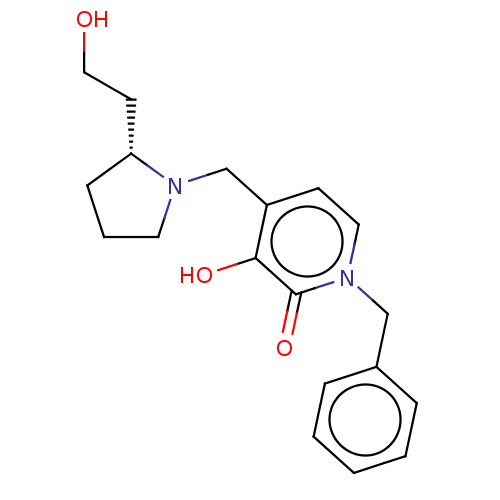
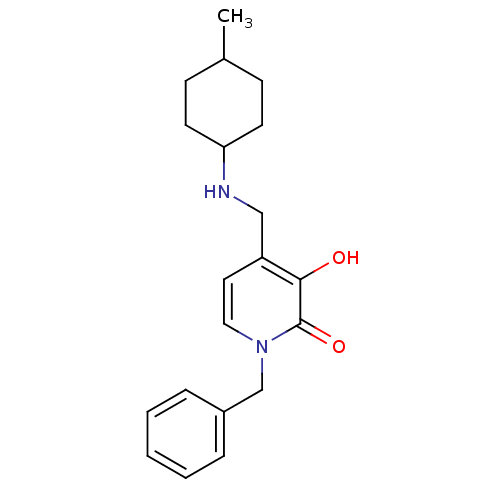
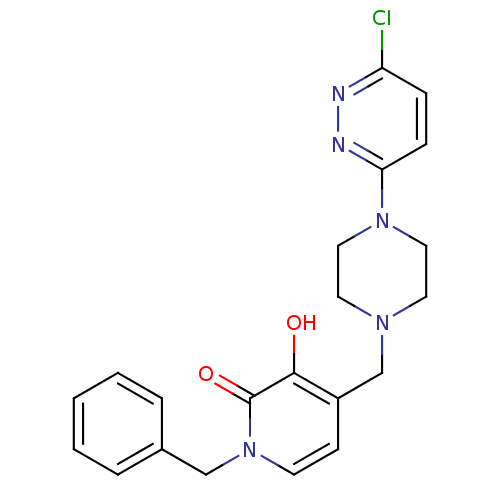
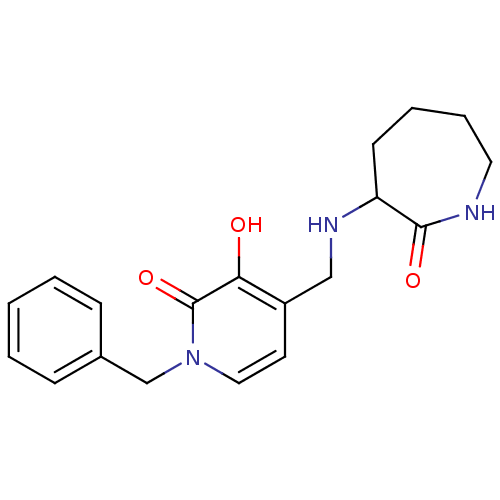
Affinity DataIC50: 1.20E+3nMpH: 7.5 T: 2°CAssay Description:The EGLN-1 (or EGLN-3) enzyme activity is determined using mass spectrometry (matrix-assisted laser desorption ionization, time-of-flight MS, MALDI-T...More data for this Ligand-Target Pair
Affinity DataIC50: 2.10E+3nMAssay Description:HEK293 cells are seeded in 96-well poly-lysine coated plates at 20,000 cells per well in DMEM (10% FBS, 1% NEAA, 0.1% glutamine). Following overnight...More data for this Ligand-Target Pair
Affinity DataIC50: 4.40E+3nMpH: 7.5 T: 2°CAssay Description:The EGLN-1 (or EGLN-3) enzyme activity is determined using mass spectrometry (matrix-assisted laser desorption ionization, time-of-flight MS, MALDI-T...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+3nMpH: 7.5 T: 2°CAssay Description:The EGLN-1 (or EGLN-3) enzyme activity is determined using mass spectrometry (matrix-assisted laser desorption ionization, time-of-flight MS, MALDI-T...More data for this Ligand-Target Pair
Affinity DataIC50: 7.40E+3nMAssay Description:HEK293 cells are seeded in 96-well poly-lysine coated plates at 20,000 cells per well in DMEM (10% FBS, 1% NEAA, 0.1% glutamine). Following overnight...More data for this Ligand-Target Pair
Affinity DataIC50: 9.00E+3nMpH: 7.5 T: 2°CAssay Description:The EGLN-1 (or EGLN-3) enzyme activity is determined using mass spectrometry (matrix-assisted laser desorption ionization, time-of-flight MS, MALDI-T...More data for this Ligand-Target Pair
Affinity DataIC50: 1.10E+4nMpH: 7.5 T: 2°CAssay Description:The EGLN-1 (or EGLN-3) enzyme activity is determined using mass spectrometry (matrix-assisted laser desorption ionization, time-of-flight MS, MALDI-T...More data for this Ligand-Target Pair
Affinity DataIC50: 1.10E+4nMpH: 7.5 T: 2°CAssay Description:The EGLN-1 (or EGLN-3) enzyme activity is determined using mass spectrometry (matrix-assisted laser desorption ionization, time-of-flight MS, MALDI-T...More data for this Ligand-Target Pair
Affinity DataIC50: 1.20E+4nMpH: 7.5 T: 2°CAssay Description:The EGLN-1 (or EGLN-3) enzyme activity is determined using mass spectrometry (matrix-assisted laser desorption ionization, time-of-flight MS, MALDI-T...More data for this Ligand-Target Pair
Affinity DataIC50: 1.20E+4nMpH: 7.5 T: 2°CAssay Description:The EGLN-1 (or EGLN-3) enzyme activity is determined using mass spectrometry (matrix-assisted laser desorption ionization, time-of-flight MS, MALDI-T...More data for this Ligand-Target Pair
