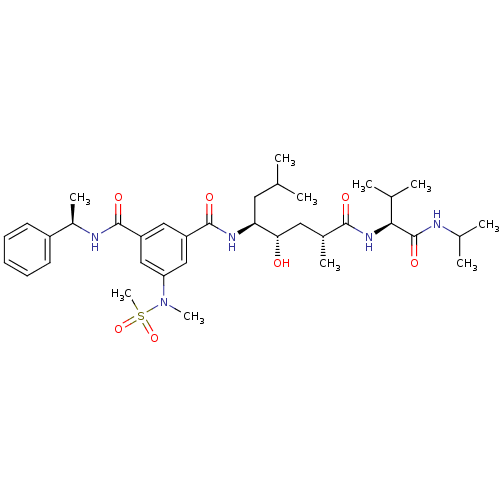
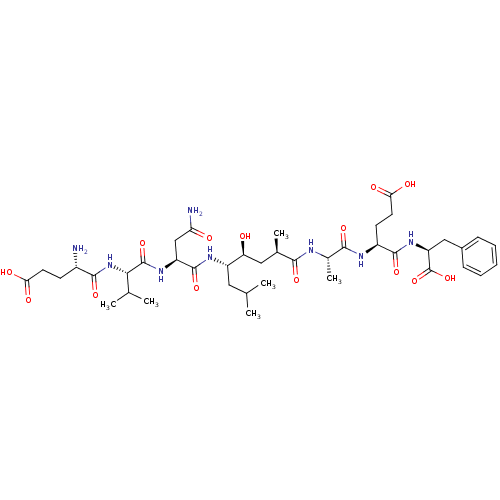
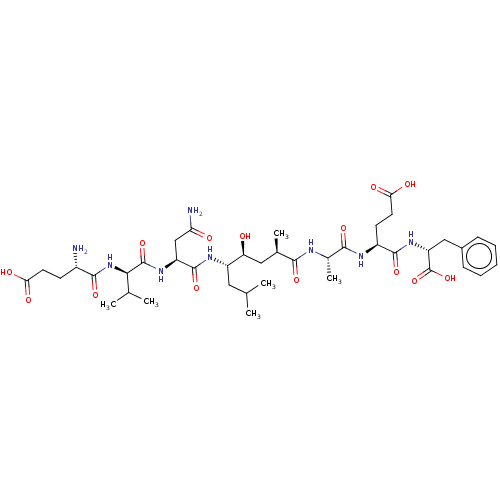
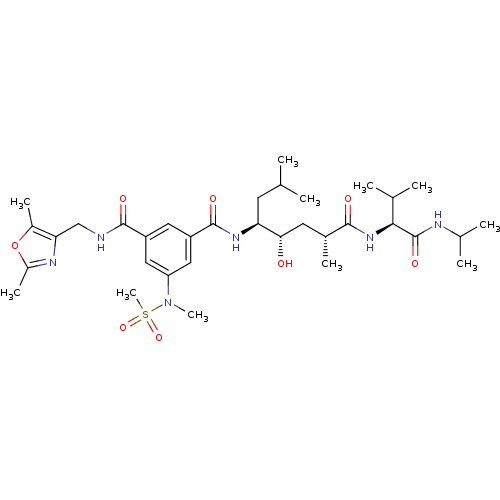
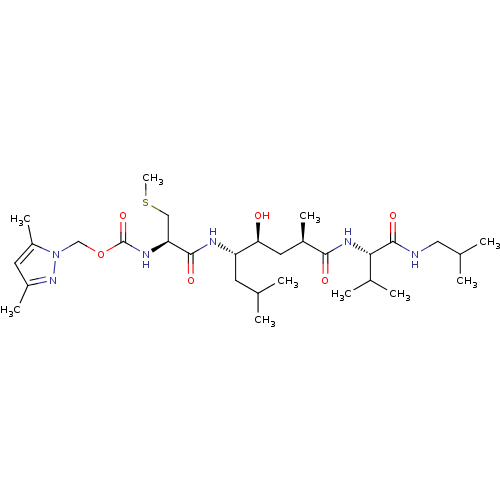
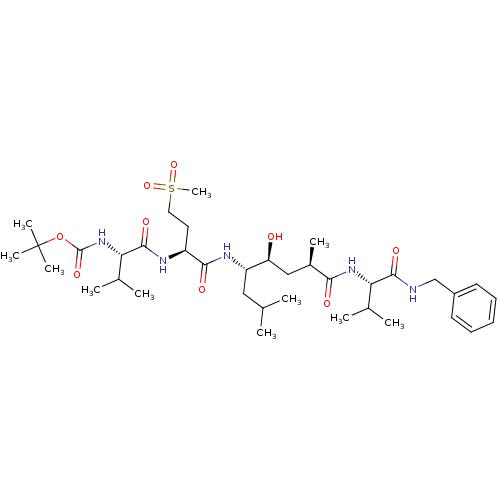
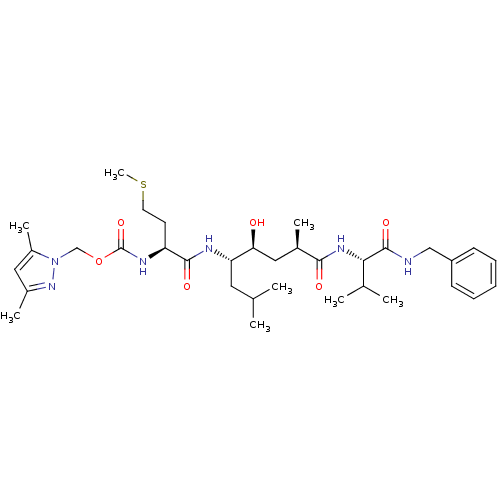
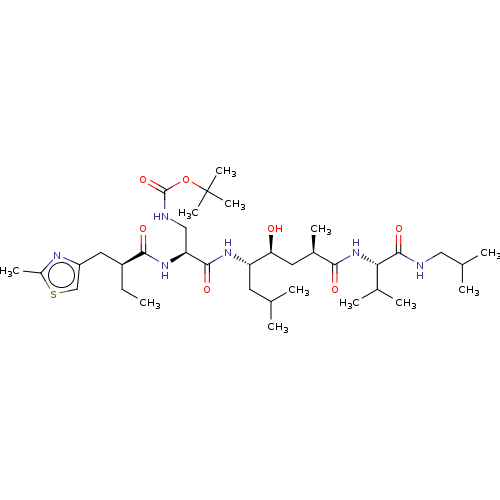
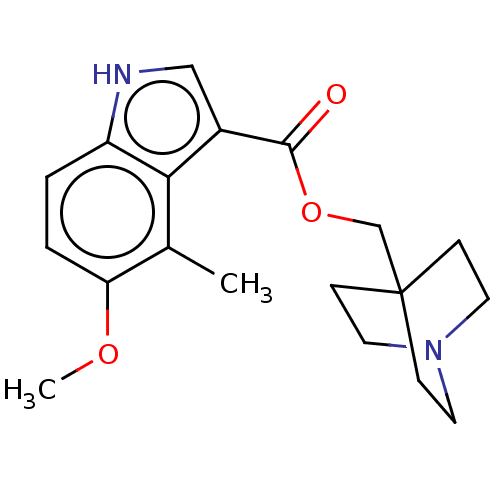
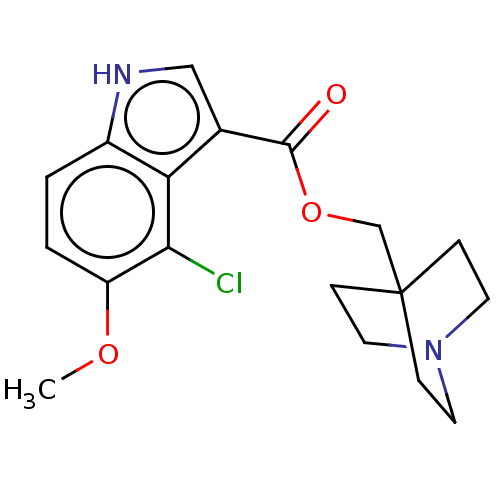
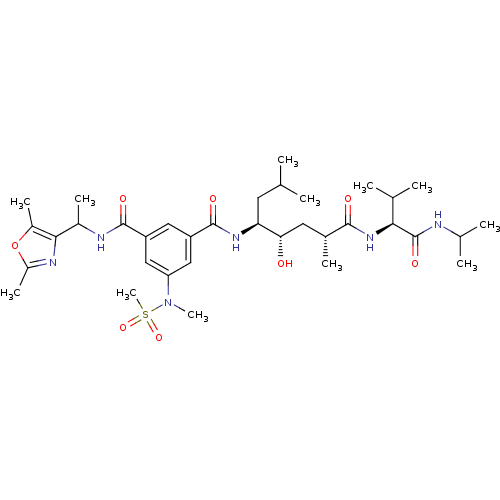
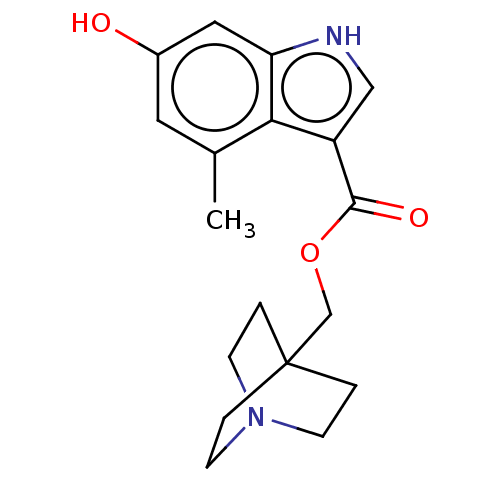
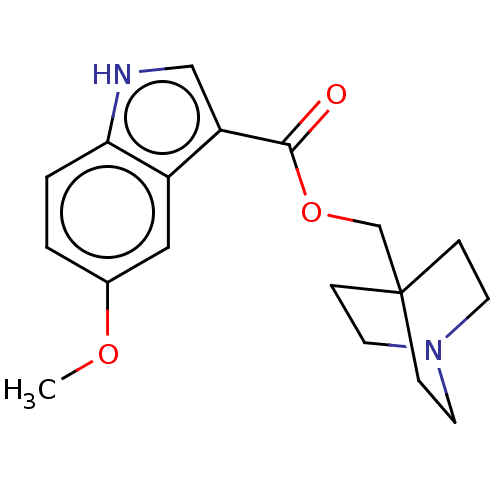
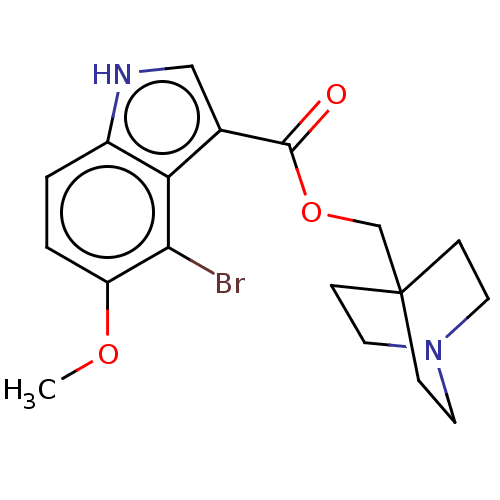
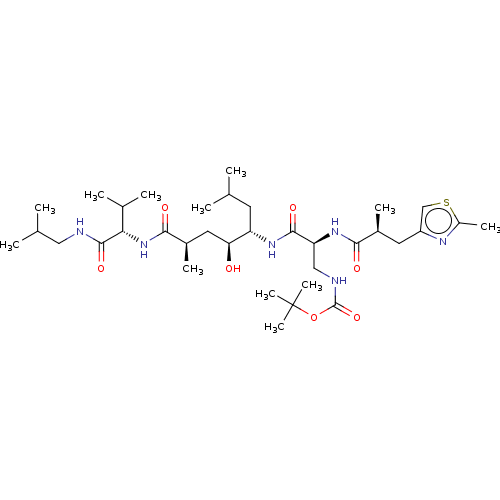
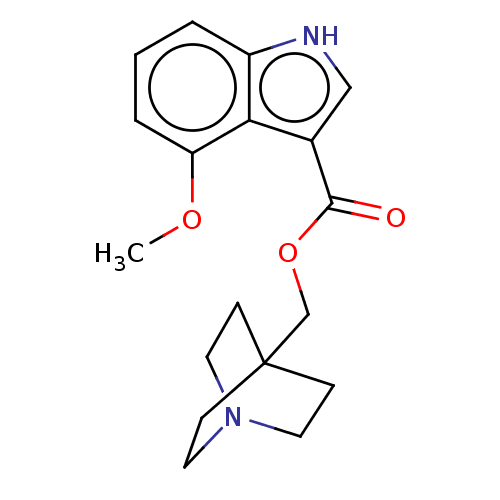
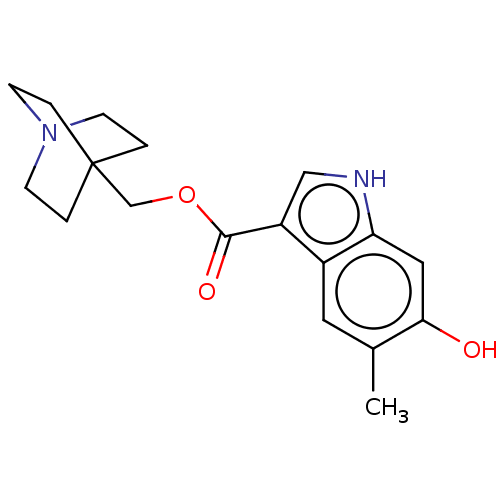
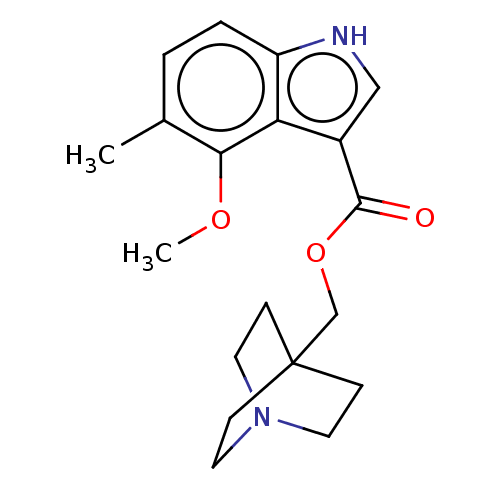
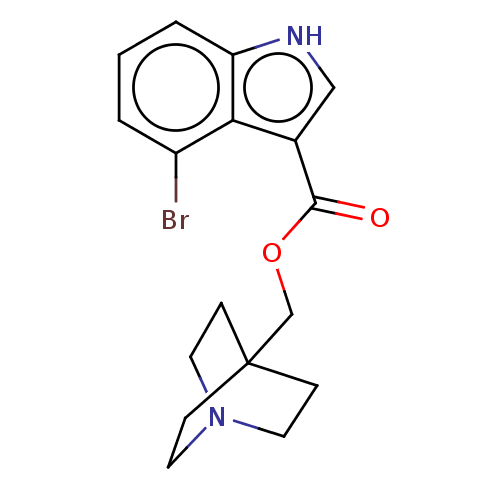
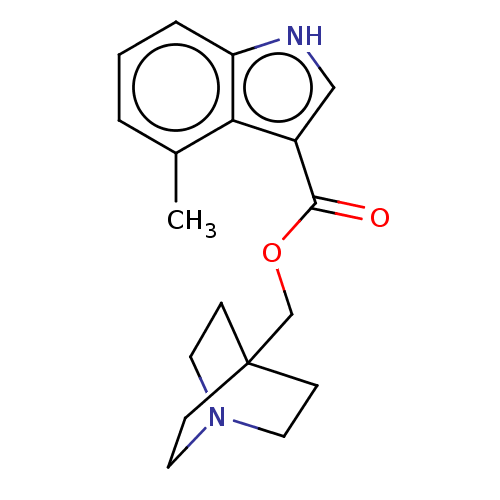
Affinity DataKi: 0.120nM ΔG°: -58.9kJ/molepH: 4.5 T: 2°CAssay Description:Enzyme activities were assayed by monitoring the hydrolysis of substrate in the presence or absence of inhibitor compounds. The hydrolysis was record...More data for this Ligand-Target Pair
Affinity DataKi: 0.25nMAssay Description:Inhibition of recombinant BACE1 (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 0.300nMAssay Description:Inhibition of recombinant BACE1 (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 0.300nM ΔG°: -56.5kJ/molepH: 4.5 T: 2°CAssay Description:Enzyme activities were assayed by monitoring the hydrolysis of substrate in the presence or absence of inhibitor compounds. The hydrolysis was record...More data for this Ligand-Target Pair
