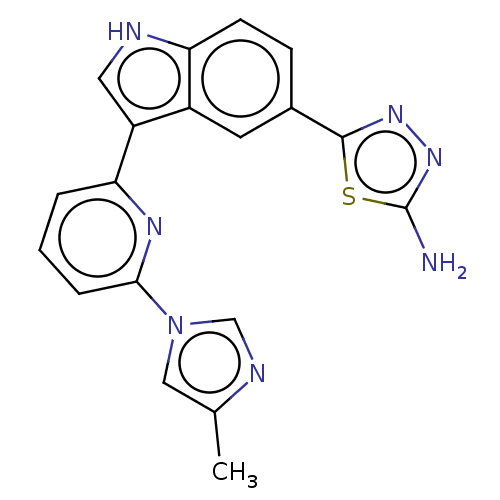
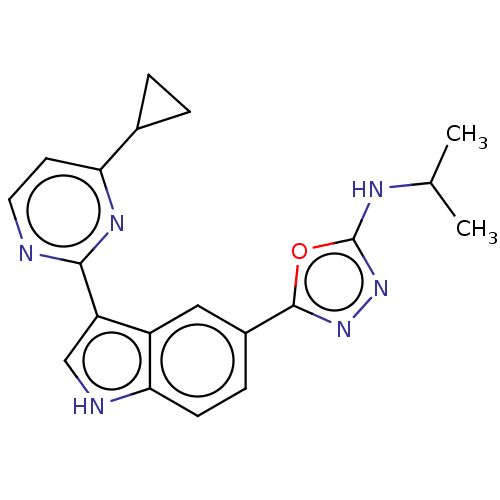
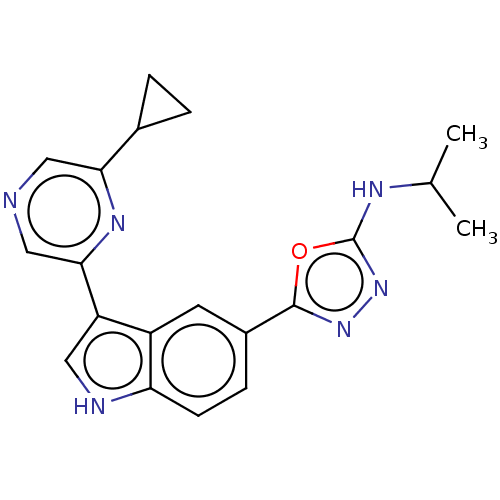
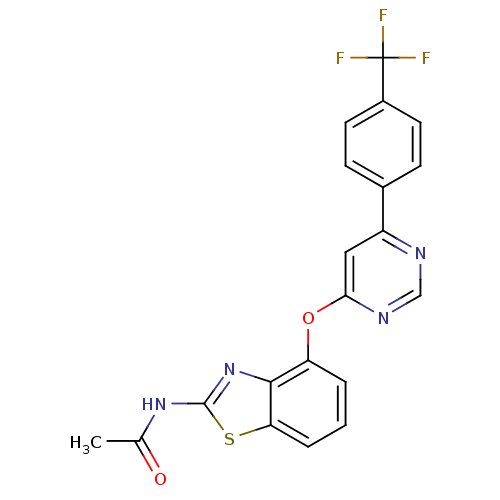
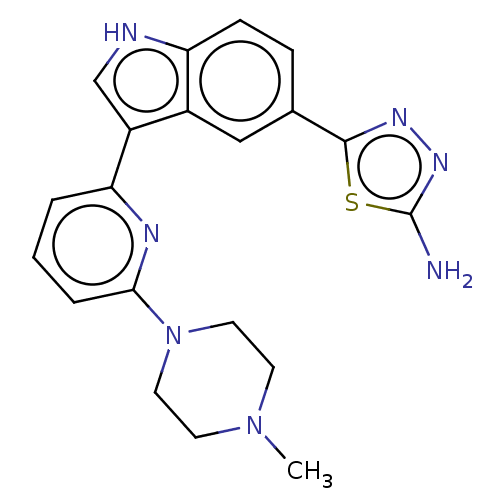
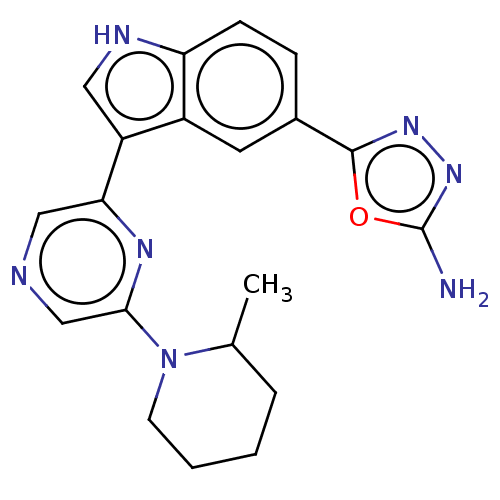
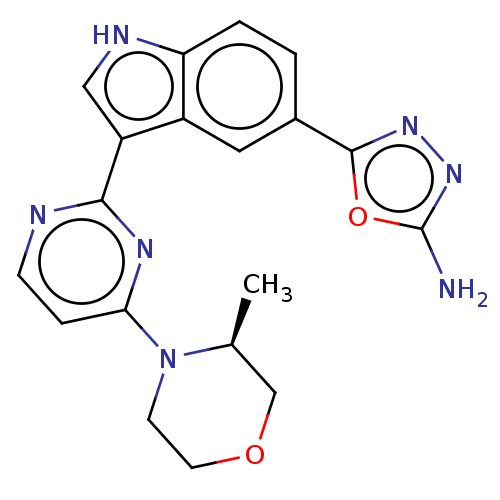
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Rattus norvegicus (rat))
Amgen
Curated by ChEMBL
Amgen
Curated by ChEMBL
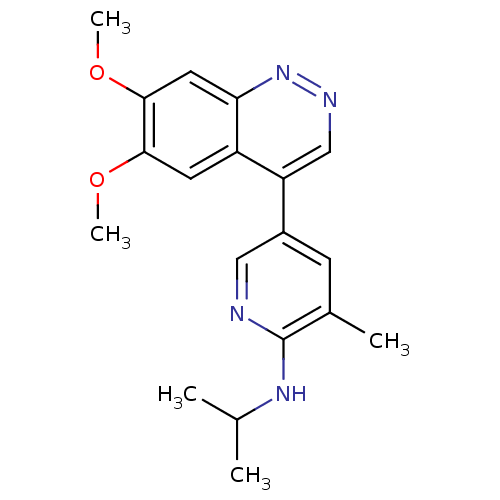
Affinity DataKi: 0.170nMAssay Description:Displacement of [3H]5-(6,7-dimethoxycinnolin-4-yl)-N-isopropyl-3-methylpyridin-2-amine from PDE10A in Sprague-Dawley rat striatumMore data for this Ligand-Target Pair
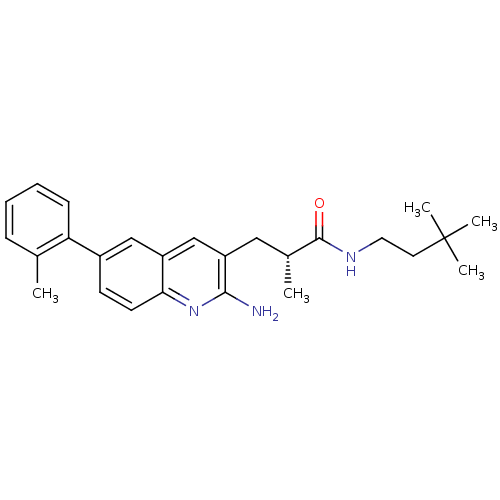
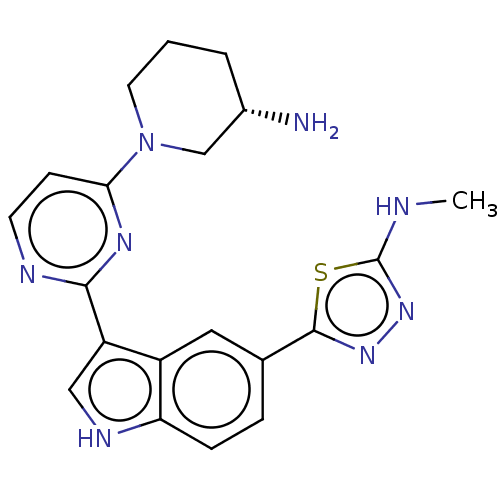
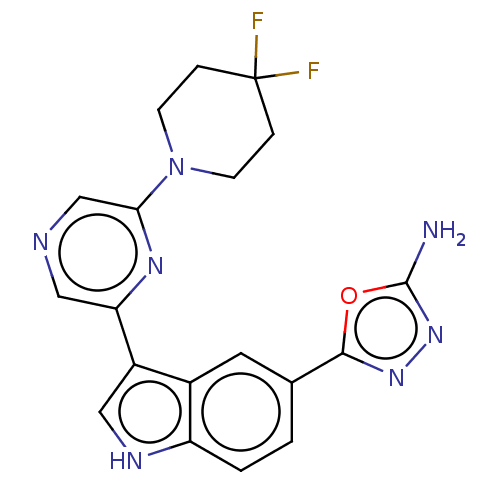
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Rattus norvegicus (rat))
Amgen
Curated by ChEMBL
Amgen
Curated by ChEMBL
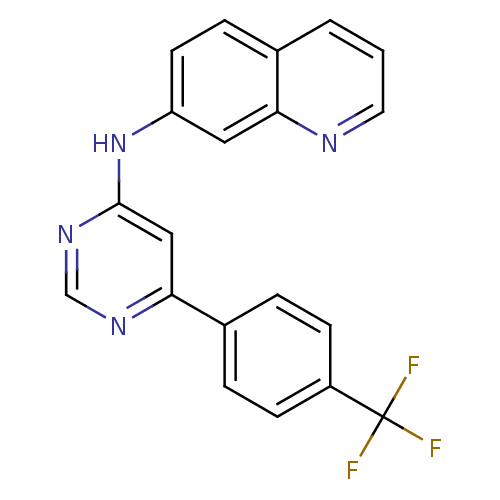
Affinity DataKi: 1.18nMAssay Description:Displacement of [3H]5-(6,7-dimethoxycinnolin-4-yl)-N-isopropyl-3-methylpyridin-2-amine from PDE10A in Sprague-Dawley rat striatumMore data for this Ligand-Target Pair
Affinity DataIC50: 0.270nMAssay Description:The assay uses purified enzyme interacting with biotinylated peptide substrate. HTRF is based on the proximity of europium cryptate (donor fluorophor...More data for this Ligand-Target Pair
Affinity DataIC50: 0.300nMAssay Description:TRPV1 activities were monitored as a function of cellular uptake of radioactive calcium (45Ca2+). The test compounds were preincubated with TRPV1 exp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.307nMAssay Description:The assay for the determination of Pim activity is based on the formation of phosphorylated biotinylated-BAD peptide at the Serine 112 residue (S112)...More data for this Ligand-Target Pair
