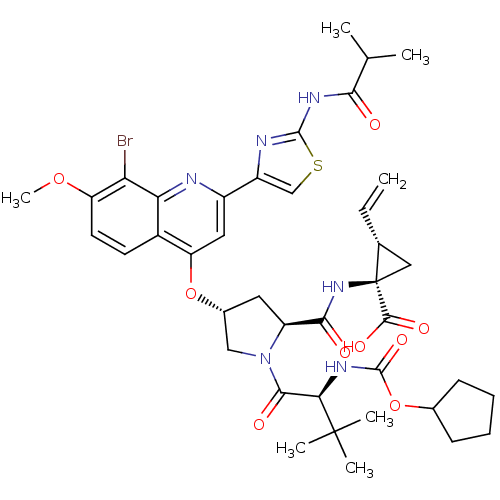
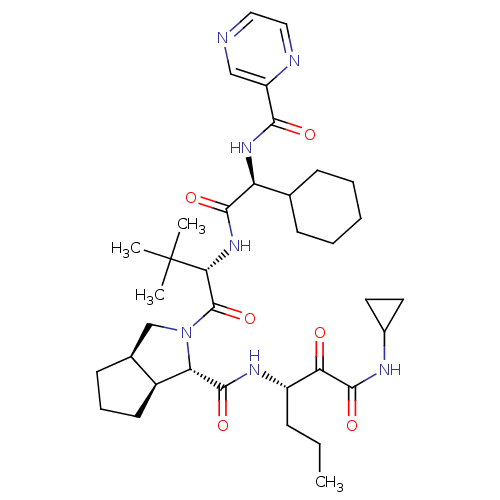
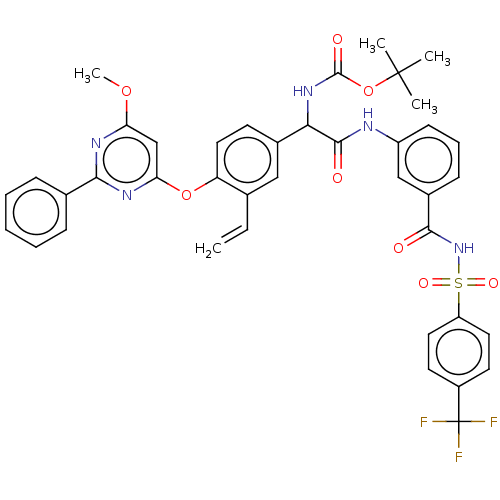
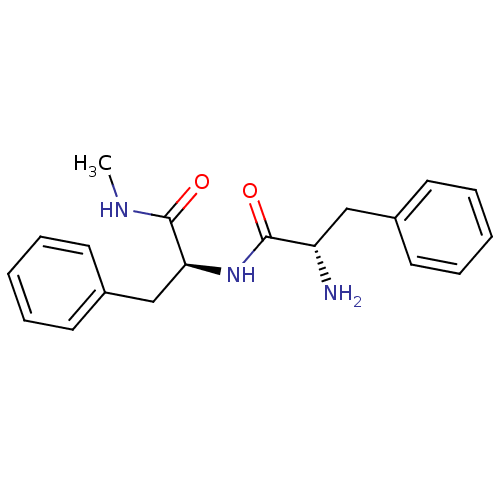
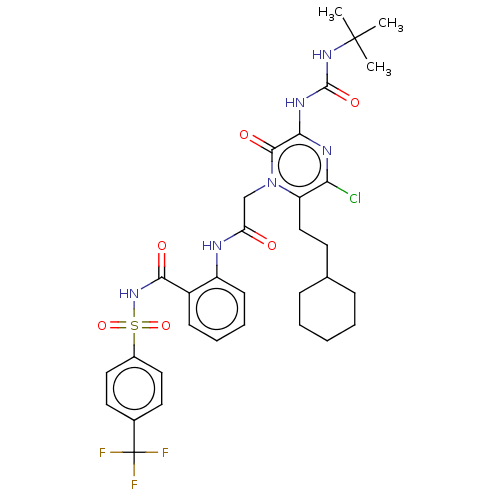
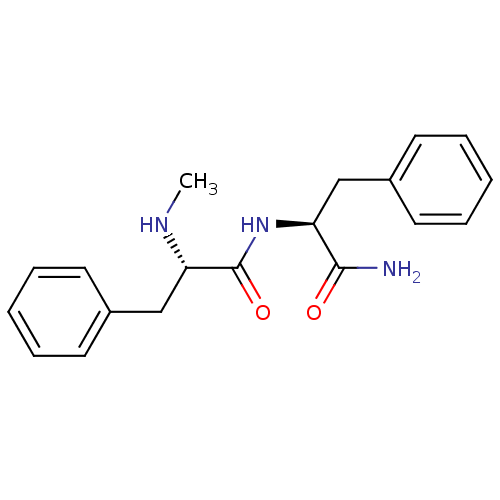
Affinity DataKi: 0.0500nMAssay Description:Inhibition of HCV genotype 1a NS3/4A protease using Ac-DED(Edans)EEAbu-psi[COO]ASK(Dabcyl)-NH2 as substrate by FRET assayMore data for this Ligand-Target Pair
Affinity DataKi: 0.300nMAssay Description:Inhibition of HCV genotype 1a NS3/4A protease using Ac-DED(Edans)EEAbu-psi[COO]ASK(Dabcyl)-NH2 as substrate by FRET assayMore data for this Ligand-Target Pair
Affinity DataKi: 0.300nMAssay Description:Binding affinity to substance P receptor (1 to 7 amino acids) binding site in rat spinal cord membranesMore data for this Ligand-Target Pair
Affinity DataKi: 0.360nMAssay Description:Inhibition of HCV genotype 1a NS3/4A protease using Ac-DED(Edans)EEAbu-psi[COO]ASK(Dabcyl)-NH2 as substrate by FRET assayMore data for this Ligand-Target Pair
Affinity DataKi: 1.20nMAssay Description:Inhibition of HCV genotype 1a NS3/4A protease using Ac-DED(Edans)EEAbu-psi[COO]ASK(Dabcyl)-NH2 as substrate by FRET assayMore data for this Ligand-Target Pair
Affinity DataKi: 1.5nMAssay Description:Binding affinity to NK1 receptor (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 1.5nMAssay Description:Displacement of [3H]-SP1-7 from NK1 receptor (unknown origin) by scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataKi: 1.60nMAssay Description:Binding affinity to substance P receptor (1 to 7 amino acids) binding site in rat spinal cord membranesMore data for this Ligand-Target Pair
Affinity DataKi: 2.20nMAssay Description:Binding affinity to NK1 receptor (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 2.20nMAssay Description:Displacement of [3H]-SP1-7 from NK1 receptor (unknown origin) by scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataKi: 2.20nMAssay Description:Displacement of [3H]SP1-7 from substance P receptor (1 to 7 amino acids) binding site in Sprague-Dawley rat spinal cord membranes after 60 mins by li...More data for this Ligand-Target Pair
Affinity DataKi: 2.80nMAssay Description:Inhibition of HCV genotype 1b NS3/4A proteaseMore data for this Ligand-Target Pair
Affinity DataKi: 3.30nMAssay Description:Displacement of [3H]SP1-7 from substance P receptor (1 to 7 amino acids) binding site in Sprague-Dawley rat spinal cord membranes after 60 mins by li...More data for this Ligand-Target Pair
TargetGenome polyprotein(Hepatitis C virus genotype 1b (isolate Con1) (HCV))