Affinity DataIC50: 1.20E+4nMpH: 7.5 T: 2°CAssay Description:The EGLN-1 (or EGLN-3) enzyme activity is determined using mass spectrometry (matrix-assisted laser desorption ionization, time-of-flight MS, MALDI-T...More data for this Ligand-Target Pair
Affinity DataIC50: 1.20E+4nMpH: 7.5 T: 2°CAssay Description:The EGLN-1 (or EGLN-3) enzyme activity is determined using mass spectrometry (matrix-assisted laser desorption ionization, time-of-flight MS, MALDI-T...More data for this Ligand-Target Pair
Affinity DataIC50: 1.20E+4nMpH: 7.5 T: 2°CAssay Description:The EGLN-1 (or EGLN-3) enzyme activity is determined using mass spectrometry (matrix-assisted laser desorption ionization, time-of-flight MS, MALDI-T...More data for this Ligand-Target Pair
Affinity DataIC50: 1.26E+4nMAssay Description:HEK293 cells are seeded in 96-well poly-lysine coated plates at 20,000 cells per well in DMEM (10% FBS, 1% NEAA, 0.1% glutamine). Following overnight...More data for this Ligand-Target Pair
Affinity DataIC50: 1.40E+4nMpH: 7.5 T: 2°CAssay Description:The EGLN-1 (or EGLN-3) enzyme activity is determined using mass spectrometry (matrix-assisted laser desorption ionization, time-of-flight MS, MALDI-T...More data for this Ligand-Target Pair
Affinity DataIC50: 1.60E+4nMpH: 7.5 T: 2°CAssay Description:The EGLN-1 (or EGLN-3) enzyme activity is determined using mass spectrometry (matrix-assisted laser desorption ionization, time-of-flight MS, MALDI-T...More data for this Ligand-Target Pair
Affinity DataIC50: 1.66E+4nMAssay Description:HEK293 cells are seeded in 96-well poly-lysine coated plates at 20,000 cells per well in DMEM (10% FBS, 1% NEAA, 0.1% glutamine). Following overnight...More data for this Ligand-Target Pair
Affinity DataIC50: 1.70E+4nMpH: 7.5 T: 2°CAssay Description:The EGLN-1 (or EGLN-3) enzyme activity is determined using mass spectrometry (matrix-assisted laser desorption ionization, time-of-flight MS, MALDI-T...More data for this Ligand-Target Pair
Affinity DataIC50: 1.80E+4nMpH: 7.5 T: 2°CAssay Description:The EGLN-1 (or EGLN-3) enzyme activity is determined using mass spectrometry (matrix-assisted laser desorption ionization, time-of-flight MS, MALDI-T...More data for this Ligand-Target Pair
Affinity DataIC50: 1.90E+4nMAssay Description:HEK293 cells are seeded in 96-well poly-lysine coated plates at 20,000 cells per well in DMEM (10% FBS, 1% NEAA, 0.1% glutamine). Following overnight...More data for this Ligand-Target Pair
Affinity DataIC50: 2.06E+4nMAssay Description:HEK293 cells are seeded in 96-well poly-lysine coated plates at 20,000 cells per well in DMEM (10% FBS, 1% NEAA, 0.1% glutamine). Following overnight...More data for this Ligand-Target Pair