Affinity DataIC50: 0.346nMAssay Description:The assay for the determination of Pim activity is based on the formation of phosphorylated biotinylated-BAD peptide at the Serine 112 residue (S112)...More data for this Ligand-Target Pair
Affinity DataIC50: 0.407nMAssay Description:The assay for the determination of Pim activity is based on the formation of phosphorylated biotinylated-BAD peptide at the Serine 112 residue (S112)...More data for this Ligand-Target Pair
Affinity DataIC50: 0.425nMAssay Description:The assay for the determination of Pim activity is based on the formation of phosphorylated biotinylated-BAD peptide at the Serine 112 residue (S112)...More data for this Ligand-Target Pair
Affinity DataIC50: 0.435nMAssay Description:The assay for the determination of Pim activity is based on the formation of phosphorylated biotinylated-BAD peptide at the Serine 112 residue (S112)...More data for this Ligand-Target Pair
Affinity DataIC50: 0.460nM EC50: 15nMpH: 7.0 T: 2°CAssay Description:The in vitro kinase activities of wild type or mutants were determined by measuring phosphorylation of biotinylated-MEK protein using Perkin-Elmer s ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.498nMAssay Description:The assay for the determination of Pim activity is based on the formation of phosphorylated biotinylated-BAD peptide at the Serine 112 residue (S112)...More data for this Ligand-Target Pair
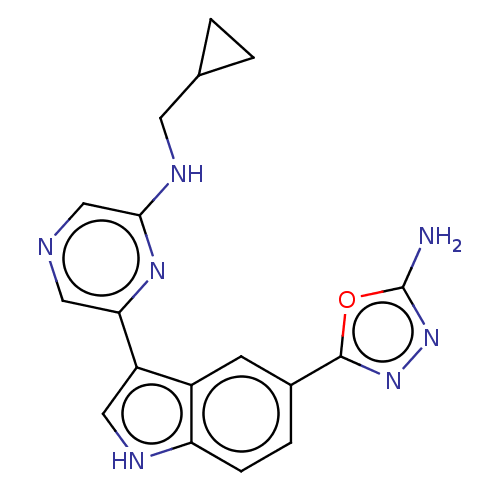
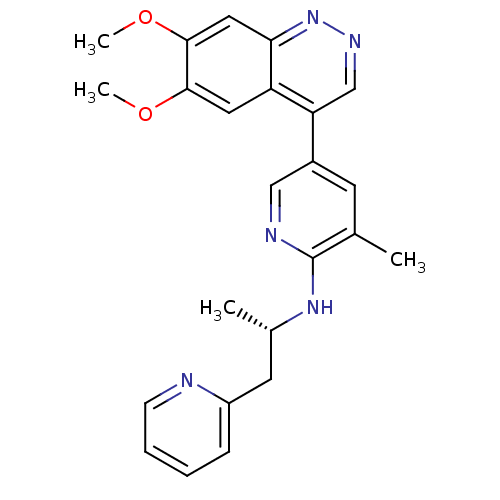
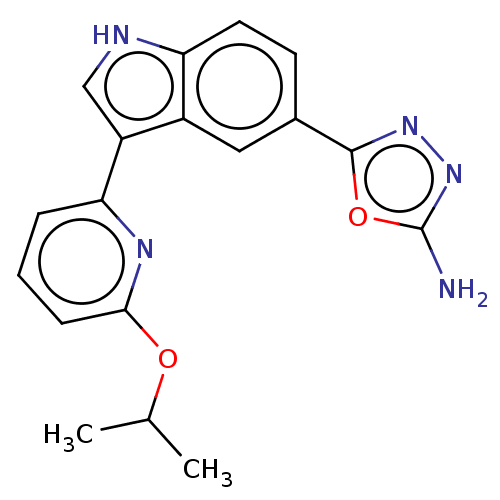
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Rattus norvegicus (rat))
Amgen
Curated by ChEMBL
Amgen
Curated by ChEMBL
Affinity DataIC50: 0.5nMAssay Description:Displacement of [3H]5-(6,7-dimethoxycinnolin-4-yl)-N-isopropyl-3-methylpyridin-2-amine from PDE10A in Sprague-Dawley rat striatumMore data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily V member 1(Rattus norvegicus (rat))
