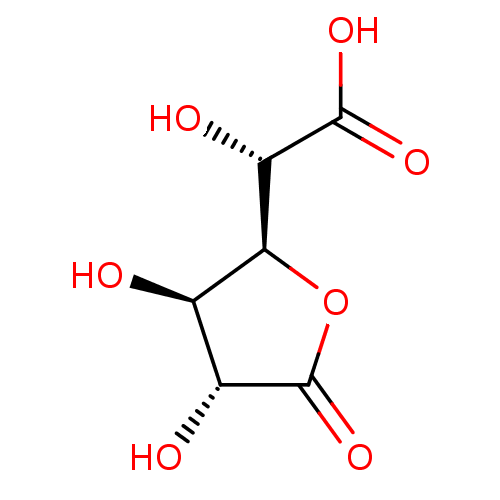
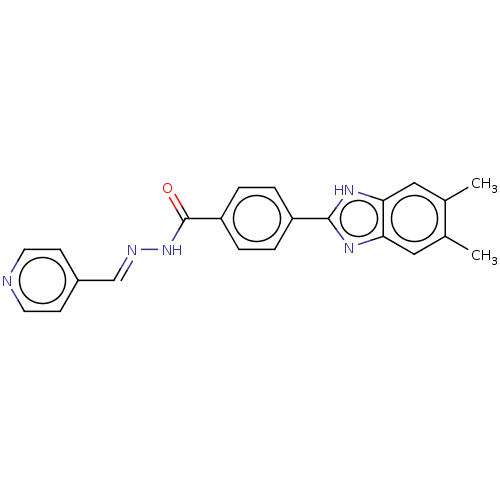
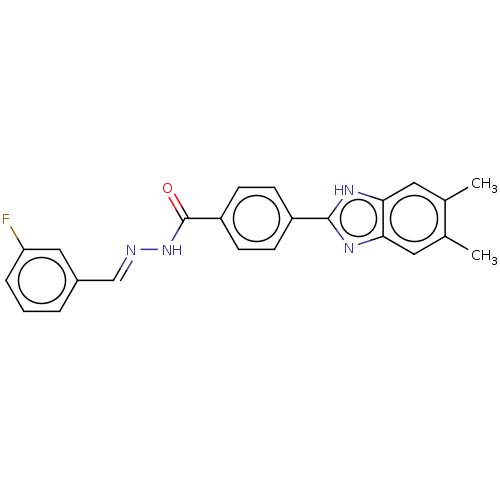
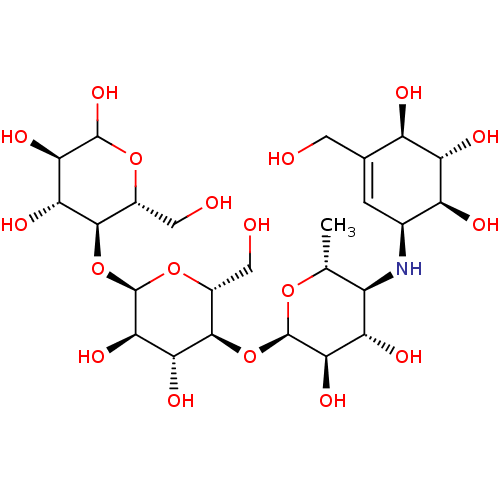
Affinity DataIC50: 2.14E+3nMAssay Description:Inhibition of beta-D-glucuronidase (unknown origin) assessed as reduction in p-nitrophenol formation using p-nitrophenyl-beta-D-glucuronide substarte...More data for this Ligand-Target Pair
Affinity DataIC50: 3.14E+3nMAssay Description:Inhibition of beta-D-glucuronidase (unknown origin) assessed as reduction in p-nitrophenol formation using p-nitrophenyl-beta-D-glucuronide substarte...More data for this Ligand-Target Pair
Affinity DataIC50: 4.12E+3nMAssay Description:Inhibition of beta-D-glucuronidase (unknown origin) assessed as reduction in p-nitrophenol formation using p-nitrophenyl-beta-D-glucuronide substarte...More data for this Ligand-Target Pair
Affinity DataIC50: 8.14E+3nMAssay Description:Inhibition of beta-D-glucuronidase (unknown origin) assessed as reduction in p-nitrophenol formation using p-nitrophenyl-beta-D-glucuronide substarte...More data for this Ligand-Target Pair
TargetOligo-1,6-glucosidase IMA1(Saccharomyces cerevisiae S288c (Baker's yeast))
Universiti Teknologi Mara (Uitm)
Universiti Teknologi Mara (Uitm)
Affinity DataIC50: 8.40E+3nMpH: 6.8Assay Description:The enzyme inhibition was evaluated according to the method previously reported by Rahim et al. [25] with slight modification. Various concentration ...More data for this Ligand-Target Pair
TargetOligo-1,6-glucosidase IMA1(Saccharomyces cerevisiae S288c (Baker's yeast))
Universiti Teknologi Mara (Uitm)
Universiti Teknologi Mara (Uitm)
Affinity DataIC50: 8.44E+3nMpH: 6.8Assay Description:The enzyme inhibition was evaluated according to the method previously reported by Rahim et al. [25] with slight modification. Various concentration ...More data for this Ligand-Target Pair
TargetOligo-1,6-glucosidase IMA1(Saccharomyces cerevisiae S288c (Baker's yeast))
Universiti Teknologi Mara (Uitm)
Universiti Teknologi Mara (Uitm)
Affinity DataIC50: 8.44E+3nMpH: 6.8Assay Description:The enzyme inhibition was evaluated according to the method previously reported by Rahim et al. [25] with slight modification. Various concentration ...More data for this Ligand-Target Pair
TargetOligo-1,6-glucosidase IMA1(Saccharomyces cerevisiae S288c (Baker's yeast))
Universiti Teknologi Mara (Uitm)
Universiti Teknologi Mara (Uitm)
Affinity DataIC50: 8.74E+3nMpH: 6.8Assay Description:The enzyme inhibition was evaluated according to the method previously reported by Rahim et al. [25] with slight modification. Various concentration ...More data for this Ligand-Target Pair
TargetOligo-1,6-glucosidase IMA1(Saccharomyces cerevisiae S288c (Baker's yeast))
Universiti Teknologi Mara (Uitm)
Universiti Teknologi Mara (Uitm)
