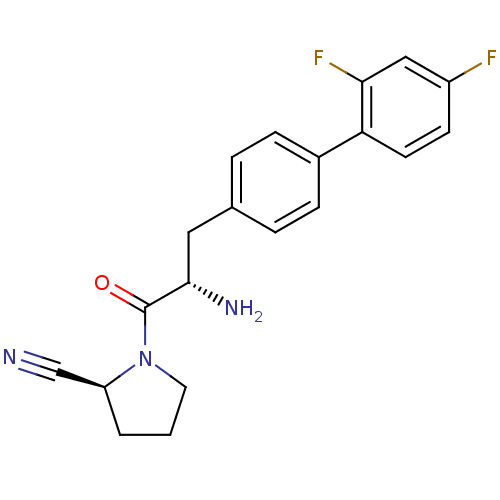
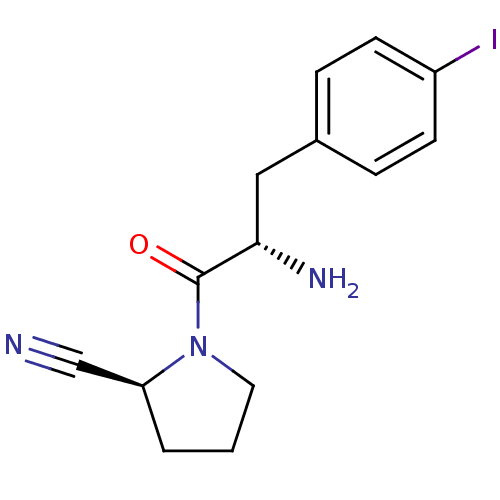
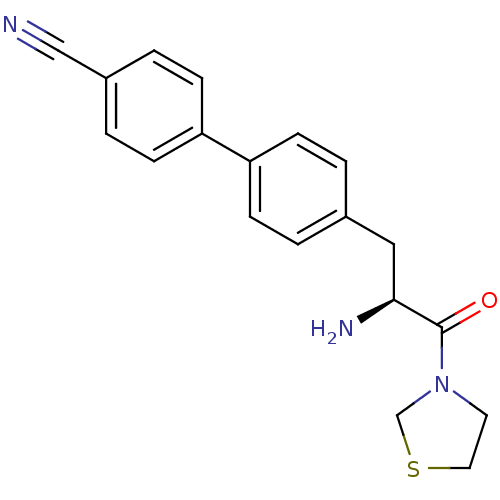
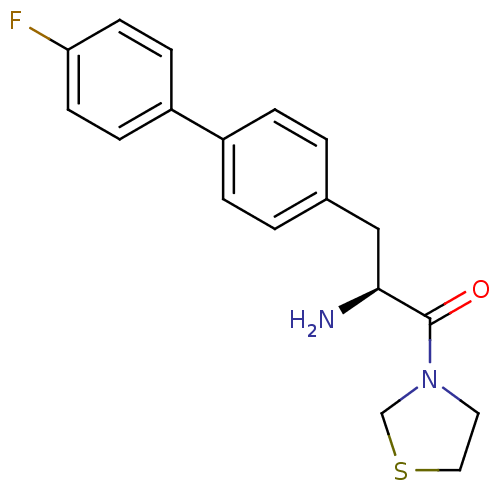
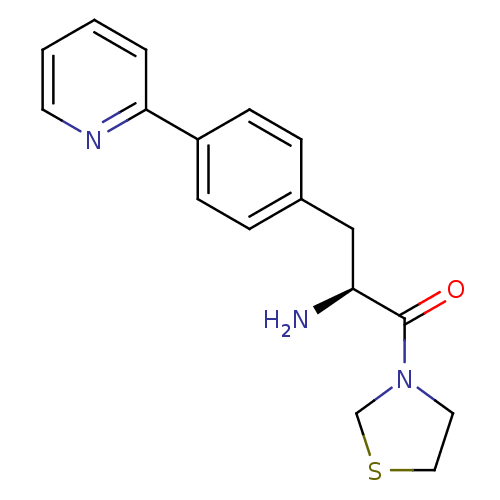
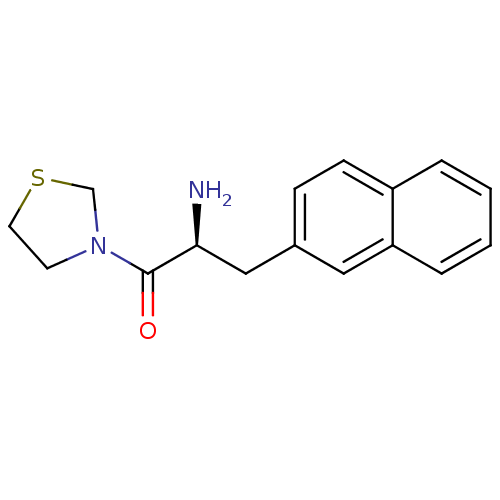
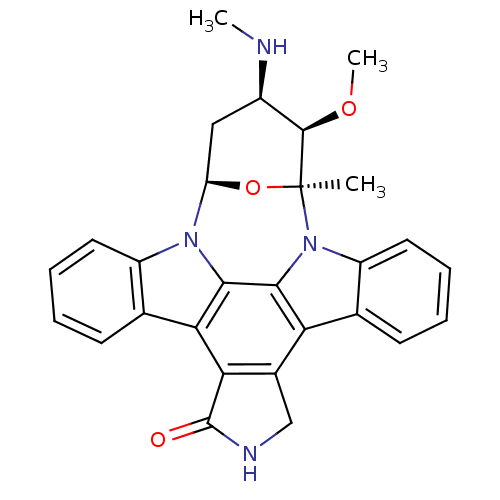
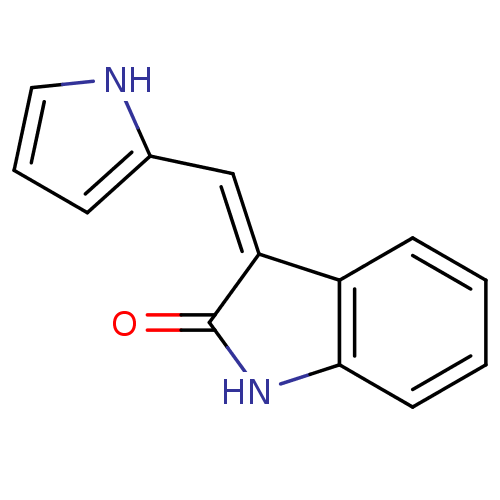
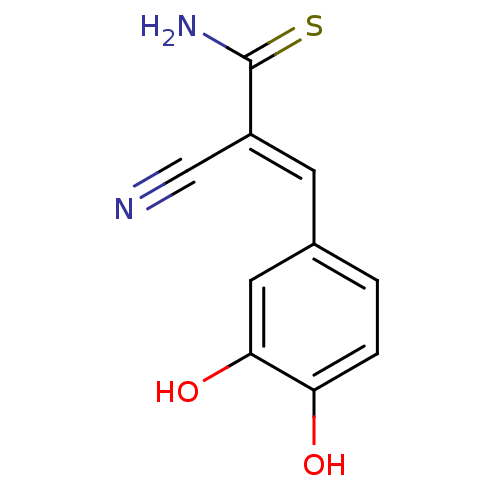
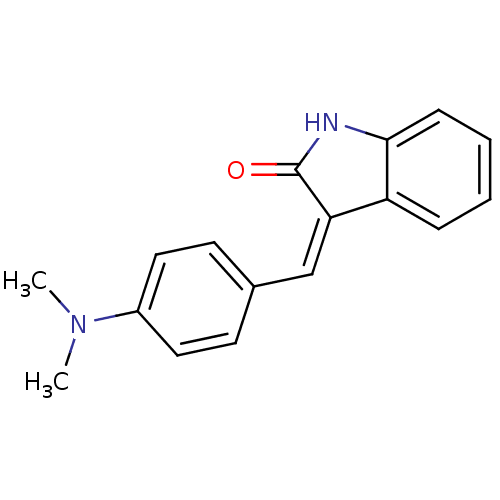
Affinity DataKi: 2.20nM ΔG°: -49.4kJ/molepH: 7.4 T: 2°CAssay Description:The progress of DPPIV inhibition by compounds was measured by recording the liberation of free pNA at 405 nm. IC50 was determined from the slope of r...More data for this Ligand-Target Pair
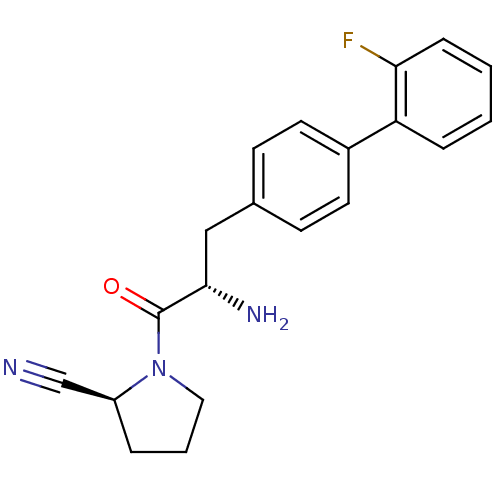
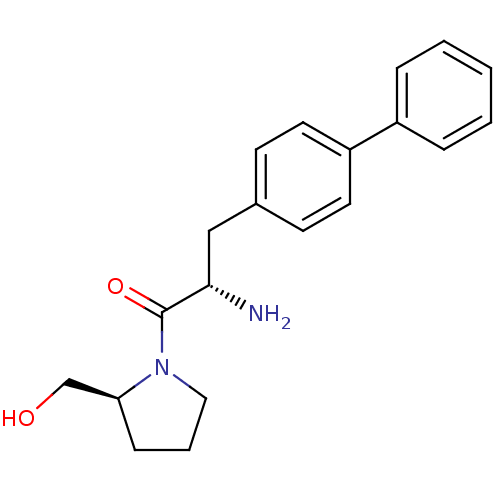
Affinity DataKi: 3.10nM ΔG°: -48.6kJ/molepH: 7.4 T: 2°CAssay Description:The progress of DPPIV inhibition by compounds was measured by recording the liberation of free pNA at 405 nm. IC50 was determined from the slope of r...More data for this Ligand-Target Pair
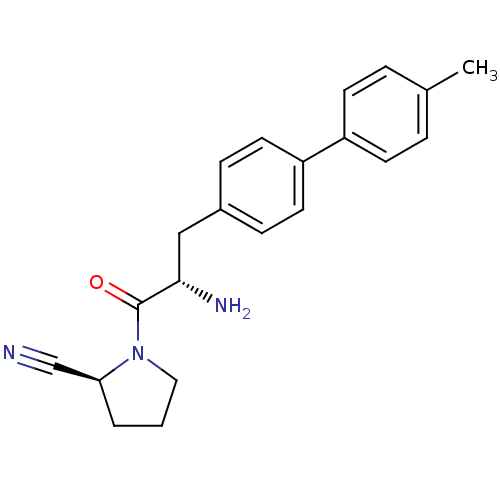
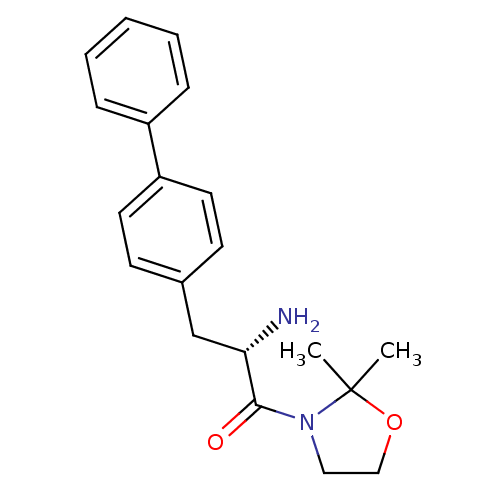
Affinity DataKi: 5.30nM ΔG°: -47.2kJ/molepH: 7.4 T: 2°CAssay Description:The progress of DPPIV inhibition by compounds was measured by recording the liberation of free pNA at 405 nm. IC50 was determined from the slope of r...More data for this Ligand-Target Pair
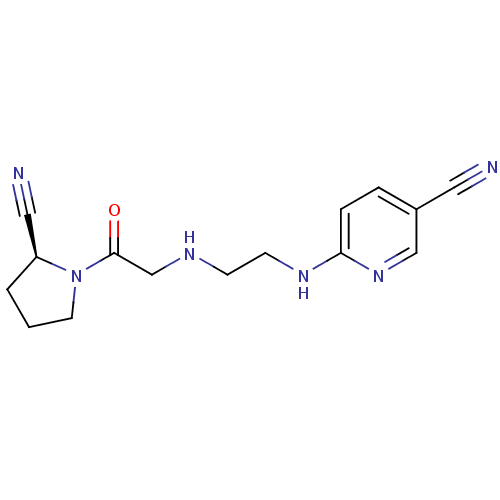
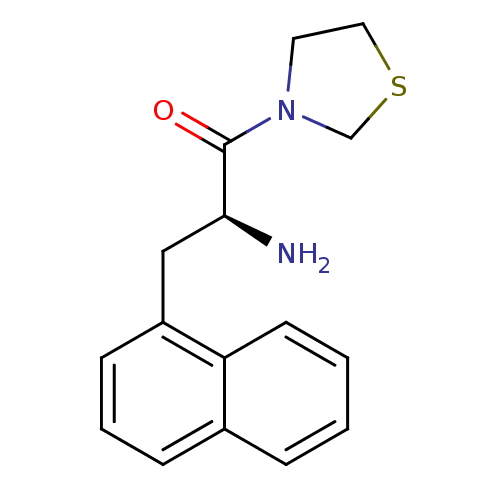
Affinity DataKi: 13nM ΔG°: -45.0kJ/molepH: 7.4 T: 2°CAssay Description:The progress of DPPIV inhibition by compounds was measured by recording the liberation of free pNA at 405 nm. IC50 was determined from the slope of r...More data for this Ligand-Target Pair
Affinity DataKi: 20nM ΔG°: -43.9kJ/molepH: 7.4 T: 2°CAssay Description:The progress of DPPIV inhibition by compounds was measured by recording the liberation of free pNA at 405 nm. IC50 was determined from the slope of r...More data for this Ligand-Target Pair
Affinity DataKi: 26nM ΔG°: -43.3kJ/molepH: 7.4 T: 2°CAssay Description:The progress of DPPIV inhibition by compounds was measured by recording the liberation of free pNA at 405 nm. IC50 was determined from the slope of r...More data for this Ligand-Target Pair