Amgen
Amgen
Affinity DataIC50: 0.5nMAssay Description:TRPV1 activities were monitored as a function of cellular uptake of radioactive calcium (45Ca2+). The test compounds were preincubated with TRPV1 exp...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily V member 1(Rattus norvegicus (rat))
Amgen
Amgen
Affinity DataIC50: 0.570nMAssay Description:TRPV1 activities were monitored as a function of cellular uptake of radioactive calcium (45Ca2+). The test compounds were preincubated with TRPV1 exp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.586nMAssay Description:The assay for the determination of Pim activity is based on the formation of phosphorylated biotinylated-BAD peptide at the Serine 112 residue (S112)...More data for this Ligand-Target Pair
Affinity DataIC50: 0.600nMpH: 7.4 T: 2°CAssay Description:TRPV1 activities were monitored as a function of cellular uptake of radioactive calcium (45Ca2+). The test compounds were preincubated with TRPV1 exp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.620nMAssay Description:TRPV1 activities were monitored as a function of cellular uptake of radioactive calcium (45Ca2+). The test compounds were preincubated with TRPV1 exp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.625nMAssay Description:The assay for the determination of Pim activity is based on the formation of phosphorylated biotinylated-BAD peptide at the Serine 112 residue (S112)...More data for this Ligand-Target Pair
Affinity DataIC50: 0.638nMAssay Description:The assay for the determination of Pim activity is based on the formation of phosphorylated biotinylated-BAD peptide at the Serine 112 residue (S112)...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily V member 1(Rattus norvegicus (rat))
Amgen
Amgen