Affinity DataKi: 1.10nM ΔG°: -53.2kJ/molepH: 4.5 T: 2°CAssay Description:Enzyme activities were assayed by monitoring the hydrolysis of substrate in the presence or absence of inhibitor compounds. The hydrolysis was record...More data for this Ligand-Target Pair
Affinity DataKi: 1.20nM ΔG°: -53.0kJ/molepH: 4.5 T: 2°CAssay Description:Enzyme activities were assayed by monitoring the hydrolysis of substrate in the presence or absence of inhibitor compounds. The hydrolysis was record...More data for this Ligand-Target Pair
Affinity DataKi: 1.30nMAssay Description:Inhibition of Cathepsin D (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 1.60nM ΔG°: -52.2kJ/molepH: 4.5 T: 2°CAssay Description:Enzyme activities were assayed by monitoring the hydrolysis of substrate in the presence or absence of inhibitor compounds. The hydrolysis was record...More data for this Ligand-Target Pair
Affinity DataKi: 1.60nM ΔG°: -52.2kJ/molepH: 4.5 T: 2°CAssay Description:Enzyme activities were assayed by monitoring the hydrolysis of substrate in the presence or absence of inhibitor compounds. The hydrolysis was record...More data for this Ligand-Target Pair
Affinity DataKi: 1.60nMAssay Description:Inhibition of recombinant BACE1 (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 1.80nM ΔG°: -51.9kJ/molepH: 4.5 T: 2°CAssay Description:Enzyme activities were assayed by monitoring the hydrolysis of substrate in the presence or absence of inhibitor compounds. The hydrolysis was record...More data for this Ligand-Target Pair
Affinity DataKi: 2.5nM ΔG°: -51.1kJ/molepH: 4.5 T: 2°CAssay Description:Enzyme activities were assayed by monitoring the hydrolysis of substrate in the presence or absence of inhibitor compounds. The hydrolysis was record...More data for this Ligand-Target Pair