Uppsala University
Curated by ChEMBL
Uppsala University
Curated by ChEMBL
Affinity DataKi: 3.90nMAssay Description:Inhibition of full-length HCV NS3 protease D168V mutantMore data for this Ligand-Target Pair
Affinity DataKi: 5.20nMAssay Description:Binding affinity to NK1 receptor (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 5.20nMAssay Description:Displacement of [3H]-SP1-7 from NK1 receptor (unknown origin) by scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataKi: 6.20nMAssay Description:Displacement of [3H]SP1-7 from substance P receptor (1 to 7 amino acids) binding site in Sprague-Dawley rat spinal cord membranes after 60 mins by li...More data for this Ligand-Target Pair
Affinity DataKi: 8.40nMAssay Description:Displacement of [3H]SP1-7 from substance P receptor (1 to 7 amino acids) binding site in Sprague-Dawley rat spinal cord membranes after 60 mins by li...More data for this Ligand-Target Pair
Affinity DataKi: 9.40nMAssay Description:Displacement of [3H]SP1-7 from substance P receptor (1 to 7 amino acids) binding site in Sprague-Dawley rat spinal cord membranes after 60 mins by li...More data for this Ligand-Target Pair
Affinity DataKi: 12nMAssay Description:Displacement of [3H]SP1-7 from substance P receptor (1 to 7 amino acids) binding site in Sprague-Dawley rat spinal cord membranes after 60 mins by li...More data for this Ligand-Target Pair
Affinity DataKi: 14nMAssay Description:Inhibition of HCV genotype 1a NS3/4A protease using Ac-DED(Edans)EEAbu-psi[COO]ASK(Dabcyl)-NH2 as substrate by FRET assayMore data for this Ligand-Target Pair
Affinity DataKi: 18nMAssay Description:Displacement of [3H]SP1-7 from substance P receptor (1 to 7 amino acids) binding site in Sprague-Dawley rat spinal cord membranes after 60 mins by li...More data for this Ligand-Target Pair
Affinity DataKi: 24nMAssay Description:Inhibition of full-length HCV NS3 protease R155K mutantMore data for this Ligand-Target Pair
Affinity DataKi: 26nMAssay Description:Displacement of [3H]SP1-7 from substance P receptor (1 to 7 amino acids) binding site in Sprague-Dawley rat spinal cord membranes after 60 mins by li...More data for this Ligand-Target Pair
Affinity DataKi: 26nMAssay Description:Inhibition of HCV genotype 1a NS3/4A protease using Ac-DED(Edans)EEAbu-psi[COO]ASK(Dabcyl)-NH2 as substrate by FRET assayMore data for this Ligand-Target Pair
Affinity DataKi: 33nMAssay Description:Inhibition of HCV genotype 1a NS3/4A protease using Ac-DED(Edans)EEAbu-psi[COO]ASK(Dabcyl)-NH2 as substrate by FRET assayMore data for this Ligand-Target Pair