Affinity DataKi: 27nM ΔG°: -43.2kJ/molepH: 7.4 T: 2°CAssay Description:The progress of DPPIV inhibition by compounds was measured by recording the liberation of free pNA at 405 nm. IC50 was determined from the slope of r...More data for this Ligand-Target Pair
Affinity DataKi: 34nM ΔG°: -42.6kJ/molepH: 7.4 T: 2°CAssay Description:The progress of DPPIV inhibition by compounds was measured by recording the liberation of free pNA at 405 nm. IC50 was determined from the slope of r...More data for this Ligand-Target Pair
Affinity DataKi: 36nM ΔG°: -42.5kJ/molepH: 7.4 T: 2°CAssay Description:The progress of DPPIV inhibition by compounds was measured by recording the liberation of free pNA at 405 nm. IC50 was determined from the slope of r...More data for this Ligand-Target Pair
Affinity DataKi: 63nM ΔG°: -41.1kJ/molepH: 7.4 T: 2°CAssay Description:The progress of DPPIV inhibition by compounds was measured by recording the liberation of free pNA at 405 nm. IC50 was determined from the slope of r...More data for this Ligand-Target Pair
Affinity DataKi: 96nM ΔG°: -40.1kJ/molepH: 7.4 T: 2°CAssay Description:The progress of DPPIV inhibition by compounds was measured by recording the liberation of free pNA at 405 nm. IC50 was determined from the slope of r...More data for this Ligand-Target Pair
Affinity DataKi: 160nM ΔG°: -38.8kJ/molepH: 7.4 T: 2°CAssay Description:The progress of DPPIV inhibition by compounds was measured by recording the liberation of free pNA at 405 nm. IC50 was determined from the slope of r...More data for this Ligand-Target Pair
Affinity DataKi: 166nM ΔG°: -38.7kJ/molepH: 7.4 T: 2°CAssay Description:The progress of DPPIV inhibition by compounds was measured by recording the liberation of free pNA at 405 nm. IC50 was determined from the slope of r...More data for this Ligand-Target Pair
Affinity DataKi: 170nM ΔG°: -38.6kJ/molepH: 7.4 T: 2°CAssay Description:The progress of DPPIV inhibition by compounds was measured by recording the liberation of free pNA at 405 nm. IC50 was determined from the slope of r...More data for this Ligand-Target Pair
Affinity DataKi: 310nM ΔG°: -37.1kJ/molepH: 7.4 T: 2°CAssay Description:The progress of DPPIV inhibition by compounds was measured by recording the liberation of free pNA at 405 nm. IC50 was determined from the slope of r...More data for this Ligand-Target Pair