Affinity DataIC50: 8.74E+3nMpH: 6.8Assay Description:The enzyme inhibition was evaluated according to the method previously reported by Rahim et al. [25] with slight modification. Various concentration ...More data for this Ligand-Target Pair
TargetOligo-1,6-glucosidase IMA1(Saccharomyces cerevisiae S288c (Baker's yeast))
Universiti Teknologi Mara (Uitm)
Universiti Teknologi Mara (Uitm)
Affinity DataIC50: 9.99E+3nMpH: 6.8Assay Description:The enzyme inhibition was evaluated according to the method previously reported by Rahim et al. [25] with slight modification. Various concentration ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.11E+4nMAssay Description:Inhibition of beta-D-glucuronidase (unknown origin) assessed as reduction in p-nitrophenol formation using p-nitrophenyl-beta-D-glucuronide substarte...More data for this Ligand-Target Pair
TargetOligo-1,6-glucosidase IMA1(Saccharomyces cerevisiae S288c (Baker's yeast))
Universiti Teknologi Mara (Uitm)
Universiti Teknologi Mara (Uitm)
Affinity DataIC50: 1.25E+4nMpH: 6.8Assay Description:The enzyme inhibition was evaluated according to the method previously reported by Rahim et al. [25] with slight modification. Various concentration ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.31E+4nMAssay Description:Inhibition of beta-D-glucuronidase (unknown origin) assessed as reduction in p-nitrophenol formation using p-nitrophenyl-beta-D-glucuronide substarte...More data for this Ligand-Target Pair
TargetOligo-1,6-glucosidase IMA1(Saccharomyces cerevisiae S288c (Baker's yeast))
Universiti Teknologi Mara (Uitm)
Universiti Teknologi Mara (Uitm)
Affinity DataIC50: 1.51E+4nMpH: 6.8Assay Description:The enzyme inhibition was evaluated according to the method previously reported by Rahim et al. [25] with slight modification. Various concentration ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.51E+4nMAssay Description:Inhibition of beta-D-glucuronidase (unknown origin) assessed as reduction in p-nitrophenol formation using p-nitrophenyl-beta-D-glucuronide substarte...More data for this Ligand-Target Pair
Affinity DataIC50: 1.61E+4nMAssay Description:Inhibition of beta-D-glucuronidase (unknown origin) assessed as reduction in p-nitrophenol formation using p-nitrophenyl-beta-D-glucuronide substarte...More data for this Ligand-Target Pair
TargetOligo-1,6-glucosidase IMA1(Saccharomyces cerevisiae S288c (Baker's yeast))
Universiti Teknologi Mara (Uitm)
Universiti Teknologi Mara (Uitm)
Affinity DataIC50: 1.82E+4nMpH: 6.8Assay Description:The enzyme inhibition was evaluated according to the method previously reported by Rahim et al. [25] with slight modification. Various concentration ...More data for this Ligand-Target Pair
TargetOligo-1,6-glucosidase IMA1(Saccharomyces cerevisiae S288c (Baker's yeast))
Universiti Teknologi Mara (Uitm)
Universiti Teknologi Mara (Uitm)