Affinity DataIC50: 0.640nMpH: 7.4 T: 2°CAssay Description:TRPV1 activities were monitored as a function of cellular uptake of radioactive calcium (45Ca2+). The test compounds were preincubated with TRPV1 exp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.650nMAssay Description:Inhibition of human recombinant BACE1 after 60 mins by FRET assayMore data for this Ligand-Target Pair
Affinity DataIC50: 0.668nMAssay Description:The assay for the determination of Pim activity is based on the formation of phosphorylated biotinylated-BAD peptide at the Serine 112 residue (S112)...More data for this Ligand-Target Pair
Affinity DataIC50: 0.700nMAssay Description:Inhibition of human recombinant BACE1 after 60 mins by FRET assayMore data for this Ligand-Target Pair
Affinity DataIC50: 0.714nMAssay Description:The assay for the determination of Pim activity is based on the formation of phosphorylated biotinylated-BAD peptide at the Serine 112 residue (S112)...More data for this Ligand-Target Pair
Affinity DataIC50: 0.734nMAssay Description:The assay for the determination of Pim activity is based on the formation of phosphorylated biotinylated-BAD peptide at the Serine 112 residue (S112)...More data for this Ligand-Target Pair
Affinity DataIC50: 0.760nMAssay Description:TRPV1 activities were monitored as a function of cellular uptake of radioactive calcium (45Ca2+). The test compounds were preincubated with TRPV1 exp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.763nMAssay Description:The assay for the determination of Pim activity is based on the formation of phosphorylated biotinylated-BAD peptide at the Serine 112 residue (S112)...More data for this Ligand-Target Pair
Affinity DataIC50: 0.800nMpH: 7.4 T: 2°CAssay Description:TRPV1 activities were monitored as a function of cellular uptake of radioactive calcium (45Ca2+). The test compounds were preincubated with TRPV1 exp...More data for this Ligand-Target Pair