Affinity DataKi: 2.5nM ΔG°: -51.1kJ/molepH: 4.5 T: 2°CAssay Description:Enzyme activities were assayed by monitoring the hydrolysis of substrate in the presence or absence of inhibitor compounds. The hydrolysis was record...More data for this Ligand-Target Pair
Affinity DataKi: 4.40nM ΔG°: -49.6kJ/molepH: 4.5 T: 2°CAssay Description:Enzyme activities were assayed by monitoring the hydrolysis of substrate in the presence or absence of inhibitor compounds. The hydrolysis was record...More data for this Ligand-Target Pair
Affinity DataKi: 5.90nM ΔG°: -48.9kJ/molepH: 4.5 T: 2°CAssay Description:Enzyme activities were assayed by monitoring the hydrolysis of substrate in the presence or absence of inhibitor compounds. The hydrolysis was record...More data for this Ligand-Target Pair
Affinity DataKi: 8nM ΔG°: -48.1kJ/molepH: 4.5 T: 2°CAssay Description:Enzyme activities were assayed by monitoring the hydrolysis of substrate in the presence or absence of inhibitor compounds. The hydrolysis was record...More data for this Ligand-Target Pair
Affinity DataKi: 9.40nM ΔG°: -47.7kJ/molepH: 4.5 T: 2°CAssay Description:Enzyme activities were assayed by monitoring the hydrolysis of substrate in the presence or absence of inhibitor compounds. The hydrolysis was record...More data for this Ligand-Target Pair