Affinity DataKi: 355nM ΔG°: -36.8kJ/molepH: 7.4 T: 2°CAssay Description:The progress of DPPIV inhibition by compounds was measured by recording the liberation of free pNA at 405 nm. IC50 was determined from the slope of r...More data for this Ligand-Target Pair
Affinity DataKi: 360nM ΔG°: -36.8kJ/molepH: 7.4 T: 2°CAssay Description:The progress of DPPIV inhibition by compounds was measured by recording the liberation of free pNA at 405 nm. IC50 was determined from the slope of r...More data for this Ligand-Target Pair
Affinity DataKi: 470nM ΔG°: -36.1kJ/molepH: 7.4 T: 2°CAssay Description:The progress of DPPIV inhibition by compounds was measured by recording the liberation of free pNA at 405 nm. IC50 was determined from the slope of r...More data for this Ligand-Target Pair
Affinity DataKi: 980nM ΔG°: -34.3kJ/molepH: 7.4 T: 2°CAssay Description:The progress of DPPIV inhibition by compounds was measured by recording the liberation of free pNA at 405 nm. IC50 was determined from the slope of r...More data for this Ligand-Target Pair
Affinity DataKi: 1.16E+3nM ΔG°: -33.9kJ/molepH: 7.4 T: 2°CAssay Description:The progress of DPPIV inhibition by compounds was measured by recording the liberation of free pNA at 405 nm. IC50 was determined from the slope of r...More data for this Ligand-Target Pair
Affinity DataKi: 8.90E+3nM ΔG°: -28.8kJ/molepH: 7.4 T: 2°CAssay Description:The progress of DPPIV inhibition by compounds was measured by recording the liberation of free pNA at 405 nm. IC50 was determined from the slope of r...More data for this Ligand-Target Pair
Affinity DataKi: >1.28E+4nM ΔG°: >-27.9kJ/molepH: 7.4 T: 2°CAssay Description:The progress of DPPIV inhibition by compounds was measured by recording the liberation of free pNA at 405 nm. IC50 was determined from the slope of r...More data for this Ligand-Target Pair
Affinity DataKi: >1.28E+4nM ΔG°: >-27.9kJ/molepH: 7.4 T: 2°CAssay Description:The progress of DPPIV inhibition by compounds was measured by recording the liberation of free pNA at 405 nm. IC50 was determined from the slope of r...More data for this Ligand-Target Pair