Affinity DataKi: 34nMAssay Description:Displacement of [3H]SP1-7 from substance P receptor (1 to 7 amino acids) binding site in Sprague-Dawley rat spinal cord membranes after 60 mins by li...More data for this Ligand-Target Pair
Affinity DataKi: 40nMAssay Description:Inhibition of HCV genotype 1a NS3/4A protease using Ac-DED(Edans)EEAbu-psi[COO]ASK(Dabcyl)-NH2 as substrate by FRET assayMore data for this Ligand-Target Pair
Affinity DataKi: 51nMAssay Description:Displacement of [3H]SP1-7 from substance P receptor (1 to 7 amino acids) binding site in Sprague-Dawley rat spinal cord membranes after 60 mins by li...More data for this Ligand-Target Pair
Affinity DataKi: 53nMAssay Description:Inhibition of Hepatitis C virus genotype 1a full length NS3/2K-NS4A protease A156T mutant using Ac-DED(Edans)EEAbupsi[COO]ASK(Dabcyl)-NH2 as substrat...More data for this Ligand-Target Pair
Affinity DataKi: 63nMAssay Description:Inhibition of HCV genotype 1a NS3/4A protease using Ac-DED(Edans)EEAbu-psi[COO]ASK(Dabcyl)-NH2 as substrate by FRET assayMore data for this Ligand-Target Pair
Affinity DataKi: 68nMAssay Description:Displacement of [3H]SP1-7 from substance P receptor (1 to 7 amino acids) binding site in Sprague-Dawley rat spinal cord membranes after 60 mins by li...More data for this Ligand-Target Pair
Affinity DataKi: 70nMAssay Description:Displacement of [3H]SP1-7 from substance P receptor (1 to 7 amino acids) binding site in Sprague-Dawley rat spinal cord membranes after 60 mins by li...More data for this Ligand-Target Pair
Affinity DataKi: 80nMAssay Description:Inhibition of HCV genotype 1a NS3/4A protease using Ac-DED(Edans)EEAbu-psi[COO]ASK(Dabcyl)-NH2 as substrate by FRET assayMore data for this Ligand-Target Pair
Affinity DataKi: 80nMAssay Description:Inhibition of HCV genotype 1a NS3/4A protease using Ac-DED(Edans)EEAbu-psi[COO]ASK(Dabcyl)-NH2 as substrate by FRET assayMore data for this Ligand-Target Pair
Affinity DataKi: 82nMAssay Description:Inhibition of full-length HCV NS3 protease R155K mutantMore data for this Ligand-Target Pair
Affinity DataKi: 93nMAssay Description:Displacement of [3H]SP1-7 from substance P receptor (1 to 7 amino acids) binding site in Sprague-Dawley rat spinal cord membranes after 60 mins by li...More data for this Ligand-Target Pair
Affinity DataKi: 95nMAssay Description:Inhibition of HCV genotype 1a NS3/4A protease using Ac-DED(Edans)EEAbu-psi[COO]ASK(Dabcyl)-NH2 as substrate by FRET assayMore data for this Ligand-Target Pair