Affinity DataKi: 11nMAssay Description:Inhibition of recombinant BACE1 (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 12.2nM ΔG°: -47.0kJ/mole IC50: 14.1nMpH: 7.4 T: 2°CAssay Description:Protocol 1: The assay was conducted following the literature reference Meyer E. M., et al., Analysis of 3-(4-Hydroxy, 2-Methoxybenzylidene) Anabasein...More data for this Ligand-Target Pair
Affinity DataKi: 14nM ΔG°: -46.6kJ/molepH: 4.5 T: 2°CAssay Description:Enzyme activities were assayed by monitoring the hydrolysis of substrate in the presence or absence of inhibitor compounds. The hydrolysis was record...More data for this Ligand-Target Pair
Affinity DataKi: 14.2nM ΔG°: -46.6kJ/molepH: 4.5 T: 2°CAssay Description:Enzyme activities were assayed by monitoring the hydrolysis of substrate in the presence or absence of inhibitor compounds. The hydrolysis was record...More data for this Ligand-Target Pair
Affinity DataKi: 15nM ΔG°: -46.5kJ/molepH: 4.5 T: 2°CAssay Description:Enzyme activities were assayed by monitoring the hydrolysis of substrate in the presence or absence of inhibitor compounds. The hydrolysis was record...More data for this Ligand-Target Pair
Affinity DataKi: 17nMAssay Description:Inhibition of recombinant BACE1 (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 20.4nM ΔG°: -45.7kJ/mole IC50: 23.6nMpH: 7.4 T: 2°CAssay Description:Protocol 1: The assay was conducted following the literature reference Meyer E. M., et al., Analysis of 3-(4-Hydroxy, 2-Methoxybenzylidene) Anabasein...More data for this Ligand-Target Pair
Affinity DataKi: 20.6nM ΔG°: -45.6kJ/mole IC50: 23.9nMpH: 7.4 T: 2°CAssay Description:Protocol 1: The assay was conducted following the literature reference Meyer E. M., et al., Analysis of 3-(4-Hydroxy, 2-Methoxybenzylidene) Anabasein...More data for this Ligand-Target Pair