Affinity DataKi: >1.28E+4nM ΔG°: >-27.9kJ/molepH: 7.4 T: 2°CAssay Description:The progress of DPPIV inhibition by compounds was measured by recording the liberation of free pNA at 405 nm. IC50 was determined from the slope of r...More data for this Ligand-Target Pair
Affinity DataKi: >1.28E+4nM ΔG°: >-27.9kJ/molepH: 7.4 T: 2°CAssay Description:The progress of DPPIV inhibition by compounds was measured by recording the liberation of free pNA at 405 nm. IC50 was determined from the slope of r...More data for this Ligand-Target Pair
Affinity DataKi: >1.28E+4nM ΔG°: >-27.9kJ/molepH: 7.4 T: 2°CAssay Description:The progress of DPPIV inhibition by compounds was measured by recording the liberation of free pNA at 405 nm. IC50 was determined from the slope of r...More data for this Ligand-Target Pair
TargetMacrophage colony-stimulating factor 1 receptor [538-678,753-922](Homo sapiens (Human))
Johnson & Johnson Pharmaceutical
Johnson & Johnson Pharmaceutical
Affinity DataIC50: 49nMpH: 7.5 T: 2°CAssay Description:The full-length cFMS cytoplasmic domain (FMS.538-972.6His) or chimera was incubated with compound in reaction buffer. Control reactions were run in e...More data for this Ligand-Target Pair
Affinity DataIC50: 57nMpH: 7.5 T: 2°CAssay Description:The full-length cFMS cytoplasmic domain (FMS.538-972.6His) or chimera was incubated with compound in reaction buffer. Control reactions were run in e...More data for this Ligand-Target Pair
TargetMacrophage colony-stimulating factor 1 receptor [538-678,753-922](Human Sapiens (Human))
Johnson & Johnson Pharmaceutical
Johnson & Johnson Pharmaceutical
Affinity DataIC50: 57nMpH: 7.5 T: 2°CAssay Description:The full-length cFMS cytoplasmic domain (FMS.538-972.6His) or chimera was incubated with compound in reaction buffer. Control reactions were run in e...More data for this Ligand-Target Pair
Affinity DataIC50: 1.20E+3nMpH: 7.5 T: 2°CAssay Description:The full-length cFMS cytoplasmic domain (FMS.538-972.6His) or chimera was incubated with compound in reaction buffer. Control reactions were run in e...More data for this Ligand-Target Pair
Affinity DataIC50: 1.20E+3nMpH: 7.5 T: 2°CAssay Description:The full-length cFMS cytoplasmic domain (FMS.538-972.6His) or chimera was incubated with compound in reaction buffer. Control reactions were run in e...More data for this Ligand-Target Pair