Affinity DataIC50: 1.83E+4nMpH: 6.8Assay Description:The enzyme inhibition was evaluated according to the method previously reported by Rahim et al. [25] with slight modification. Various concentration ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.92E+4nMAssay Description:Inhibition of beta-D-glucuronidase (unknown origin) assessed as reduction in p-nitrophenol formation using p-nitrophenyl-beta-D-glucuronide substarte...More data for this Ligand-Target Pair
TargetOligo-1,6-glucosidase IMA1(Saccharomyces cerevisiae S288c (Baker's yeast))
Universiti Teknologi Mara (Uitm)
Universiti Teknologi Mara (Uitm)
Affinity DataIC50: 1.94E+4nMpH: 6.8Assay Description:The enzyme inhibition was evaluated according to the method previously reported by Rahim et al. [25] with slight modification. Various concentration ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.01E+4nMAssay Description:Inhibition of beta-D-glucuronidase (unknown origin) assessed as reduction in p-nitrophenol formation using p-nitrophenyl-beta-D-glucuronide substarte...More data for this Ligand-Target Pair
TargetOligo-1,6-glucosidase IMA1(Saccharomyces cerevisiae S288c (Baker's yeast))
Universiti Teknologi Mara (Uitm)
Universiti Teknologi Mara (Uitm)
Affinity DataIC50: 2.24E+4nMpH: 6.8Assay Description:The enzyme inhibition was evaluated according to the method previously reported by Rahim et al. [25] with slight modification. Various concentration ...More data for this Ligand-Target Pair
TargetOligo-1,6-glucosidase IMA1(Saccharomyces cerevisiae S288c (Baker's yeast))
Universiti Teknologi Mara (Uitm)
Universiti Teknologi Mara (Uitm)
Affinity DataIC50: 2.35E+4nMpH: 6.8Assay Description:The enzyme inhibition was evaluated according to the method previously reported by Rahim et al. [25] with slight modification. Various concentration ...More data for this Ligand-Target Pair
TargetOligo-1,6-glucosidase IMA1(Saccharomyces cerevisiae S288c (Baker's yeast))
Universiti Teknologi Mara (Uitm)
Universiti Teknologi Mara (Uitm)
Affinity DataIC50: 2.48E+4nMpH: 6.8Assay Description:The enzyme inhibition was evaluated according to the method previously reported by Rahim et al. [25] with slight modification. Various concentration ...More data for this Ligand-Target Pair
TargetOligo-1,6-glucosidase IMA1(Saccharomyces cerevisiae S288c (Baker's yeast))
Universiti Teknologi Mara (Uitm)
Universiti Teknologi Mara (Uitm)
Affinity DataIC50: 2.48E+4nMpH: 6.8Assay Description:The enzyme inhibition was evaluated according to the method previously reported by Rahim et al. [25] with slight modification. Various concentration ...More data for this Ligand-Target Pair
TargetOligo-1,6-glucosidase IMA1(Saccharomyces cerevisiae S288c (Baker's yeast))
Universiti Teknologi Mara (Uitm)
Universiti Teknologi Mara (Uitm)
Affinity DataIC50: 2.57E+4nMpH: 6.8Assay Description:The enzyme inhibition was evaluated according to the method previously reported by Rahim et al. [25] with slight modification. Various concentration ...More data for this Ligand-Target Pair