Affinity DataKi: 22.9nM ΔG°: -45.4kJ/mole IC50: 26.6nMpH: 7.4 T: 2°CAssay Description:Protocol 1: The assay was conducted following the literature reference Meyer E. M., et al., Analysis of 3-(4-Hydroxy, 2-Methoxybenzylidene) Anabasein...More data for this Ligand-Target Pair
Affinity DataKi: 25nM ΔG°: -45.1kJ/molepH: 4.5 T: 2°CAssay Description:Enzyme activities were assayed by monitoring the hydrolysis of substrate in the presence or absence of inhibitor compounds. The hydrolysis was record...More data for this Ligand-Target Pair
Affinity DataKi: 25.1nM ΔG°: -45.1kJ/molepH: 4.5 T: 2°CAssay Description:Enzyme activities were assayed by monitoring the hydrolysis of substrate in the presence or absence of inhibitor compounds. The hydrolysis was record...More data for this Ligand-Target Pair
Affinity DataKi: 26.6nM ΔG°: -45.0kJ/molepH: 4.5 T: 2°CAssay Description:Enzyme activities were assayed by monitoring the hydrolysis of substrate in the presence or absence of inhibitor compounds. The hydrolysis was record...More data for this Ligand-Target Pair
Affinity DataKi: 27nM ΔG°: -44.9kJ/molepH: 4.5 T: 2°CAssay Description:Enzyme activities were assayed by monitoring the hydrolysis of substrate in the presence or absence of inhibitor compounds. The hydrolysis was record...More data for this Ligand-Target Pair
Affinity DataKi: 27.6nM ΔG°: -44.9kJ/mole IC50: 31.8nMpH: 7.4 T: 2°CAssay Description:Protocol 1: The assay was conducted following the literature reference Meyer E. M., et al., Analysis of 3-(4-Hydroxy, 2-Methoxybenzylidene) Anabasein...More data for this Ligand-Target Pair