Ligand Info
TargetMacrophage colony-stimulating factor 1 receptor [538-678,753-922](Human Sapiens (Human))
Johnson & Johnson Pharmaceutical
Johnson & Johnson Pharmaceutical
Affinity DataIC50: 1.20E+3nMpH: 7.5 T: 2°CAssay Description:The full-length cFMS cytoplasmic domain (FMS.538-972.6His) or chimera was incubated with compound in reaction buffer. Control reactions were run in e...More data for this Ligand-Target Pair
Ligand Info
TargetMacrophage colony-stimulating factor 1 receptor [538-678,753-922](Homo sapiens (Human))
Johnson & Johnson Pharmaceutical
Johnson & Johnson Pharmaceutical
Affinity DataIC50: 1.30E+3nMpH: 7.5 T: 2°CAssay Description:The full-length cFMS cytoplasmic domain (FMS.538-972.6His) or chimera was incubated with compound in reaction buffer. Control reactions were run in e...More data for this Ligand-Target Pair
TargetMacrophage colony-stimulating factor 1 receptor [538-678,753-922](Homo sapiens (Human))
Johnson & Johnson Pharmaceutical
Johnson & Johnson Pharmaceutical
Affinity DataIC50: 1.30E+3nMpH: 7.5 T: 2°CAssay Description:The full-length cFMS cytoplasmic domain (FMS.538-972.6His) or chimera was incubated with compound in reaction buffer. Control reactions were run in e...More data for this Ligand-Target Pair
Ligand Info
TargetMacrophage colony-stimulating factor 1 receptor [538-678,753-922](Human Sapiens (Human))
Johnson & Johnson Pharmaceutical
Johnson & Johnson Pharmaceutical
Affinity DataIC50: 1.40E+3nMpH: 7.5 T: 2°CAssay Description:The full-length cFMS cytoplasmic domain (FMS.538-972.6His) or chimera was incubated with compound in reaction buffer. Control reactions were run in e...More data for this Ligand-Target Pair
Affinity DataIC50: 2.90E+3nMpH: 7.5 T: 2°CAssay Description:The full-length cFMS cytoplasmic domain (FMS.538-972.6His) or chimera was incubated with compound in reaction buffer. Control reactions were run in e...More data for this Ligand-Target Pair
TargetMacrophage colony-stimulating factor 1 receptor [538-678,753-922](Human Sapiens (Human))
Johnson & Johnson Pharmaceutical
Johnson & Johnson Pharmaceutical