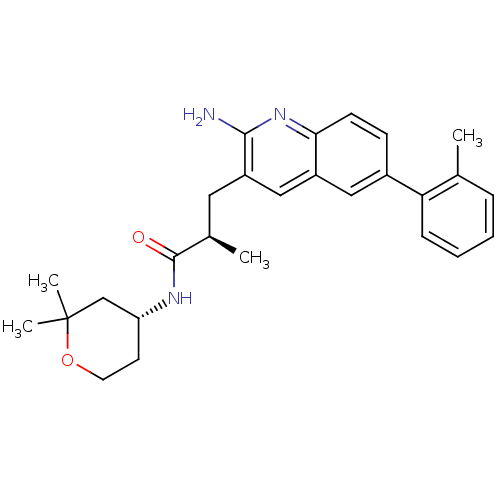
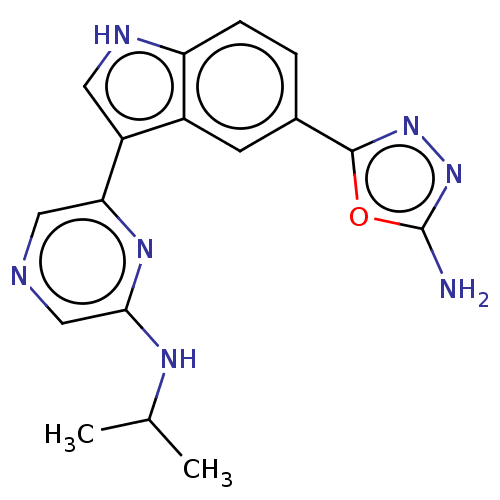
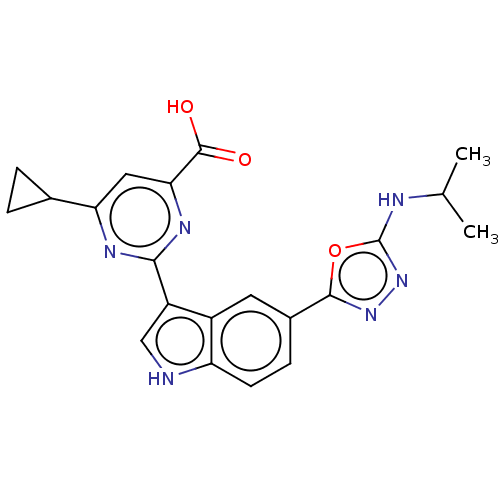
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Homo sapiens (Human))
Amgen
Curated by ChEMBL
Amgen
Curated by ChEMBL
Affinity DataIC50: 0.800nMAssay Description:Inhibition of human recombinant PDE10A using cAMP as substrate incubated for 30 mins prior to substrate addition measured after 3 hrs to overnight by...More data for this Ligand-Target Pair
TargetTransient receptor potential cation channel subfamily V member 1(Rattus norvegicus (rat))
Amgen
Amgen
Affinity DataIC50: 0.900nMAssay Description:TRPV1 activities were monitored as a function of cellular uptake of radioactive calcium (45Ca2+). The test compounds were preincubated with TRPV1 exp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.900nMAssay Description:TRPV1 activities were monitored as a function of cellular uptake of radioactive calcium (45Ca2+). The test compounds were preincubated with TRPV1 exp...More data for this Ligand-Target Pair
Affinity DataIC50: 0.901nMAssay Description:The assay for the determination of Pim activity is based on the formation of phosphorylated biotinylated-BAD peptide at the Serine 112 residue (S112)...More data for this Ligand-Target Pair
Affinity DataIC50: 0.974nMAssay Description:The assay for the determination of Pim activity is based on the formation of phosphorylated biotinylated-BAD peptide at the Serine 112 residue (S112)...More data for this Ligand-Target Pair
Affinity DataIC50: 0.984nMAssay Description:The assay for the determination of Pim activity is based on the formation of phosphorylated biotinylated-BAD peptide at the Serine 112 residue (S112)...More data for this Ligand-Target Pair