Affinity DataKi: 27.9nM ΔG°: -44.9kJ/mole IC50: 31.8nMpH: 7.4 T: 2°CAssay Description:Protocol 1: The assay was conducted following the literature reference Meyer E. M., et al., Analysis of 3-(4-Hydroxy, 2-Methoxybenzylidene) Anabasein...More data for this Ligand-Target Pair
Affinity DataKi: 28.3nM ΔG°: -44.8kJ/mole IC50: 32.6nMpH: 7.4 T: 2°CAssay Description:Protocol 1: The assay was conducted following the literature reference Meyer E. M., et al., Analysis of 3-(4-Hydroxy, 2-Methoxybenzylidene) Anabasein...More data for this Ligand-Target Pair
Affinity DataKi: 28.4nM ΔG°: -44.8kJ/molepH: 4.5 T: 2°CAssay Description:Enzyme activities were assayed by monitoring the hydrolysis of substrate in the presence or absence of inhibitor compounds. The hydrolysis was record...More data for this Ligand-Target Pair
Affinity DataKi: 28.5nM ΔG°: -44.8kJ/mole IC50: 33.3nMpH: 7.4 T: 2°CAssay Description:Protocol 1: The assay was conducted following the literature reference Meyer E. M., et al., Analysis of 3-(4-Hydroxy, 2-Methoxybenzylidene) Anabasein...More data for this Ligand-Target Pair
Affinity DataKi: 28.6nM ΔG°: -44.8kJ/mole IC50: 33.4nMpH: 7.4 T: 2°CAssay Description:Protocol 1: The assay was conducted following the literature reference Meyer E. M., et al., Analysis of 3-(4-Hydroxy, 2-Methoxybenzylidene) Anabasein...More data for this Ligand-Target Pair
Affinity DataKi: 29nMAssay Description:Inhibition of recombinant BACE1 (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 31nM ΔG°: -44.6kJ/molepH: 4.5 T: 2°CAssay Description:Enzyme activities were assayed by monitoring the hydrolysis of substrate in the presence or absence of inhibitor compounds. The hydrolysis was record...More data for this Ligand-Target Pair