Affinity DataIC50: 3.00E+3nMpH: 7.5 T: 2°CAssay Description:The full-length cFMS cytoplasmic domain (FMS.538-972.6His) or chimera was incubated with compound in reaction buffer. Control reactions were run in e...More data for this Ligand-Target Pair
TargetMacrophage colony-stimulating factor 1 receptor [538-678,753-922](Homo sapiens (Human))
Johnson & Johnson Pharmaceutical
Johnson & Johnson Pharmaceutical
Affinity DataIC50: 3.30E+3nMpH: 7.5 T: 2°CAssay Description:The full-length cFMS cytoplasmic domain (FMS.538-972.6His) or chimera was incubated with compound in reaction buffer. Control reactions were run in e...More data for this Ligand-Target Pair
TargetMacrophage colony-stimulating factor 1 receptor [538-678,753-922](Human Sapiens (Human))
Johnson & Johnson Pharmaceutical
Johnson & Johnson Pharmaceutical
Affinity DataIC50: 3.90E+3nMpH: 7.5 T: 2°CAssay Description:The full-length cFMS cytoplasmic domain (FMS.538-972.6His) or chimera was incubated with compound in reaction buffer. Control reactions were run in e...More data for this Ligand-Target Pair
TargetMacrophage colony-stimulating factor 1 receptor [538-678,753-922](Human Sapiens (Human))
Johnson & Johnson Pharmaceutical
Johnson & Johnson Pharmaceutical
Affinity DataIC50: 4.20E+3nMpH: 7.5 T: 2°CAssay Description:The full-length cFMS cytoplasmic domain (FMS.538-972.6His) or chimera was incubated with compound in reaction buffer. Control reactions were run in e...More data for this Ligand-Target Pair
TargetMacrophage colony-stimulating factor 1 receptor [538-678,753-922](Homo sapiens (Human))
Johnson & Johnson Pharmaceutical
Johnson & Johnson Pharmaceutical
Affinity DataIC50: 4.50E+3nMpH: 7.5 T: 2°CAssay Description:The full-length cFMS cytoplasmic domain (FMS.538-972.6His) or chimera was incubated with compound in reaction buffer. Control reactions were run in e...More data for this Ligand-Target Pair
TargetMacrophage colony-stimulating factor 1 receptor [538-678,753-922](Homo sapiens (Human))
Johnson & Johnson Pharmaceutical
Johnson & Johnson Pharmaceutical
Affinity DataIC50: 4.60E+3nMpH: 7.5 T: 2°CAssay Description:The full-length cFMS cytoplasmic domain (FMS.538-972.6His) or chimera was incubated with compound in reaction buffer. Control reactions were run in e...More data for this Ligand-Target Pair