Affinity DataIC50: 0.998nMAssay Description:The assay for the determination of Pim activity is based on the formation of phosphorylated biotinylated-BAD peptide at the Serine 112 residue (S112)...More data for this Ligand-Target Pair
Affinity DataIC50: 1nMAssay Description:Inhibition of full length recombinant PIM3 (unknown origin) using biotinylated-BAD peptide as substrate assessed as phosphorylation at serine 112 aft...More data for this Ligand-Target Pair
Affinity DataIC50: 1nMpH: 7.5 T: 2°CAssay Description:The assay uses purified enzyme interacting with biotinylated peptide substrate. HTRF is based on the proximity of europium cryptate (donor fluorophor...More data for this Ligand-Target Pair
Affinity DataIC50: 1nMAssay Description:Inhibition of full length recombinant PIM3 (unknown origin) using biotinylated-BAD peptide as substrate assessed as phosphorylation at serine 112 aft...More data for this Ligand-Target Pair
Affinity DataIC50: 1nMAssay Description:Inhibition of full length recombinant PIM1 (unknown origin) using biotinylated-BAD peptide as substrate assessed as phosphorylation at serine 112 aft...More data for this Ligand-Target Pair
Affinity DataIC50: 1.01nMAssay Description:The assay for the determination of Pim activity is based on the formation of phosphorylated biotinylated-BAD peptide at the Serine 112 residue (S112)...More data for this Ligand-Target Pair
Affinity DataIC50: 1.05nMAssay Description:The assay for the determination of Pim activity is based on the formation of phosphorylated biotinylated-BAD peptide at the Serine 112 residue (S112)...More data for this Ligand-Target Pair
Affinity DataIC50: 1.06nMAssay Description:The assay for the determination of Pim activity is based on the formation of phosphorylated biotinylated-BAD peptide at the Serine 112 residue (S112)...More data for this Ligand-Target Pair