Affinity DataKi: 32nMAssay Description:Inhibition of recombinant BACE1 (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 36nM ΔG°: -44.2kJ/molepH: 4.5 T: 2°CAssay Description:Enzyme activities were assayed by monitoring the hydrolysis of substrate in the presence or absence of inhibitor compounds. The hydrolysis was record...More data for this Ligand-Target Pair
Affinity DataKi: 40nMAssay Description:Inhibition of recombinant BACE1 (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 41nM ΔG°: -43.9kJ/molepH: 4.5 T: 2°CAssay Description:Enzyme activities were assayed by monitoring the hydrolysis of substrate in the presence or absence of inhibitor compounds. The hydrolysis was record...More data for this Ligand-Target Pair
Affinity DataKi: 42.3nM ΔG°: -43.8kJ/mole IC50: 48.9nMpH: 7.4 T: 2°CAssay Description:Protocol 1: The assay was conducted following the literature reference Meyer E. M., et al., Analysis of 3-(4-Hydroxy, 2-Methoxybenzylidene) Anabasein...More data for this Ligand-Target Pair
Affinity DataKi: 43.2nM ΔG°: -43.7kJ/mole IC50: 49.6nMpH: 7.4 T: 2°CAssay Description:Protocol 1: The assay was conducted following the literature reference Meyer E. M., et al., Analysis of 3-(4-Hydroxy, 2-Methoxybenzylidene) Anabasein...More data for this Ligand-Target Pair
Affinity DataKi: 44.8nM ΔG°: -43.6kJ/mole IC50: 52.3nMpH: 7.4 T: 2°CAssay Description:Protocol 1: The assay was conducted following the literature reference Meyer E. M., et al., Analysis of 3-(4-Hydroxy, 2-Methoxybenzylidene) Anabasein...More data for this Ligand-Target Pair