Affinity DataIC50: 2.10E+4nMpH: 7.5 T: 2°CAssay Description:The EGLN-1 (or EGLN-3) enzyme activity is determined using mass spectrometry (matrix-assisted laser desorption ionization, time-of-flight MS, MALDI-T...More data for this Ligand-Target Pair
Affinity DataIC50: 2.70E+4nMAssay Description:HEK293 cells are seeded in 96-well poly-lysine coated plates at 20,000 cells per well in DMEM (10% FBS, 1% NEAA, 0.1% glutamine). Following overnight...More data for this Ligand-Target Pair
Affinity DataIC50: 2.74E+4nMAssay Description:HEK293 cells are seeded in 96-well poly-lysine coated plates at 20,000 cells per well in DMEM (10% FBS, 1% NEAA, 0.1% glutamine). Following overnight...More data for this Ligand-Target Pair
Affinity DataIC50: 2.92E+4nMAssay Description:HEK293 cells are seeded in 96-well poly-lysine coated plates at 20,000 cells per well in DMEM (10% FBS, 1% NEAA, 0.1% glutamine). Following overnight...More data for this Ligand-Target Pair
Affinity DataIC50: 4.20E+4nMAssay Description:HEK293 cells are seeded in 96-well poly-lysine coated plates at 20,000 cells per well in DMEM (10% FBS, 1% NEAA, 0.1% glutamine). Following overnight...More data for this Ligand-Target Pair
Affinity DataIC50: 4.25E+4nMAssay Description:HEK293 cells are seeded in 96-well poly-lysine coated plates at 20,000 cells per well in DMEM (10% FBS, 1% NEAA, 0.1% glutamine). Following overnight...More data for this Ligand-Target Pair
Affinity DataIC50: 5.30E+4nMAssay Description:HEK293 cells are seeded in 96-well poly-lysine coated plates at 20,000 cells per well in DMEM (10% FBS, 1% NEAA, 0.1% glutamine). Following overnight...More data for this Ligand-Target Pair
Affinity DataIC50: 5.30E+4nMAssay Description:HEK293 cells are seeded in 96-well poly-lysine coated plates at 20,000 cells per well in DMEM (10% FBS, 1% NEAA, 0.1% glutamine). Following overnight...More data for this Ligand-Target Pair
Affinity DataIC50: 6.29E+4nMAssay Description:HEK293 cells are seeded in 96-well poly-lysine coated plates at 20,000 cells per well in DMEM (10% FBS, 1% NEAA, 0.1% glutamine). Following overnight...More data for this Ligand-Target Pair
Affinity DataIC50: 7.80E+4nMAssay Description:HEK293 cells are seeded in 96-well poly-lysine coated plates at 20,000 cells per well in DMEM (10% FBS, 1% NEAA, 0.1% glutamine). Following overnight...More data for this Ligand-Target Pair
Affinity DataIC50: >1.00E+5nMAssay Description:HEK293 cells are seeded in 96-well poly-lysine coated plates at 20,000 cells per well in DMEM (10% FBS, 1% NEAA, 0.1% glutamine). Following overnight...More data for this Ligand-Target Pair