Affinity DataIC50: 4.70E+3nMpH: 7.5 T: 2°CAssay Description:The full-length cFMS cytoplasmic domain (FMS.538-972.6His) or chimera was incubated with compound in reaction buffer. Control reactions were run in e...More data for this Ligand-Target Pair
Affinity DataIC50: 4.80E+3nMpH: 7.5 T: 2°CAssay Description:The full-length cFMS cytoplasmic domain (FMS.538-972.6His) or chimera was incubated with compound in reaction buffer. Control reactions were run in e...More data for this Ligand-Target Pair
TargetMacrophage colony-stimulating factor 1 receptor [538-678,753-922](Homo sapiens (Human))
Johnson & Johnson Pharmaceutical
Johnson & Johnson Pharmaceutical
Affinity DataIC50: 8.30E+3nMpH: 7.5 T: 2°CAssay Description:The full-length cFMS cytoplasmic domain (FMS.538-972.6His) or chimera was incubated with compound in reaction buffer. Control reactions were run in e...More data for this Ligand-Target Pair
TargetMacrophage colony-stimulating factor 1 receptor [538-678,753-922](Human Sapiens (Human))
Johnson & Johnson Pharmaceutical
Johnson & Johnson Pharmaceutical
Affinity DataIC50: 9.40E+3nMpH: 7.5 T: 2°CAssay Description:The full-length cFMS cytoplasmic domain (FMS.538-972.6His) or chimera was incubated with compound in reaction buffer. Control reactions were run in e...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMpH: 7.5 T: 2°CAssay Description:The full-length cFMS cytoplasmic domain (FMS.538-972.6His) or chimera was incubated with compound in reaction buffer. Control reactions were run in e...More data for this Ligand-Target Pair
Affinity DataEC50: 1.07E+4nMAssay Description:The DISCOVERX PATHHUNTER� β-Arrestin assay is used to measure β-arrestin-2 activity. The technology is based on Enzyme Fragment Complementa...More data for this Ligand-Target Pair
Affinity DataEC50: 67nMAssay Description:G-protein signaling was measured via second messenger cAMP modulation. Detection of cAMP modulation was accomplished in the PathHunter human OPRM1 (&...More data for this Ligand-Target Pair
Affinity DataEC50: >1.00E+5nMAssay Description:The DISCOVERX PATHHUNTER� β-Arrestin assay is used to measure β-arrestin-2 activity. The technology is based on Enzyme Fragment Complementa...More data for this Ligand-Target Pair