TargetOligo-1,6-glucosidase IMA1(Saccharomyces cerevisiae S288c (Baker's yeast))
Universiti Teknologi Mara (Uitm)
Universiti Teknologi Mara (Uitm)
Affinity DataIC50: 2.60E+4nMpH: 6.8Assay Description:The enzyme inhibition was evaluated according to the method previously reported by Rahim et al. [25] with slight modification. Various concentration ...More data for this Ligand-Target Pair
TargetOligo-1,6-glucosidase IMA1(Saccharomyces cerevisiae S288c (Baker's yeast))
Universiti Teknologi Mara (Uitm)
Universiti Teknologi Mara (Uitm)
Affinity DataIC50: 2.90E+4nMpH: 6.8Assay Description:The enzyme inhibition was evaluated according to the method previously reported by Rahim et al. [25] with slight modification. Various concentration ...More data for this Ligand-Target Pair
Affinity DataIC50: 3.01E+4nMAssay Description:Inhibition of beta-D-glucuronidase (unknown origin) assessed as reduction in p-nitrophenol formation using p-nitrophenyl-beta-D-glucuronide substarte...More data for this Ligand-Target Pair
Affinity DataIC50: 3.11E+4nMAssay Description:Inhibition of beta-D-glucuronidase (unknown origin) assessed as reduction in p-nitrophenol formation using p-nitrophenyl-beta-D-glucuronide substarte...More data for this Ligand-Target Pair
TargetOligo-1,6-glucosidase IMA1(Saccharomyces cerevisiae S288c (Baker's yeast))
Universiti Teknologi Mara (Uitm)
Universiti Teknologi Mara (Uitm)
Affinity DataIC50: 3.21E+4nMpH: 6.8Assay Description:The enzyme inhibition was evaluated according to the method previously reported by Rahim et al. [25] with slight modification. Various concentration ...More data for this Ligand-Target Pair
Affinity DataIC50: 3.21E+4nMAssay Description:Inhibition of beta-D-glucuronidase (unknown origin) assessed as reduction in p-nitrophenol formation using p-nitrophenyl-beta-D-glucuronide substarte...More data for this Ligand-Target Pair
TargetOligo-1,6-glucosidase IMA1(Saccharomyces cerevisiae S288c (Baker's yeast))
Universiti Teknologi Mara (Uitm)
Universiti Teknologi Mara (Uitm)
Affinity DataIC50: 3.25E+4nMpH: 6.8Assay Description:The enzyme inhibition was evaluated according to the method previously reported by Rahim et al. [25] with slight modification. Various concentration ...More data for this Ligand-Target Pair
TargetOligo-1,6-glucosidase IMA1(Saccharomyces cerevisiae S288c (Baker's yeast))
Universiti Teknologi Mara (Uitm)
Universiti Teknologi Mara (Uitm)
Affinity DataIC50: 3.26E+4nMpH: 6.8Assay Description:The enzyme inhibition was evaluated according to the method previously reported by Rahim et al. [25] with slight modification. Various concentration ...More data for this Ligand-Target Pair
TargetOligo-1,6-glucosidase IMA1(Saccharomyces cerevisiae S288c (Baker's yeast))
Universiti Teknologi Mara (Uitm)
Universiti Teknologi Mara (Uitm)