Affinity DataKi: 110nMAssay Description:Inhibition of HCV genotype 1a NS3/4A protease using Ac-DED(Edans)EEAbu-psi[COO]ASK(Dabcyl)-NH2 as substrate by FRET assayMore data for this Ligand-Target Pair
Affinity DataKi: 110nMAssay Description:Inhibition of HCV genotype 1a NS3/4A protease using Ac-DED(Edans)EEAbu-psi[COO]ASK(Dabcyl)-NH2 as substrate by FRET assayMore data for this Ligand-Target Pair
Affinity DataKi: 110nMAssay Description:Inhibition of Hepatitis C virus genotype 1a full length NS3/2K-NS4A protease D168V mutant using Ac-DED(Edans)EEAbupsi[COO]ASK(Dabcyl)-NH2 as substrat...More data for this Ligand-Target Pair
Affinity DataKi: 110nMAssay Description:Inhibition of Hepatitis C virus genotype 1a full length NS3/2K-NS4A protease A156T mutant using Ac-DED(Edans)EEAbupsi[COO]ASK(Dabcyl)-NH2 as substrat...More data for this Ligand-Target Pair
Affinity DataKi: 120nMAssay Description:Inhibition of full-length HCV NS3 protease A156T mutantMore data for this Ligand-Target Pair
Affinity DataKi: 130nMAssay Description:Inhibition of HCV genotype 1a NS3/4A protease using Ac-DED(Edans)EEAbu-psi[COO]ASK(Dabcyl)-NH2 as substrate by FRET assayMore data for this Ligand-Target Pair
Affinity DataKi: 136nMAssay Description:Displacement of [3H]SP1-7 from substance P receptor (1 to 7 amino acids) binding site in Sprague-Dawley rat spinal cord membranes after 60 mins by li...More data for this Ligand-Target Pair
Affinity DataKi: 160nMAssay Description:Inhibition of HCV NS3/4A protease A156T mutant using Ac-DED(Edans)EEAbu-psi[COO]ASK(Dabcyl)-NH2 as substrate by FRET assayMore data for this Ligand-Target Pair
Affinity DataKi: 170nMAssay Description:Inhibition of full-length HCV NS3 protease A156T mutantMore data for this Ligand-Target Pair
Affinity DataKi: 180nMAssay Description:Inhibition of Hepatitis C virus genotype 1a full length NS3/2K-NS4A protease D168V mutant using Ac-DED(Edans)EEAbupsi[COO]ASK(Dabcyl)-NH2 as substrat...More data for this Ligand-Target Pair
Affinity DataKi: 189nMAssay Description:Displacement of [3H]SP1-7 from substance P receptor (1 to 7 amino acids) binding site in Sprague-Dawley rat spinal cord membranes after 60 mins by li...More data for this Ligand-Target Pair
Affinity DataKi: 190nMAssay Description:Inhibition of HCV genotype 1a NS3/4A protease using Ac-DED(Edans)EEAbu-psi[COO]ASK(Dabcyl)-NH2 as substrate by FRET assayMore data for this Ligand-Target Pair

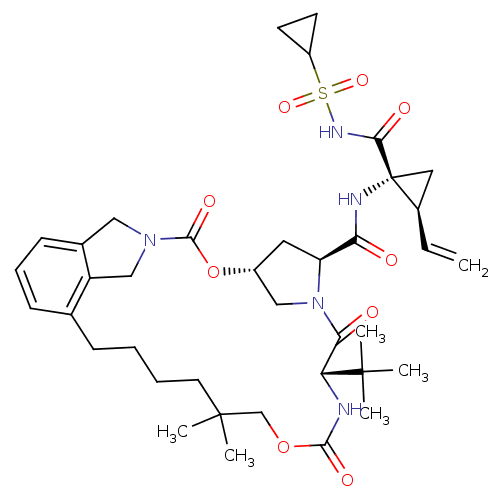
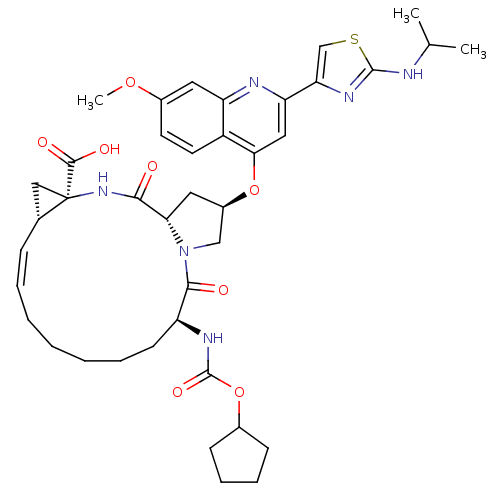

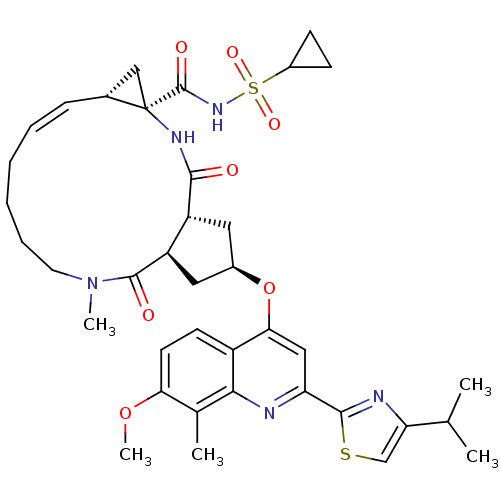
 3D Structure (crystal)
3D Structure (crystal)