Affinity DataKi: 45nM ΔG°: -43.6kJ/mole IC50: 52.5nMpH: 7.4 T: 2°CAssay Description:Protocol 1: The assay was conducted following the literature reference Meyer E. M., et al., Analysis of 3-(4-Hydroxy, 2-Methoxybenzylidene) Anabasein...More data for this Ligand-Target Pair
Affinity DataKi: 46nM ΔG°: -43.6kJ/mole IC50: 53.6nMpH: 7.4 T: 2°CAssay Description:Protocol 1: The assay was conducted following the literature reference Meyer E. M., et al., Analysis of 3-(4-Hydroxy, 2-Methoxybenzylidene) Anabasein...More data for this Ligand-Target Pair
Affinity DataKi: 50.1nM ΔG°: -43.3kJ/molepH: 4.5 T: 2°CAssay Description:Enzyme activities were assayed by monitoring the hydrolysis of substrate in the presence or absence of inhibitor compounds. The hydrolysis was record...More data for this Ligand-Target Pair
Affinity DataKi: 50.4nM ΔG°: -43.3kJ/mole IC50: 58.1nMpH: 7.4 T: 2°CAssay Description:Protocol 1: The assay was conducted following the literature reference Meyer E. M., et al., Analysis of 3-(4-Hydroxy, 2-Methoxybenzylidene) Anabasein...More data for this Ligand-Target Pair
Affinity DataKi: 50.9nM ΔG°: -43.3kJ/mole IC50: 58.8nMpH: 7.4 T: 2°CAssay Description:Protocol 1: The assay was conducted following the literature reference Meyer E. M., et al., Analysis of 3-(4-Hydroxy, 2-Methoxybenzylidene) Anabasein...More data for this Ligand-Target Pair

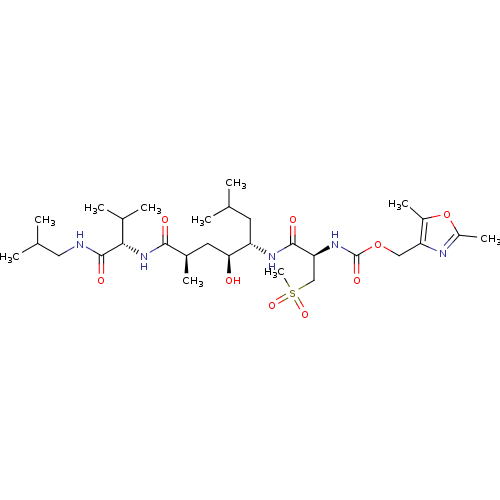
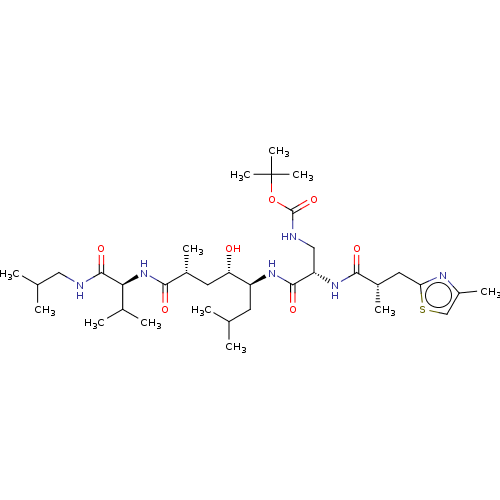
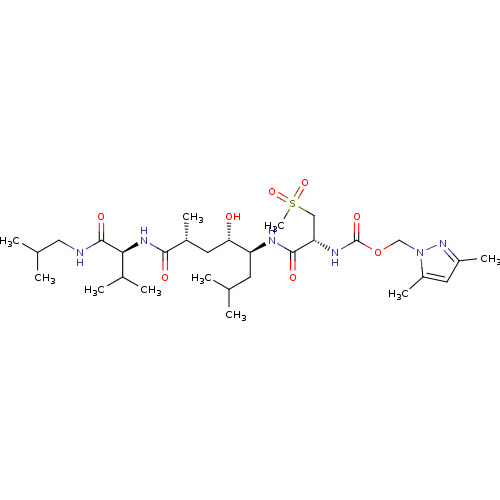
 3D Structure (crystal)
3D Structure (crystal)