Affinity DataIC50: 1.07nMAssay Description:The assay for the determination of Pim activity is based on the formation of phosphorylated biotinylated-BAD peptide at the Serine 112 residue (S112)...More data for this Ligand-Target Pair
Affinity DataIC50: 1.08nMAssay Description:The assay for the determination of Pim activity is based on the formation of phosphorylated biotinylated-BAD peptide at the Serine 112 residue (S112)...More data for this Ligand-Target Pair
Affinity DataIC50: 1.11nMAssay Description:The assay for the determination of Pim activity is based on the formation of phosphorylated biotinylated-BAD peptide at the Serine 112 residue (S112)...More data for this Ligand-Target Pair
Affinity DataIC50: 1.15nMAssay Description:The assay for the determination of Pim activity is based on the formation of phosphorylated biotinylated-BAD peptide at the Serine 112 residue (S112)...More data for this Ligand-Target Pair
Affinity DataIC50: 1.16nMAssay Description:The assay for the determination of Pim activity is based on the formation of phosphorylated biotinylated-BAD peptide at the Serine 112 residue (S112)...More data for this Ligand-Target Pair
Affinity DataIC50: 1.16nMAssay Description:The assay for the determination of Pim activity is based on the formation of phosphorylated biotinylated-BAD peptide at the Serine 112 residue (S112)...More data for this Ligand-Target Pair
Affinity DataIC50: 1.19nMAssay Description:The assay for the determination of Pim activity is based on the formation of phosphorylated biotinylated-BAD peptide at the Serine 112 residue (S112)...More data for this Ligand-Target Pair
Affinity DataIC50: 1.20nMAssay Description:The assay for the determination of Pim activity is based on the formation of phosphorylated biotinylated-BAD peptide at the Serine 112 residue (S112)...More data for this Ligand-Target Pair

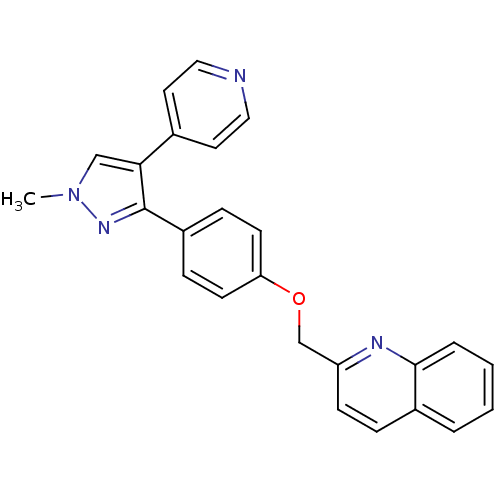
 3D Structure (crystal)
3D Structure (crystal)