Affinity DataIC50: 3.45E+4nMpH: 6.8Assay Description:The enzyme inhibition was evaluated according to the method previously reported by Rahim et al. [25] with slight modification. Various concentration ...More data for this Ligand-Target Pair
Affinity DataIC50: 4.01E+4nMAssay Description:Inhibition of beta-D-glucuronidase (unknown origin) assessed as reduction in p-nitrophenol formation using p-nitrophenyl-beta-D-glucuronide substarte...More data for this Ligand-Target Pair
TargetOligo-1,6-glucosidase IMA1(Saccharomyces cerevisiae S288c (Baker's yeast))
Universiti Teknologi Mara (Uitm)
Universiti Teknologi Mara (Uitm)
Affinity DataIC50: 4.60E+4nMpH: 6.8Assay Description:The enzyme inhibition was evaluated according to the method previously reported by Rahim et al. [25] with slight modification. Various concentration ...More data for this Ligand-Target Pair
Affinity DataIC50: 4.61E+4nMAssay Description:Inhibition of beta-D-glucuronidase (unknown origin) assessed as reduction in p-nitrophenol formation using p-nitrophenyl-beta-D-glucuronide substarte...More data for this Ligand-Target Pair
Affinity DataIC50: 4.84E+4nMAssay Description:Inhibition of beta-D-glucuronidase (unknown origin) assessed as reduction in p-nitrophenol formation using p-nitrophenyl-beta-D-glucuronide substarte...More data for this Ligand-Target Pair
TargetOligo-1,6-glucosidase IMA1(Saccharomyces cerevisiae S288c (Baker's yeast))
Universiti Teknologi Mara (Uitm)
Universiti Teknologi Mara (Uitm)
Affinity DataIC50: 1.48E+5nMpH: 6.8Assay Description:The enzyme inhibition was evaluated according to the method previously reported by Rahim et al. [25] with slight modification. Various concentration ...More data for this Ligand-Target Pair
TargetOligo-1,6-glucosidase IMA1(Saccharomyces cerevisiae S288c (Baker's yeast))
Universiti Teknologi Mara (Uitm)
Universiti Teknologi Mara (Uitm)
Affinity DataIC50: 1.58E+5nMpH: 6.8Assay Description:The enzyme inhibition was evaluated according to the method previously reported by Rahim et al. [25] with slight modification. Various concentration ...More data for this Ligand-Target Pair
TargetOligo-1,6-glucosidase IMA1(Saccharomyces cerevisiae S288c (Baker's yeast))
Universiti Teknologi Mara (Uitm)
Universiti Teknologi Mara (Uitm)
Affinity DataIC50: 1.80E+5nMpH: 6.8Assay Description:The enzyme inhibition was evaluated according to the method previously reported by Rahim et al. [25] with slight modification. Various concentration ...More data for this Ligand-Target Pair
TargetOligo-1,6-glucosidase IMA1(Saccharomyces cerevisiae S288c (Baker's yeast))
Universiti Teknologi Mara (Uitm)
Universiti Teknologi Mara (Uitm)
Affinity DataIC50: 7.75E+5nMpH: 6.8Assay Description:The enzyme inhibition was evaluated according to the method previously reported by Rahim et al. [25] with slight modification. Various concentration ...More data for this Ligand-Target Pair